

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.5
(638 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
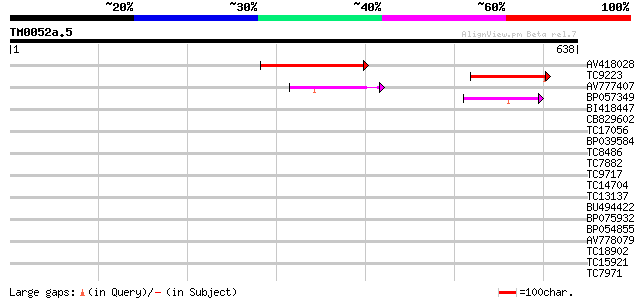
Sequences producing significant alignments: (bits) Value
AV418028 179 1e-45
TC9223 similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol 3-O-gluc... 115 2e-26
AV777407 92 3e-19
BP057349 55 3e-08
BI418447 34 0.063
CB829602 33 0.11
TC17056 similar to UP|Q94BZ2 (Q94BZ2) At1g30970/F17F8_14, partia... 32 0.31
BP039584 31 0.53
TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Ara... 30 0.91
TC7882 homologue to UP|O49875 (O49875) Adenine nucleotide transl... 30 1.2
TC9717 weakly similar to UP|Q9LNI1 (Q9LNI1) F6F3.22 protein, par... 29 2.6
TC14704 28 3.5
TC13137 weakly similar to UP|Q8LKG3 (Q8LKG3) UDP-glucosyltransfe... 28 5.9
BU494422 28 5.9
BP075932 28 5.9
BP054855 27 7.7
AV778079 27 7.7
TC18902 weakly similar to GB|AAP37786.1|30725528|BT008427 At1g51... 27 7.7
TC15921 weakly similar to GB|AAB64701.1|603415|SCE9163 Yer174cp ... 27 7.7
TC7971 homologue to PIR|C86216|C86216 protein T23G18.6 [imported... 27 7.7
>AV418028
Length = 366
Score = 179 bits (454), Expect = 1e-45
Identities = 79/121 (65%), Positives = 97/121 (79%)
Frame = +2
Query: 283 LETGVPFRAQAIIANPPAYGHAHVAEALGVPLHIFFTMPWTPTYDFPHPLARVPQSAGYW 342
L++GV F+A AIIANPPAYGH HVAEAL +P+HIFFTMPWTPT +FPHPL+RV Q AGY
Sbjct: 2 LDSGVDFKADAIIANPPAYGHTHVAEALKIPIHIFFTMPWTPTAEFPHPLSRVKQPAGYR 181
Query: 343 MSYIIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRGSISHLPTGYMWSPHVVPKPSD 402
+SY IVD + W G+R +IND RKK+LKL P+ Y S +GS + +P Y+WSPH+VPKP D
Sbjct: 182 LSYQIVDSLIWLGIRDMINDLRKKRLKLRPVTYLSGSQGSDTDIPHAYIWSPHLVPKPKD 361
Query: 403 W 403
W
Sbjct: 362 W 364
>TC9223 similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol
3-O-glucosyltransferase, partial (16%)
Length = 659
Score = 115 bits (288), Expect = 2e-26
Identities = 54/98 (55%), Positives = 69/98 (70%), Gaps = 8/98 (8%)
Frame = +1
Query: 519 KAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEVKSRVML 578
KA CPTTIVPFFGDQ FWGDR+H++ +GP PIP+ + ++ L NAI FML P+VK R +
Sbjct: 1 KAACPTTIVPFFGDQPFWGDRVHDRGVGPPPIPVDEFSLPKLVNAINFMLDPKVKERAIE 180
Query: 579 IAKLVENEDGVAAAVDAFHRHL--------PDELPLPT 608
+AK +ENEDGV AV AF + L PD+ PLP+
Sbjct: 181 LAKAMENEDGVTGAVKAFFKQLPQTRNKTEPDQQPLPS 294
>AV777407
Length = 520
Score = 92.0 bits (227), Expect = 3e-19
Identities = 52/136 (38%), Positives = 68/136 (49%), Gaps = 30/136 (22%)
Frame = -2
Query: 316 IFFTMPWTPTYDFPHPLARVPQSAGY------------------------------WMSY 345
+F P PT +FPHPL+RV Q AGY +SY
Sbjct: 519 VFHIFPPRPTAEFPHPLSRVKQPAGYRVSSFI*PGSDLALNFVAFMCD*ILIIYNSQLSY 340
Query: 346 IIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRGSISHLPTGYMWSPHVVPKPSDWGP 405
IVD + W G+R +IND RKK+LKL P+ Y S +GS + +P Y+WSPH+VPKP
Sbjct: 339 QIVDSLIWLGIRDMINDLRKKRLKLRPVTYLSGSQGSDTDIPHAYIWSPHLVPKPKG--- 169
Query: 406 LVDVVGYCFLSLASKY 421
CF SL+S +
Sbjct: 168 -------CFRSLSSHF 142
>BP057349
Length = 493
Score = 55.1 bits (131), Expect = 3e-08
Identities = 33/102 (32%), Positives = 50/102 (48%), Gaps = 12/102 (11%)
Frame = -2
Query: 511 AGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVEN----------- 559
+GTTA L+AG P + PF DQF+W +R++ + P P+ L +
Sbjct: 453 SGTTAAALQAGTPQVVCPFMLDQFYWAERMYWLGISPEPLSRHHLLPDKNDDRSVQEAAH 274
