

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
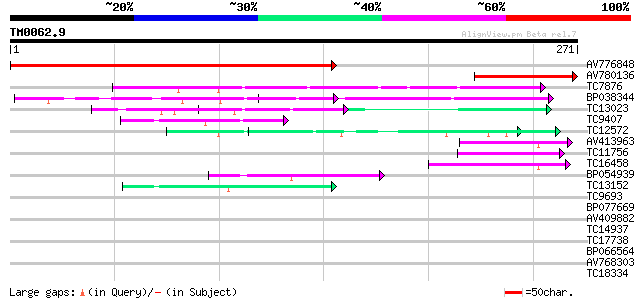
Query= TM0062.9
(271 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV776848 293 2e-80
AV780136 94 2e-20
TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-bi... 56 7e-09
BP038344 52 1e-07
TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragm... 47 3e-06
TC9407 weakly similar to UP|TAF5_YEAST (P38129) Transcription in... 45 1e-05
TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory pro... 45 2e-05
AV413963 44 3e-05
TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snR... 42 1e-04
TC16458 similar to UP|O48679 (O48679) F3I6.5 protein, partial (20%) 40 4e-04
BP054939 40 4e-04
TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial... 40 5e-04
TC9693 similar to UP|COPP_MOUSE (O55029) Coatomer beta' subunit ... 39 0.001
BP077669 39 0.001
AV409882 38 0.002
TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5,... 38 0.002
TC17738 similar to UP|O48679 (O48679) F3I6.5 protein, partial (8%) 35 0.010
BP066564 35 0.010
AV768303 35 0.018
TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1 (AT3... 35 0.018
>AV776848
Length = 584
Score = 293 bits (750), Expect = 2e-80
Identities = 139/156 (89%), Positives = 147/156 (94%)
Frame = +3
Query: 1 MHRVGSAGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSR 60
MHRVGSAGNT++STRPRKEKR TYVLNDSD TKHCAGINCLA+L SA SDGSDYLFTGSR
Sbjct: 117 MHRVGSAGNTSNSTRPRKEKRFTYVLNDSDGTKHCAGINCLALLTSAASDGSDYLFTGSR 296
Query: 61 DGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTR 120
DGRLKRWAL EDA TCSATFESHVDWVNDAVLVGD+TLVSCSSDTTLKTW+AFS G+CTR
Sbjct: 297 DGRLKRWALAEDAVTCSATFESHVDWVNDAVLVGDNTLVSCSSDTTLKTWNAFSVGSCTR 476
Query: 121 TLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDL 156
TLRQHSDYVTCLAA+ KN NIVASGGLGGEVF+WDL
Sbjct: 477 TLRQHSDYVTCLAAAGKNCNIVASGGLGGEVFIWDL 584
>AV780136
Length = 558
Score = 94.4 bits (233), Expect = 2e-20
Identities = 44/49 (89%), Positives = 48/49 (97%)
Frame = -1
Query: 223 ALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDST 271
ALAMNEGGT+LVSGGTEKV+RVWDPRSGSKT+KL+GH DNIRALLLDST
Sbjct: 558 ALAMNEGGTVLVSGGTEKVIRVWDPRSGSKTLKLRGHADNIRALLLDST 412
Score = 35.0 bits (79), Expect = 0.014
Identities = 15/52 (28%), Positives = 29/52 (54%)
Frame = -1
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+GH +++ AL ++ G +SG ++ ++R+WD HTD++ AL
Sbjct: 456 RGHADNIRALLLDSTGRYCLSGSSDSMIRLWDIGQPRCLHTYAVHTDSVWAL 301
>TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1379
Score = 55.8 bits (133), Expect = 7e-09
Identities = 60/211 (28%), Positives = 88/211 (41%), Gaps = 4/211 (1%)
Frame = +3
Query: 50 DGSDYLFTGSRDGRLKRWALGEDAATCSAT---FESHVDWVNDAVLVGDSTL-VSCSSDT 105
D SD + T SRD + W L ++ T H +V D VL D +S S D
Sbjct: 147 DNSDMIVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHFVQDVVLSSDGQFALSGSWDG 326
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
L+ WD + GT R H+ V +A S N IV S + +W+ +
Sbjct: 327 ELRLWD-LAAGTSARRFVGHTKDVLSVAFSIDNRQIV-SASRDRTIKLWNTLGECKYTIQ 500
