

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
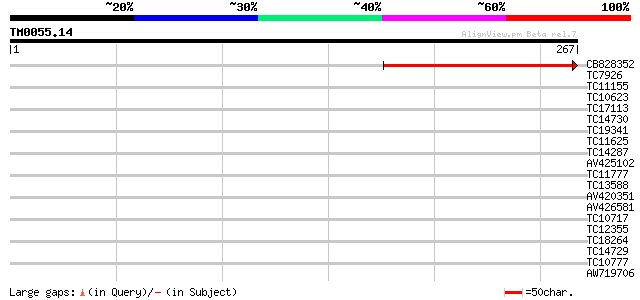
Query= TM0055.14
(267 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CB828352 191 8e-50
TC7926 homologue to UP|PGKH_TOBAC (Q42961) Phosphoglycerate kina... 35 0.013
TC11155 similar to UP|O24447 (O24447) Carbamoyl phosphate synthe... 35 0.017
TC10623 similar to GB|AAK98802.1|27372514|AY045774 tubby-like pr... 33 0.050
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 33 0.050
TC14730 similar to UP|Q9LG49 (Q9LG49) ESTs AU029606(E31139), par... 33 0.050
TC19341 similar to UP|Q9CAD1 (Q9CAD1) GMP synthase, partial (28%) 33 0.066
TC11625 weakly similar to UP|O81015 (O81015) At2g26920 protein, ... 33 0.066
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 32 0.086
AV425102 32 0.11
TC11777 similar to GB|AAM78073.1|21928089|AY125563 AT4g24660/F22... 32 0.15
TC13588 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall... 32 0.15
AV420351 31 0.19
AV426581 31 0.19
TC10717 similar to UP|Q8S564 (Q8S564) 4-coumarate:coenzyme A lig... 31 0.19
TC12355 similar to UP|PR18_HUMAN (Q99633) Pre-mRNA splicing fact... 31 0.19
TC18264 similar to UP|Q9SP93 (Q9SP93) Thiol protease, partial (4%) 31 0.25
TC14729 homologue to UP|Q8W590 (Q8W590) At2g32080/F22D22.17, par... 31 0.25
TC10777 similar to UP|Q9SCL3 (Q9SCL3) Pre-mRNA splicing factor S... 31 0.25
AW719706 31 0.25
>CB828352
Length = 496
Score = 191 bits (486), Expect = 8e-50
Identities = 91/91 (100%), Positives = 91/91 (100%)
Frame = +3
Query: 177 DEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQEC 236
DEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQEC
Sbjct: 3 DEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQEC 182
Query: 237 FAVDVKVKAALQEPNTEAWKRLCVNFLKDRL 267
FAVDVKVKAALQEPNTEAWKRLCVNFLKDRL
Sbjct: 183 FAVDVKVKAALQEPNTEAWKRLCVNFLKDRL 275
>TC7926 homologue to UP|PGKH_TOBAC (Q42961) Phosphoglycerate kinase,
chloroplast precursor , partial (86%)
Length = 1857
Score = 35.0 bits (79), Expect = 0.013
Identities = 13/18 (72%), Positives = 15/18 (83%)
Frame = -1
Query: 10 RELEMESGCGGGGGGGGG 27
RE++ GCGGGGGGGGG
Sbjct: 222 REVDAGGGCGGGGGGGGG 169
Score = 28.1 bits (61), Expect = 1.6
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = -1
Query: 13 EMESGCGGGGGGGGGKG 29
E+++G G GGGGGGG G
Sbjct: 219 EVDAGGGCGGGGGGGGG 169
>TC11155 similar to UP|O24447 (O24447) Carbamoyl phosphate synthetase small
subunit , partial (30%)
Length = 902
Score = 34.7 bits (78), Expect = 0.017
Identities = 17/37 (45%), Positives = 23/37 (61%), Gaps = 2/37 (5%)
Frame = +3
Query: 111 VHTLDSINKKI--LGICFGHQILGRVLGGKVGRSSTG 145
V T+ +I K+ GIC GHQ+LG+ LGGK + G
Sbjct: 30 VETVKNIIGKVPVFGICMGHQLLGQALGGKTFKMKFG 140
>TC10623 similar to GB|AAK98802.1|27372514|AY045774 tubby-like protein 3
{Arabidopsis thaliana;} , partial (27%)
Length = 692
Score = 33.1 bits (74), Expect = 0.050
Identities = 13/22 (59%), Positives = 17/22 (77%)
Frame = +3
Query: 11 ELEMESGCGGGGGGGGGKGKRY 32
E+E+ G GGGG GGGG G+R+
Sbjct: 288 EIEVAEGGGGGGRGGGGGGRRW 353
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 33.1 bits (74), Expect = 0.050