Query: 560 -LSNAIKFMLQPEVKSRVMLIAKLVENEDGVAAAVDAFHRHL 600
LS AI L VK++ + IA+ + EDGV+ A+ L
Sbjct: 273 VLSLAIHDALSSRVKTQAIEIAERLSLEDGVSEAIKCLKEEL 148
>BI418447
Length = 443
Score = 34.3 bits (77), Expect = 0.063
Identities = 25/104 (24%), Positives = 41/104 (39%), Gaps = 1/104 (0%)
Frame = +1
Query: 513 TTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEV 572
TTAT G G Q + + + + L + + NA+KF+ P+V
Sbjct: 118 TTATDENQGAEIVQPTNLGQQNATEEPVKQSSTTSVFVNTEPLREDQIQNAVKFLSHPKV 297
Query: 573 K-SRVMLIAKLVENEDGVAAAVDAFHRHLPDELPLPTPSPQEED 615
K S V+ +E + +D R +PD+ P S +D
Sbjct: 298 KGSPVIYRRSFLEKKGLTKEEIDEAFRRVPDDAPTVQTSGVNQD 429
>CB829602
Length = 542
Score = 33.5 bits (75), Expect = 0.11
Identities = 28/126 (22%), Positives = 51/126 (40%), Gaps = 12/126 (9%)
Frame = +1
Query: 421 YKPREDFVQWIKKGPPP--LYFGFGSMPLEDPTRTTDVILEALKDTEQ-----RGIIDRG 473
+K + ++W+ P LY FGS+ P + ++ ++ R + G
Sbjct: 76 WKEEPECLKWLDSQEPSSVLYVNFGSVINMTPQQLVELAWGIANSKKKFIWVIRPDLVEG 255
Query: 474 WGNL---GSLAEVPDNVFLLEECPHDWLFPQCS--AVVHHGGAGTTATGLKAGCPTTIVP 528
++ +AE D +L CP + + + + H G +T + +G P P
Sbjct: 256 EASIVLPEIVAETKDRGIMLSWCPQEQVLKHSALGGFLTHCGWNSTIESISSGVPLICSP 435
Query: 529 FFGDQF 534
FF DQF
Sbjct: 436 FFNDQF 453
>TC17056 similar to UP|Q94BZ2 (Q94BZ2) At1g30970/F17F8_14, partial (40%)
Length = 736
Score = 32.0 bits (71), Expect = 0.31
Identities = 30/108 (27%), Positives = 44/108 (39%), Gaps = 1/108 (0%)
Frame = +2
Query: 226 KSAGIDFYPLGG-DPRILAGYMARNKGLIPSGPGEVSIQLKQLKDIIDSLLPACTAPDLE 284
+S I+ Y + G P LA + + +PS +V + QL + ++P P L
Sbjct: 317 ESTDIEIYGMQGIPPDELAAHYGEEEDDVPSKAAKVDVPSAQL---VGGMVP----PPLG 475
Query: 285 TGVPFRAQAIIANPPAYGHAHVAEALGVPLHIFFTMPWTPTYDFPHPL 332
G P R+ + PP Y HA A VP P P P P+
Sbjct: 476 PGYPPRS-TLGPMPPMYNHAVPPSAWPVPPRPQPWFPQPPAVSMPPPV 616
>BP039584
Length = 545
Score = 31.2 bits (69), Expect = 0.53
Identities = 14/36 (38%), Positives = 18/36 (49%)
Frame = -3
Query: 500 PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFF 535
P VV H G TT + AG P +P F +QF+
Sbjct: 507 PAIGGVVTHCGWNTTLESVIAGLPMATMPLFAEQFY 400
>TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;}, partial (25%)
Length = 856
Score = 30.4 bits (67), Expect = 0.91
Identities = 23/90 (25%), Positives = 42/90 (46%), Gaps = 8/90 (8%)
Frame = +1
Query: 203 AKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDPRILA--------GYMARNKGLIP 254
AK L YG +++ TH N ++V GGD +ILA ++++ L
Sbjct: 154 AKFLCSYGGKIQPRTHDNQLSYV----------GGDTKILAVDRNTKFPTFLSKLAALCD 303
Query: 255 SGPGEVSIQLKQLKDIIDSLLPACTAPDLE 284
+ P E++ + + + +D+L+ DLE
Sbjct: 304 AAPQELTFKYQLPGEDLDALISVTNDDDLE 393
>TC7882 homologue to UP|O49875 (O49875) Adenine nucleotide translocator,
complete
Length = 1656
Score = 30.0 bits (66), Expect = 1.2
Identities = 20/68 (29%), Positives = 27/68 (39%)
Frame = +1
Query: 498 LFPQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNV 557
+ P C A V A TA+ + G P FF D G + API +L +
Sbjct: 361 VMPSCRASVDLSAAAATASPVFVGAPAEKSHFFIDFLMGGVSAAVSKTAAAPIERVKLLI 540
Query: 558 ENLSNAIK 565
+N IK
Sbjct: 541 QNQDEMIK 564
>TC9717 weakly similar to UP|Q9LNI1 (Q9LNI1) F6F3.22 protein, partial (18%)
Length = 902
Score = 28.9 bits (63), Expect = 2.6
Identities = 13/39 (33%), Positives = 20/39 (50%)
Frame = +2
Query: 508 HGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELG 546