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
DA D S + S ++L T + S S ++ V N A GH V +A
Sbjct: 501 DNDAHSDWVSC-VRFSPSTLQPTIV-SASWDRTVKVWNLTNCKLRNTLA-GHSGYVNTVA 671
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKL 256
++ G+L SGG + V+ +WD G + L
Sbjct: 672 VSPDGSLCASGGKDGVILLWDLAEGKRLYSL 764
Score = 38.5 bits (88), Expect = 0.001
Identities = 29/143 (20%), Positives = 53/143 (36%)
Frame = +3
Query: 121 TLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGING 180
T+R H+D VT +A NS+++ + + +W L T +D + G+
Sbjct: 99 TMRAHTDQVTAIATPIDNSDMIVTASRDKSIILWHL-------------TKEDKTYGV-- 233
Query: 181 SGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEK 240
P L GH V + ++ G +SG +
Sbjct: 234 -----PRRRLT------------------------GHSHFVQDVVLSSDGQFALSGSWDG 326
Query: 241 VVRVWDPRSGSKTMKLKGHTDNI 263
+R+WD +G+ + GHT ++
Sbjct: 327 ELRLWDLAAGTSARRFVGHTKDV 395
>BP038344
Length = 525
Score = 51.6 bits (122), Expect = 1e-07
Identities = 50/159 (31%), Positives = 72/159 (44%), Gaps = 4/159 (2%)
Frame = +2
Query: 3 RVGSAGNTNSSTRPR-KEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRD 61
RV +A NSS P K R L T H A + C+ S+ L + S D
Sbjct: 14 RVRAAAMANSSDEPPYKPYRHLKTL-----TAHDAAVACVKF-----SNDGTLLASASLD 163
Query: 62 GRLKRWALGEDAATCSATFE--SHVDWVNDAVLVGDSTLV-SCSSDTTLKTWDAFSTGTC 118
L W+ +AT S H + ++D DS + S S D TL+ WDA + G C
Sbjct: 164 KTLIIWS----SATLSLLHRLTGHSEGISDLAWSSDSHYICSASDDRTLRIWDA-TGGDC 328
Query: 119 TRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE 157
+TLR H+ V C+ + + SN + SG V VW+++
Sbjct: 329 VKTLRGHTHAVFCVNFNPQ-SNYIVSGSFDETVRVWEVK 442
Score = 39.7 bits (91), Expect = 5e-04
Identities = 32/141 (22%), Positives = 58/141 (40%)
Frame = +2
Query: 120 RTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGIN 179
+TL H V C+ S + ++AS L + +W +SA + S GI+
Sbjct: 80 KTLTAHDAAVACVKFSN-DGTLLASASLDKTLIIW------SSATLSLLHRLTGHSEGIS 238
Query: 180 GSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTE 239
S + S S ++ + + +GH +V+ + N +VSG +
Sbjct: 239 DLAWSSDSHYICSASDDRTLRIWDATGGDCVK-TLRGHTHAVFCVNFNPQSNYIVSGSFD 415
Query: 240 KVVRVWDPRSGSKTMKLKGHT 260
+ VRVW+ ++G + HT
Sbjct: 416 ETVRVWEVKTGRCIHTIIAHT 478
Score = 32.3 bits (72), Expect = 0.088
Identities = 16/54 (29%), Positives = 26/54 (47%)
Frame = +2
Query: 217 HKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
H +V + + GTLL S +K + +W + S +L GH++ I L S
Sbjct: 95 HDAAVACVKFSNDGTLLASASLDKTLIIWSSATLSLLHRLTGHSEGISDLAWSS 256
>TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragment),
partial (18%)
Length = 591
Score = 47.4 bits (111), Expect = 3e-06
Identities = 41/169 (24%), Positives = 67/169 (39%)
Frame = +1
Query: 91 VLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGE 150
V+ D + ++C+ ++K D+ + + TL+ S+ VT LA S + N++ S G +
Sbjct: 181 VVSSDGSFIACACGESIKIVDS-ANASIRSTLQGDSESVTALALSP-DDNLLFSSGHSRQ 354
Query: 151 VFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYI 210
+ VWDL S KC+ +
Sbjct: 355 IRVWDL-----STLKCVRSW---------------------------------------- 399
