

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.2
(68 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
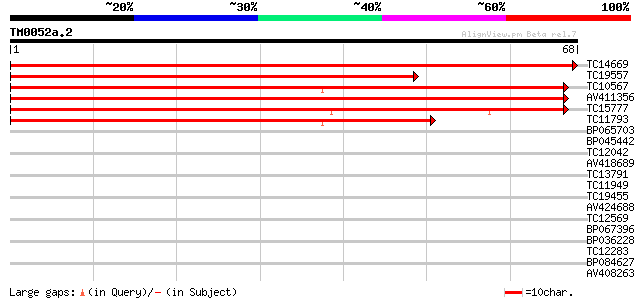
TC14669 similar to PIR|E86326|E86326 protein F18O14.3 [imported]... 123 6e-30
TC19557 similar to PIR|E86326|E86326 protein F18O14.3 [imported]... 96 2e-21
TC10567 similar to UP|AAR99376 (AAR99376) Ring domain containing... 90 7e-20
AV411356 89 2e-19
TC15777 similar to UP|AAR99376 (AAR99376) Ring domain containing... 88 4e-19
TC11793 similar to UP|AAR99376 (AAR99376) Ring domain containing... 72 3e-14
BP065703 34 0.006
BP045442 33 0.008
TC12042 similar to GB|CAB87983.1|7576235|ATH276134 Pex10p {Arabi... 33 0.008
AV418689 33 0.010
TC13791 weakly similar to UP|RO52_HUMAN (P19474) 52 kDa Ro prote... 33 0.010
TC11949 33 0.010
TC19455 similar to UP|Q8S8Z6 (Q8S8Z6) Syringolide-induced protei... 33 0.014
AV424688 33 0.014
TC12569 UP|Q40221 (Q40221) Protein containing C-terminal RING-fi... 33 0.014
BP067396 32 0.018
BP036228 32 0.023
TC12283 similar to UP|AAR23700 (AAR23700) At1g65040, partial (21%) 32 0.023
BP084627 32 0.030
AV408263 32 0.030
>TC14669 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(77%)
Length = 1120
Score = 123 bits (309), Expect = 6e-30
Identities = 53/68 (77%), Positives = 59/68 (85%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
CNICFDLAQDP+ITLCG+LF WPCLYKWLHFHS S++CPVC AL EE+KLVPL+GRG TS
Sbjct: 128 CNICFDLAQDPIITLCGHLFCWPCLYKWLHFHSQSRECPVCKALVEEEKLVPLYGRGKTS 307
Query: 61 SSSSSSSI 68
S S SI
Sbjct: 308 SDPRSRSI 331
>TC19557 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(24%)
Length = 457
Score = 95.5 bits (236), Expect = 2e-21
Identities = 38/49 (77%), Positives = 44/49 (89%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDK 49
CNICF+LAQDPVITLCG+LF WPCLY+WLH HS SQ+CPVC AL +E+K
Sbjct: 311 CNICFELAQDPVITLCGHLFCWPCLYRWLHHHSQSQECPVCKALIQEEK 457
>TC10567 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (27%)
Length = 423
Score = 90.1 bits (222), Expect = 7e-20
Identities = 39/75 (52%), Positives = 50/75 (66%), Gaps = 8/75 (10%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQ--------QCPVCNALAEEDKLVP 52
C+IC D QDPV+TLCG+L+ WPC+YKWL+FHS+S QCPVC + E LVP
Sbjct: 86 CSICLDRVQDPVVTLCGHLYCWPCIYKWLNFHSVSSDHEEKKEPQCPVCKSEISESSLVP 265
Query: 53 LHGRGNTSSSSSSSS 67
L+GR T+ S S +
Sbjct: 266 LYGRDQTALPSKSKA 310
>AV411356
Length = 358
Score = 88.6 bits (218), Expect = 2e-19
Identities = 34/67 (50%), Positives = 46/67 (67%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
CNIC DLA++PV+T CG+LF W C+Y+WLH HS +++CPVC + P++GRGN
Sbjct: 44 CNICLDLAREPVVTCCGHLFCWTCVYRWLHLHSDAKECPVCKGEVTLKSVTPIYGRGNNG 223
Query: 61 SSSSSSS 67
SS S
Sbjct: 224 RSSEEDS 244
>TC15777 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (34%)
Length = 566
Score = 87.8 bits (216), Expect = 4e-19
Identities = 39/76 (51%), Positives = 51/76 (66%), Gaps = 9/76 (11%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQ--------CPVCNALAEEDKLVP 52