H G + GL+ GCP ++PF +Q + K+LG
Sbjct: 248 HCGWSSVIEGLQVGCPLIMLPFPNEQMLVARFMEGKKLG 364
>TC14704
Length = 587
Score = 28.5 bits (62), Expect = 3.5
Identities = 14/28 (50%), Positives = 20/28 (71%)
Frame = +3
Query: 147 MERSASVPSELLELQSFGEPTVSGSLSS 174
+ERSAS+ S+ +E + E + SGSLSS
Sbjct: 48 LERSASISSDNIESELVTEASASGSLSS 131
>TC13137 weakly similar to UP|Q8LKG3 (Q8LKG3) UDP-glucosyltransferase,
partial (7%)
Length = 503
Score = 27.7 bits (60), Expect = 5.9
Identities = 11/37 (29%), Positives = 19/37 (50%)
Frame = +3
Query: 510 GAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELG 546
G + L+ GCP ++PF +Q + EK++G
Sbjct: 6 GWSSVIEALQVGCPLIMLPFHNEQVLVAKLMEEKKVG 116
>BU494422
Length = 564
Score = 27.7 bits (60), Expect = 5.9
Identities = 18/58 (31%), Positives = 27/58 (46%)
Frame = -1
Query: 471 DRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVP 528
D G+ L+E + L P WL C+A+ HHG G+T + C +T+ P
Sbjct: 294 DNGYEKRSLLSEA-STLLLQPSFPSKWL-AFCAALQHHGPFGSTCS---CSCHSTLSP 136
>BP075932
Length = 560
Score = 27.7 bits (60), Expect = 5.9
Identities = 16/52 (30%), Positives = 29/52 (55%), Gaps = 6/52 (11%)
Frame = +2
Query: 570 PEVKSRVMLIAKLVENEDGVAAAV----DAFHRHLPDELP--LPTPSPQEED 615
P + S + L+ +L+ ++ + + + A H HL ++LP LP P P +ED
Sbjct: 203 PALFSSMKLMVELITSKQIIPSRILRSLGASHHHLQEQLP*WLPFP*PMKED 358
>BP054855
Length = 516
Score = 27.3 bits (59), Expect = 7.7
Identities = 12/31 (38%), Positives = 14/31 (44%)
Frame = -3
Query: 604 LPLPTPSPQEEDQINPLQWFFLQIGKWCCAP 634
LPLP P+ N L G+WC AP
Sbjct: 367 LPLPQMLPRSCASTN*YSCILLSFGRWCTAP 275
>AV778079
Length = 601
Score = 27.3 bits (59), Expect = 7.7
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +2
Query: 599 HLPDELPLPTPSPQEEDQINP 619
H P E+P+P PSP+E+ P
Sbjct: 242 HNPMEVPVPIPSPREDQGARP 304
>TC18902 weakly similar to GB|AAP37786.1|30725528|BT008427 At1g51560
{Arabidopsis thaliana;}, partial (11%)
Length = 575
Score = 27.3 bits (59), Expect = 7.7
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = -1
Query: 611 PQEEDQINPLQWFFLQIGKWC 631
P+E +QI WF +Q+G WC
Sbjct: 380 PEETNQIK-YNWFVVQVGLWC 321
>TC15921 weakly similar to GB|AAB64701.1|603415|SCE9163 Yer174cp
{Saccharomyces cerevisiae;} , partial (10%)
Length = 580
Score = 27.3 bits (59), Expect = 7.7
Identities = 13/32 (40%), Positives = 15/32 (46%)
Frame = -1
Query: 313 PLHIFFTMPWTPTYDFPHPLARVPQSAGYWMS 344
PLH+F T W Y P P+ S G W S
Sbjct: 388 PLHLFHT--WRSIYHNPQPVHHPQISGGMWAS 299
>TC7971 homologue to PIR|C86216|C86216 protein T23G18.6 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(98%)
Length = 1662
Score = 27.3 bits (59), Expect = 7.7
Identities = 12/23 (52%), Positives = 14/23 (60%)
Frame = -1
Query: 610 SPQEEDQINPLQWFFLQIGKWCC 632
SPQEE Q N +W+ G WCC
Sbjct: 414 SPQEEHQ*NYCKWY---QGVWCC 355
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,581,968
Number of Sequences: 28460
Number of extensions: 201414
Number of successful extensions: 950
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 942
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 949
length of query: 638
length of database: 4,897,600
effective HSP length: 96
effective length of query: 542
effective length of database: 2,165,440
effective search space: 1173668480
effective search space used: 1173668480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0052a.5