Query: 211 PIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
KGH+ V ++ + G LL +GG ++ V VWD G T KGH
Sbjct: 400 ----KGHEGPVMCMSCHPSGGLLATGGADRKVLVWDVDGGYCTHFFKGH 534
Score = 41.2 bits (95), Expect = 2e-04
Identities = 34/131 (25%), Positives = 59/131 (44%), Gaps = 8/131 (6%)
Frame = +1
Query: 40 CLAVLKSAVSDGSDYLFTGSRDGRLKRWALGE-----DAATCS--ATFESHVDWVNDAVL 92
C+ L+ + G + S DG A GE D+A S +T + + V L
Sbjct: 139 CVPALQQFYTGGP---YVVSSDGSFIACACGESIKIVDSANASIRSTLQGDSESVTALAL 309
Query: 93 VGDSTLVSCSSDTT-LKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEV 151
D L+ S + ++ WD ST C R+ + H V C++ + ++A+GG +V
Sbjct: 310 SPDDNLLFSSGHSRQIRVWD-LSTLKCVRSWKGHEGPVMCMSC-HPSGGLLATGGADRKV 483
Query: 152 FVWDLEAAHAS 162
VWD++ + +
Sbjct: 484 LVWDVDGGYCT 516
>TC9407 weakly similar to UP|TAF5_YEAST (P38129) Transcription initiation
factor TFIID subunit 5 (TBP-associated factor 5)
(TBP-associated factor 90 kDa) (TAFII-90), partial (4%)
Length = 530
Score = 45.4 bits (106), Expect = 1e-05
Identities = 30/81 (37%), Positives = 39/81 (48%), Gaps = 1/81 (1%)
Frame = +2
Query: 54 YLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVL-VGDSTLVSCSSDTTLKTWDA 112
YL T S D +K W + D T T H WV D V V + L++ SSDTT + W +
Sbjct: 56 YLATASADHTVKIWNV--DGFTLDKTLIGHQRWVWDCVFSVDGAYLITASSDTTARLW-S 226
Query: 113 FSTGTCTRTLRQHSDYVTCLA 133
STG + + H TC A
Sbjct: 227 MSTGEDIKVYQGHHKATTCCA 289
Score = 26.9 bits (58), Expect = 3.7
Identities = 10/44 (22%), Positives = 20/44 (44%)
Frame = +2
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
GH+ V+ + G L++ ++ R+W +G +GH
Sbjct: 134 GHQRWVWDCVFSVDGAYLITASSDTTARLWSMSTGEDIKVYQGH 265
>TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (51%)
Length = 770
Score = 44.7 bits (104), Expect = 2e-05
Identities = 43/181 (23%), Positives = 73/181 (39%), Gaps = 11/181 (6%)
Frame = +3
Query: 76 CSATFESHVDWVNDAVLVGDSTL-VSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAA 134
C +H D V D +L VS S D + WDA STG C +TL +
Sbjct: 15 CLKVLPAHSDPVTAVDFNRDGSLIVSSSYDGLCRIWDA-STGHCMKTLIDDENPPVSFVK 191
Query: 135 SEKNSNIVASGGLGGEVFVWDLE--------AAHASAAKCIDATDDDTSNGINGSGNSLP 186
N+ + G L + +W+ H ++ CI +T T+NG G
Sbjct: 192 FSPNAKFILVGTLDNNLRLWNYSTGRFLKTYTGHVNSKYCISST-FSTTNGKYVVGG--- 359
Query: 187 LTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSG--GTEKVVRV 244
S + I + Q + + +GH ++V +++ + ++ SG G +K V++
Sbjct: 360 -------SEDHGIYLWELQTRKIVQ-KLEGHSDTVVSVSCHPTENMIASGALGNDKTVKI 515
Query: 245 W 245
W
Sbjct: 516 W 518
Score = 43.5 bits (101), Expect = 4e-05
Identities = 38/160 (23%), Positives = 64/160 (39%), Gaps = 11/160 (6%)
Frame = +3
Query: 115 TGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDT 174
+G C + L HSD VT + + S IV+S G +WD H C+ DD
Sbjct: 6 SGKCLKVLPAHSDPVTAVDFNRDGSLIVSSS-YDGLCRIWDASTGH-----CMKTLIDDE 167
Query: 175 SNGINGSGNSLPLTSLRSISSSNSISVHTTQNQ--------GYIPIAAKGHKESVYALAM 226
+ P++ ++ ++ I V T N G GH S Y ++
Sbjct: 168 NP---------PVSFVKFSPNAKFILVGTLDNNLRLWNYSTGRFLKTYTGHVNSKYCISS 320