Identities = 16/38 (42%), Positives = 22/38 (57%)
Frame = -2
Query: 15 ESGCGGGGGGGGGKGKRYALLMCGEDSEYLLKKHGGCF 52
+ G GGGGGGGG G + + + G+ + K GGCF
Sbjct: 788 KGGKNGGGGGGGGGGVKPGITIGGKGN----KGEGGCF 687
>TC14730 similar to UP|Q9LG49 (Q9LG49) ESTs AU029606(E31139), partial (59%)
Length = 781
Score = 33.1 bits (74), Expect = 0.050
Identities = 16/33 (48%), Positives = 17/33 (51%)
Frame = +3
Query: 5 TMLLERELEMESGCGGGGGGGGGKGKRYALLMC 37
T L EME GGGGGGG G G L+C
Sbjct: 168 TSLSSGAAEMEGNSGGGGGGGSGGGGSDVELLC 266
>TC19341 similar to UP|Q9CAD1 (Q9CAD1) GMP synthase, partial (28%)
Length = 484
Score = 32.7 bits (73), Expect = 0.066
Identities = 21/79 (26%), Positives = 42/79 (52%), Gaps = 2/79 (2%)
Frame = +3
Query: 121 ILGICFGHQILGRVLGG--KVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDE 178
+LGIC+G Q+L + LGG +VG + + + T++ S+ + + + ++ H DE
Sbjct: 201 VLGICYGLQLLVQRLGGDVRVGHTQEYGRMEI-TVEHPSALFPSYKVGHKQVVWMSHGDE 377
Query: 179 VRNLPPKAEIIGWSEKTGI 197
LPP ++ S++ +
Sbjct: 378 AAALPPGFNVVARSQQGSV 434
>TC11625 weakly similar to UP|O81015 (O81015) At2g26920 protein, partial
(8%)
Length = 410
Score = 32.7 bits (73), Expect = 0.066
Identities = 14/21 (66%), Positives = 15/21 (70%)
Frame = +2
Query: 9 ERELEMESGCGGGGGGGGGKG 29
ERE + G GGGGGGGGG G
Sbjct: 236 ERERSVVDGGGGGGGGGGGGG 298
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(75%)
Length = 1999
Score = 32.3 bits (72), Expect = 0.086
Identities = 20/42 (47%), Positives = 22/42 (51%), Gaps = 3/42 (7%)
Frame = -1
Query: 15 ESGCGGGGGGGGGK---GKRYALLMCGEDSEYLLKKHGGCFG 53
E G GGGGGGGGG G+R GED Y K+ G G
Sbjct: 385 EEGGGGGGGGGGGVAS*GER*GRF*RGEDGRYPDKRDFGI*G 260
>AV425102
Length = 426
Score = 32.0 bits (71), Expect = 0.11
Identities = 14/22 (63%), Positives = 15/22 (67%)
Frame = -1
Query: 10 RELEMESGCGGGGGGGGGKGKR 31
R E E GGGGGGGGG G+R
Sbjct: 111 RGYEAEISGGGGGGGGGGGGRR 46
>TC11777 similar to GB|AAM78073.1|21928089|AY125563 AT4g24660/F22K18_140
{Arabidopsis thaliana;}, partial (35%)
Length = 588
Score = 31.6 bits (70), Expect = 0.15
Identities = 15/35 (42%), Positives = 18/35 (50%)
Frame = +3
Query: 11 ELEMESGCGGGGGGGGGKGKRYALLMCGEDSEYLL 45
E +M + GGGGGGG KRY E E +L
Sbjct: 57 EEDMSNPSSSGGGGGGGMKKRYRTKFTPEQKEKML 161
>TC13588 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall structural
protein 1.8 precursor (GRP 1.8), partial (6%)
Length = 517
Score = 31.6 bits (70), Expect = 0.15
Identities = 13/18 (72%), Positives = 13/18 (72%)
Frame = +1
Query: 10 RELEMESGCGGGGGGGGG 27
RE S CGGGGGGGGG
Sbjct: 361 REWRYRS*CGGGGGGGGG 414
>AV420351
Length = 188
Score = 31.2 bits (69), Expect = 0.19
Identities = 11/11 (100%), Positives = 11/11 (100%)
Frame = -2
Query: 17 GCGGGGGGGGG 27
GCGGGGGGGGG
Sbjct: 94 GCGGGGGGGGG 62
Score = 30.8 bits (68), Expect = 0.25
Identities = 14/22 (63%), Positives = 14/22 (63%)
Frame = -2
Query: 8 LERELEMESGCGGGGGGGGGKG 29
L E E GCGG GGGGGG G
Sbjct: 130 LSGEEGAEYGCGGCGGGGGGGG 65
>AV426581
Length = 429
Score = 31.2 bits (69), Expect = 0.19
Identities = 23/46 (50%), Positives = 26/46 (56%), Gaps = 5/46 (10%)
Frame = -1
Query: 2 KGRTMLLERELEMESGCG-----GGGGGGGGKGKRYALLMCGEDSE 42