CNIC D QDPV+TLCG+L+ WPC+YKWL+F+S+S CPVC + + LVP
Sbjct: 289 CNICLDCVQDPVVTLCGHLYCWPCIYKWLNFYSISSDYEEKEEPICPVCKSEISQSSLVP 468
Query: 53 LHGR-GNTSSSSSSSS 67
L+GR G T+S S S +
Sbjct: 469 LYGREGQTTSPSKSKA 516
>TC11793 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (23%)
Length = 527
Score = 71.6 bits (174), Expect = 3e-14
Identities = 30/59 (50%), Positives = 38/59 (63%), Gaps = 8/59 (13%)
Frame = +3
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQ--------QCPVCNALAEEDKLV 51
CNIC D QDPV+TLCG+L+ WPC+YKWL+ +S S QCPVC + + LV
Sbjct: 351 CNICLDCVQDPVVTLCGHLYCWPCIYKWLNSNSASSENVEQNKPQCPVCKSEVSQSSLV 527
>BP065703
Length = 480
Score = 33.9 bits (76), Expect = 0.006
Identities = 15/43 (34%), Positives = 23/43 (52%)
Frame = -3
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNA 43
C +C D ++ VI C +LF PC+ + L ++CP C A
Sbjct: 319 CTVCSDRPKEVVIVKCYHLFCNPCIQRNLELR--HRKCPACGA 197
>BP045442
Length = 488
Score = 33.5 bits (75), Expect = 0.008
Identities = 18/52 (34%), Positives = 28/52 (53%), Gaps = 4/52 (7%)
Frame = -1
Query: 1 CNIC---FDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNA-LAEED 48
C++C F+ + CG+LF CL KWL + +++ CP+C L ED
Sbjct: 437 CSVCLTKFEPESEINCLPCGHLFHKACLEKWLDYWNIT--CPLCRTPLMPED 288
>TC12042 similar to GB|CAB87983.1|7576235|ATH276134 Pex10p {Arabidopsis
thaliana;} , partial (19%)
Length = 582
Score = 33.5 bits (75), Expect = 0.008
Identities = 12/39 (30%), Positives = 18/39 (45%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCP 39
C +C P T CG++F W C+ +W + S P
Sbjct: 148 CTLCLSNRPHPTATSCGHVFCWNCITEWCNEKSRVPSLP 264
>AV418689
Length = 371
Score = 33.1 bits (74), Expect = 0.010
Identities = 17/54 (31%), Positives = 25/54 (45%)
Frame = +3
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLH 54
C I DL +DPV G + + KW F + CPV N + + ++P H
Sbjct: 207 CPISLDLMKDPVTLSTGITYDRDSVEKW--FDDGNYTCPVTNQIVKNFDMIPNH 362
>TC13791 weakly similar to UP|RO52_HUMAN (P19474) 52 kDa Ro protein (Sjogren
syndrome type A antigen) (SS-A) (Ro(SS-A)) (52 kDa
ribonucleoprotein autoantigen Ro/SS-A), partial (5%)
Length = 545
Score = 33.1 bits (74), Expect = 0.010
Identities = 19/56 (33%), Positives = 26/56 (45%), Gaps = 10/56 (17%)
Frame = +3
Query: 1 CNICFDLAQDPVITLCGYLFRWPC--------LYKWLHFHSLSQQCPVC--NALAE 46
C IC D DPV CG++F + C + L Q+CP+C NA+ E
Sbjct: 60 CPICLDTVFDPVSLTCGHIFCYMCACSAASVSIVDGLKSAVTKQKCPLCRENAVYE 227
>TC11949
Length = 780
Score = 33.1 bits (74), Expect = 0.010
Identities = 15/49 (30%), Positives = 23/49 (46%), Gaps = 8/49 (16%)
Frame = +3
Query: 1 CNICFDLAQDPVITLCGYLFRWPC--------LYKWLHFHSLSQQCPVC 41
C+IC D DPV CG++F + C + L ++CP+C
Sbjct: 129 CSICMDTVFDPVSLTCGHIFCYSCACSAASVTIVDGLKAAXPKEKCPLC 275
>TC19455 similar to UP|Q8S8Z6 (Q8S8Z6) Syringolide-induced protein 13-1-1,
partial (25%)
Length = 485
Score = 32.7 bits (73), Expect = 0.014
Identities = 16/54 (29%), Positives = 27/54 (49%)