Query: 227 N---EGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
G +V G + + +W+ ++ KL+GH+D +
Sbjct: 321 TFSTTNGKYVVGGSEDHGIYLWELQTRKIVQKLEGHSDTV 440
>AV413963
Length = 313
Score = 43.9 bits (102), Expect = 3e-05
Identities = 22/58 (37%), Positives = 33/58 (55%), Gaps = 4/58 (6%)
Frame = +2
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGS----KTMKLKGHTDNIRALLLD 269
GHK+ V+++A N GT L SG ++ R+W K ++LKGHTD++ L D
Sbjct: 71 GHKKKVHSVAWNCIGTKLASGSVDQTARIWHVEQHGHGKVKDIELKGHTDSVDQLCWD 244
>TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snRNP-specific
40 kDa protein (hPrp8-binding) {Homo sapiens;} , partial
(43%)
Length = 719
Score = 42.0 bits (97), Expect = 1e-04
Identities = 19/51 (37%), Positives = 27/51 (52%)
Frame = +1
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRA 265
KGHK +V L GT +VS +K VRVWD +G + K+ H + +
Sbjct: 424 KGHKNAVLDLHWTTDGTQIVSASPDKTVRVWDVETGKQVKKMVEHLSYVNS 576
Score = 35.8 bits (81), Expect = 0.008
Identities = 15/49 (30%), Positives = 27/49 (54%), Gaps = 1/49 (2%)
Frame = +1
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKT-MKLKGHTDNI 263
GH+ +Y + N GT++ SG ++ + +W+ K M LKGH + +
Sbjct: 298 GHQSVIYTMKFNPAGTVIASGSHDREIFLWNVHGECKNFMVLKGHKNAV 444
>TC16458 similar to UP|O48679 (O48679) F3I6.5 protein, partial (20%)
Length = 539
Score = 40.0 bits (92), Expect = 4e-04
Identities = 23/74 (31%), Positives = 38/74 (51%), Gaps = 6/74 (8%)
Frame = +3
Query: 201 VHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGS------KTM 254
VH + + + + HK +V ALA+NE G++L SG ++ + VW+ G
Sbjct: 6 VHAGEKKHSLLETLEKHKSAVNALALNEDGSVLYSGACDRSILVWERDGGDGGGEMVVVG 185
Query: 255 KLKGHTDNIRALLL 268
L+GHT I L++
Sbjct: 186 ALRGHTKAILCLVV 227
>BP054939
Length = 514
Score = 40.0 bits (92), Expect = 4e-04
Identities = 26/87 (29%), Positives = 42/87 (47%), Gaps = 3/87 (3%)
Frame = +1
Query: 96 STLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLA---ASEKNSNIVASGGLGGEVF 152
+ LV+ S D +++ W G+C R LR H+ V L+ E +S ++ASGG V
Sbjct: 178 NVLVTSSCDHSIRLW---RKGSCLRCLRGHNGPVLSLSNKLLGEGSSKVLASGGEDSTVR 348
Query: 153 VWDLEAAHASAAKCIDATDDDTSNGIN 179
+W L ++ + AT G+N
Sbjct: 349 LWSLGSSGKRGQHALKATFYGHEKGVN 429
Score = 26.6 bits (57), Expect = 4.8
Identities = 15/42 (35%), Positives = 23/42 (54%), Gaps = 4/42 (9%)
Frame = +1
Query: 215 KGHKESVYALA---MNEGGT-LLVSGGTEKVVRVWDPRSGSK 252
+GH V +L+ + EG + +L SGG + VR+W S K
Sbjct: 250 RGHNGPVLSLSNKLLGEGSSKVLASGGEDSTVRLWSLGSSGK 375
>TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial (60%)
Length = 453
Score = 39.7 bits (91), Expect = 5e-04
Identities = 28/103 (27%), Positives = 41/103 (39%), Gaps = 1/103 (0%)
Frame = +2
Query: 55 LFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSS-DTTLKTWDAF 113
L +G D + W D+ AT E H + D ++ SS D T+K WD
Sbjct: 5 LASGGHDKKAVLWYT--DSLKQKATLEEHSALITDVRFSPSMPRLATSSFDKTVKVWDVD 178
Query: 114 STGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDL 156
+ G RT HS V L +++ S GE+ W +
Sbjct: 179 NPGYSLRTFTGHSASVMSLDFHPNKEDLICSCDSDGEIRYWSI 307
>TC9693 similar to UP|COPP_MOUSE (O55029) Coatomer beta' subunit