K R M LE E+E+E G GGGGGGGG G AL ED+E
Sbjct: 231 KKRQMDLEVEVEVEEKEGFEFKNGGGGGGGGGGPD*AL---QEDAE 103
>TC10717 similar to UP|Q8S564 (Q8S564) 4-coumarate:coenzyme A ligase ,
partial (32%)
Length = 691
Score = 31.2 bits (69), Expect = 0.19
Identities = 16/35 (45%), Positives = 19/35 (53%), Gaps = 5/35 (14%)
Frame = -1
Query: 3 GRTMLLERELEMESGCGGGGGGG-----GGKGKRY 32
G + LER++ S CGGGGGG GG G Y
Sbjct: 154 GMSGSLERKMNCSSSCGGGGGGDMPENEGGGGCNY 50
>TC12355 similar to UP|PR18_HUMAN (Q99633) Pre-mRNA splicing factor 18
(PRP18 homolog) (hPRP18), partial (8%)
Length = 563
Score = 31.2 bits (69), Expect = 0.19
Identities = 22/57 (38%), Positives = 27/57 (46%), Gaps = 13/57 (22%)
Frame = -1
Query: 17 GCGGGGGGGGGKGK--------RYALLMCGEDSEYLLKKHGG-----CFGFFTRMLA 60
GCGGG GGGGG G ++ALL+ E + L G GFF + LA
Sbjct: 269 GCGGGVGGGGGGGGGAF*RFGFQFALLLFAEFLDLFLLDLGALEEFLAAGFFGKGLA 99
>TC18264 similar to UP|Q9SP93 (Q9SP93) Thiol protease, partial (4%)
Length = 570
Score = 30.8 bits (68), Expect = 0.25
Identities = 12/15 (80%), Positives = 13/15 (86%)
Frame = +2
Query: 17 GCGGGGGGGGGKGKR 31
G GGGGGGGGG G+R
Sbjct: 473 GDGGGGGGGGGGGRR 517
Score = 26.9 bits (58), Expect = 3.6
Identities = 12/17 (70%), Positives = 12/17 (70%)
Frame = +2
Query: 15 ESGCGGGGGGGGGKGKR 31
E G GGGGGGGG G R
Sbjct: 464 E*GGDGGGGGGGGGGGR 514
Score = 26.6 bits (57), Expect = 4.7
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +2
Query: 15 ESGCGGGGGGGGGKGKR 31
+ G GGGGGGGG + K+
Sbjct: 476 DGGGGGGGGGGGRRVKK 526
>TC14729 homologue to UP|Q8W590 (Q8W590) At2g32080/F22D22.17, partial (51%)
Length = 749
Score = 30.8 bits (68), Expect = 0.25
Identities = 13/19 (68%), Positives = 14/19 (73%)
Frame = +3
Query: 11 ELEMESGCGGGGGGGGGKG 29
E+E SG GGGGG GGG G
Sbjct: 237 EMEGNSGGGGGGGSGGGGG 293
Score = 30.8 bits (68), Expect = 0.25
Identities = 14/25 (56%), Positives = 14/25 (56%)
Frame = +3
Query: 5 TMLLERELEMESGCGGGGGGGGGKG 29
T L EME GGGGGGG G G
Sbjct: 213 TSLSSGAAEMEGNSGGGGGGGSGGG 287
>TC10777 similar to UP|Q9SCL3 (Q9SCL3) Pre-mRNA splicing factor SF2-like
protein, partial (46%)
Length = 597
Score = 30.8 bits (68), Expect = 0.25
Identities = 13/18 (72%), Positives = 13/18 (72%)
Frame = +1
Query: 17 GCGGGGGGGGGKGKRYAL 34
G GGGGGGGGG G R L
Sbjct: 514 GYGGGGGGGGGGGGRIGL 567
>AW719706
Length = 432
Score = 30.8 bits (68), Expect = 0.25
Identities = 11/13 (84%), Positives = 11/13 (84%)
Frame = -3
Query: 17 GCGGGGGGGGGKG 29
GCGGGG GGGG G
Sbjct: 169 GCGGGGSGGGGDG 131
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.143 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,568,450
Number of Sequences: 28460
Number of extensions: 90366
Number of successful extensions: 2679
Number of sequences better than 10.0: 187
Number of HSP's better than 10.0 without gapping: 1182
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1952
length of query: 267
length of database: 4,897,600
effective HSP length: 89
effective length of query: 178
effective length of database: 2,364,660
effective search space: 420909480
effective search space used: 420909480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0055.14