Frame = +1
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLH 54
C + DL +DPV G + + KW+ + +++CPV N + L+P H
Sbjct: 178 CPVSLDLMKDPVTLPTGITYDRTSIEKWI--EAGNKKCPVTNQILTTFDLIPNH 333
>AV424688
Length = 414
Score = 32.7 bits (73), Expect = 0.014
Identities = 12/29 (41%), Positives = 21/29 (72%)
Frame = +3
Query: 32 HSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
+S +++CPVCN E LVP++G+ ++S
Sbjct: 6 YSNARECPVCNGEVTEADLVPIYGKSSSS 92
>TC12569 UP|Q40221 (Q40221) Protein containing C-terminal RING-finger,
complete
Length = 2092
Score = 32.7 bits (73), Expect = 0.014
Identities = 21/55 (38%), Positives = 31/55 (56%), Gaps = 5/55 (9%)
Frame = +2
Query: 1 CNICFDLAQ--DPVITL--CGYLFRWPCLYKWLHFHSLSQQCPVCNA-LAEEDKL 50
C IC + + D V TL CG+ + C+ KWL S+ + CP+C A + EDK+
Sbjct: 1784 CVICLEEYKNMDDVGTLKTCGHDYHVNCIKKWL---SMKKLCPICKASVMPEDKM 1939
>BP067396
Length = 506
Score = 32.3 bits (72), Expect = 0.018
Identities = 15/45 (33%), Positives = 24/45 (53%), Gaps = 4/45 (8%)
Frame = -1
Query: 1 CNICF-DLAQDPVITL---CGYLFRWPCLYKWLHFHSLSQQCPVC 41
C+IC D + + + CG+++ C+ WL FHS CP+C
Sbjct: 386 CSICLGDYKEADTLRMLPDCGHVYHLACVDPWLKFHS---TCPIC 261
>BP036228
Length = 626
Score = 32.0 bits (71), Expect = 0.023
Identities = 12/29 (41%), Positives = 15/29 (51%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWL 29
C+ C L PV T CG+ F C KW+
Sbjct: 422 CSFCMQLPDRPVTTPCGHNFCLKCFEKWI 508
>TC12283 similar to UP|AAR23700 (AAR23700) At1g65040, partial (21%)
Length = 390
Score = 32.0 bits (71), Expect = 0.023
Identities = 16/44 (36%), Positives = 19/44 (42%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNAL 44
C IC + +CG+LF CL WL CP C AL
Sbjct: 137 CIICREEMTTAKKLICGHLFHVHCLRSWL---ERQHTCPTCRAL 259
>BP084627
Length = 327
Score = 31.6 bits (70), Expect = 0.030
Identities = 17/54 (31%), Positives = 28/54 (51%)
Frame = -2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLH 54
C+IC D ++ VIT C +LF C+ K S ++CP C + + P++
Sbjct: 326 CSICQDRTKEVVITKCYHLFCNTCMLKVT--GSRHRKCPQCGTSFGANDVKPVY 171
>AV408263
Length = 331
Score = 31.6 bits (70), Expect = 0.030
Identities = 13/29 (44%), Positives = 16/29 (54%)
Frame = +2
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWL 29
C I +L QDPVI G + C+ KWL
Sbjct: 116 CPISLELMQDPVIVSTGQTYERSCIEKWL 202
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.137 0.473
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,185,768
Number of Sequences: 28460
Number of extensions: 41307
Number of successful extensions: 457
Number of sequences better than 10.0: 100
Number of HSP's better than 10.0 without gapping: 445
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 453
length of query: 68
length of database: 4,897,600
effective HSP length: 44
effective length of query: 24
effective length of database: 3,645,360
effective search space: 87488640
effective search space used: 87488640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0052a.2