(Beta'-coat protein) (Beta'-COP) (p102), partial (20%)
Length = 743
Score = 38.5 bits (88), Expect = 0.001
Identities = 24/78 (30%), Positives = 38/78 (47%), Gaps = 1/78 (1%)
Frame = +1
Query: 80 FESHVDWVND-AVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKN 138
FE+H D++ AV ++S S D +K WD CT+ HS YV + + K+
Sbjct: 490 FEAHTDYIRCVAVHPTLPYVLSSSDDMLIKLWDWEKGWICTQICEAHSHYVMQVTFNPKD 669
Query: 139 SNIVASGGLGGEVFVWDL 156
+N AS L + + +L
Sbjct: 670 TNTFASASLDRTIRIRNL 723
>BP077669
Length = 488
Score = 38.5 bits (88), Expect = 0.001
Identities = 24/70 (34%), Positives = 31/70 (44%), Gaps = 1/70 (1%)
Frame = -2
Query: 43 VLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST-LVSC 101
+LK ++ L T S D ++ W L T H WV D DS+ LV+
Sbjct: 310 ILKCKIAPNCTQLATCSSDSTVRLWDLTSQCRL-QKTLNGHKKWVWDCTYNSDSSYLVTA 134
Query: 102 SSDTTLKTWD 111
SSD T K WD
Sbjct: 133 SSDLTAKLWD 104
Score = 35.4 bits (80), Expect = 0.010
Identities = 20/59 (33%), Positives = 31/59 (51%), Gaps = 1/59 (1%)
Frame = -2
Query: 98 LVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYV-TCLAASEKNSNIVASGGLGGEVFVWD 155
L +CSSD+T++ WD S +TL H +V C S+ + + AS L + +WD
Sbjct: 274 LATCSSDSTVRLWDLTSQCRLQKTLNGHKKWVWDCTYNSDSSYLVTASSDLTAK--LWD 104
Score = 31.6 bits (70), Expect = 0.15
Identities = 16/70 (22%), Positives = 32/70 (44%)
Frame = -2
Query: 188 TSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDP 247
T L + SS +++ + +Q + GHK+ V+ N + LV+ ++ ++WD
Sbjct: 280 TQLATCSSDSTVRLWDLTSQCRLQKTLNGHKKWVWDCTYNSDSSYLVTASSDLTAKLWDC 101
Query: 248 RSGSKTMKLK 257
G + K
Sbjct: 100 ARGEYIKEYK 71
>AV409882
Length = 422
Score = 38.1 bits (87), Expect = 0.002
Identities = 23/69 (33%), Positives = 34/69 (48%), Gaps = 1/69 (1%)
Frame = +3
Query: 80 FESHVDWVND-AVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKN 138
FE+H D++ AV ++S S D +K WD CT+ HS YV + + K+
Sbjct: 210 FEAHTDYIRCVAVHPTLPYVLSSSDDMLIKLWDWEKGWICTQIFEGHSHYVMQVTFNPKD 389
Query: 139 SNIVASGGL 147
+N AS L
Sbjct: 390 TNTFASASL 416
>TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5, TAC
clone:K8K14 (AT5g67320/K8K14_4), partial (19%)
Length = 1089
Score = 37.7 bits (86), Expect = 0.002
Identities = 21/72 (29%), Positives = 35/72 (48%), Gaps = 1/72 (1%)
Frame = +1
Query: 180 GSGNSLPLTSLRSISSSNSISVHTTQNQ-GYIPIAAKGHKESVYALAMNEGGTLLVSGGT 238
G G S P L S+S +V + G + + GH++ VY++A + G L SG
Sbjct: 19 GPGTSNPNKKLVLASASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFSPNGEYLASGSP 198
Query: 239 EKVVRVWDPRSG 250
+K + +W + G
Sbjct: 199 DKSIHIWSLKEG 234
Score = 26.6 bits (57), Expect = 4.8
Identities = 10/32 (31%), Positives = 15/32 (46%)
Frame = +1
Query: 232 LLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
+L S + V++WD G L GH D +
Sbjct: 52 VLASASFDSTVKLWDVEVGKLIYSLNGHRDGV 147
>TC17738 similar to UP|O48679 (O48679) F3I6.5 protein, partial (8%)
Length = 599
Score = 35.4 bits (80), Expect = 0.010
Identities = 35/120 (29%), Positives = 51/120 (42%), Gaps = 11/120 (9%)
Frame = +2
Query: 55 LFTGSRDGRLKRWALGEDA--ATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDA 112
LF+G+ D + W + A S H + + V D L+S S+D T++ W
Sbjct: 11 LFSGACDRSILVWEREDSANHMVVSGALRGHEKAILCLINVSDF-LLSGSADRTVRVWKR 187
Query: 113 FSTGT--CTRTLRQHSDYVTCLAA---SEKNSN----IVASGGLGGEVFVWDLEAAHASA 163
G C L H V LAA S++NS + SG L GE+ VW + +A
Sbjct: 188 GFDGPFCCLAVLDGHRKPVKSLAAVAESDQNSPDGVVSIFSGSLDGEIRVWQVSLTSLTA 367
>BP066564
Length = 492
Score = 35.4 bits (80), Expect = 0.010
Identities = 18/44 (40%), Positives = 23/44 (51%), Gaps = 2/44 (4%)
Frame = +3
Query: 228 EGGTLLVSGGTEKVVRVWDP--RSGSKTMKLKGHTDNIRALLLD 269
E +SG T+ V++WDP R LKGHT IRA+ D
Sbjct: 30 EDAGFFISGSTDCSVKIWDPSLRGSELRATLKGHTRTIRAISSD 161
>AV768303
Length = 550
Score = 34.7 bits (78), Expect = 0.018
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = +1
Query: 217 HKESVYALAMNEGGTLLVSGGTEKVVRVWDP 247
HK+S+ A+ G L VS G +K V++W P
Sbjct: 13 HKDSIRAIRFAANGKLFVSAGDDKTVKIWSP 105
>TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1
(AT3g18860/MCB22_3), partial (21%)
Length = 595
Score = 34.7 bits (78), Expect = 0.018
Identities = 27/86 (31%), Positives = 39/86 (44%), Gaps = 8/86 (9%)
Frame = +2
Query: 83 HVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTG-TCTRTLRQHSDYVTCLAASEKNSNI 141
H D V + G+ + + S D T++ W + ++ L HS +V L N +
Sbjct: 146 HEDDVRGICVCGNDGIATSSRDKTVRFWSPENRKFVSSKVLVGHSSFVGPLVWIPPNPEL 325
Query: 142 ----VASGGLGGEVFVWDL---EAAH 160
VASGG+ V VWDL E AH
Sbjct: 326 PQGGVASGGMDTLVLVWDLSTGEKAH 403
Score = 34.3 bits (77), Expect = 0.023
Identities = 29/106 (27%), Positives = 46/106 (43%), Gaps = 8/106 (7%)
Frame = +2
Query: 172 DDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAK-GHKESVYALA----- 225
+D GI GN TS R ++ + +N+ ++ GH V L
Sbjct: 149 EDDVRGICVCGNDGIATSSRD----KTVRFWSPENRKFVSSKVLVGHSSFVGPLVWIPPN 316
Query: 226 --MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
+ +GG + SGG + +V VWD +G K LKGH + ++ D
Sbjct: 317 PELPQGG--VASGGMDTLVLVWDLSTGEKAHTLKGHQLQVTSIAFD 448
Score = 33.5 bits (75), Expect = 0.039
Identities = 25/89 (28%), Positives = 37/89 (41%), Gaps = 6/89 (6%)
Frame = +2
Query: 51 GSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST------LVSCSSD 104
G+D + T SRD ++ W+ S H +V V + + + S D
Sbjct: 179 GNDGIATSSRDKTVRFWSPENRKFVSSKVLVGHSSFVGPLVWIPPNPELPQGGVASGGMD 358
Query: 105 TTLKTWDAFSTGTCTRTLRQHSDYVTCLA 133
T + WD STG TL+ H VT +A
Sbjct: 359 TLVLVWD-LSTGEKAHTLKGHQLQVTSIA 442
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.127 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,609,390
Number of Sequences: 28460
Number of extensions: 62363
Number of successful extensions: 340
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 294
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 333
length of query: 271
length of database: 4,897,600
effective HSP length: 89
effective length of query: 182
effective length of database: 2,364,660
effective search space: 430368120
effective search space used: 430368120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0062.9


