

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.1
(387 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
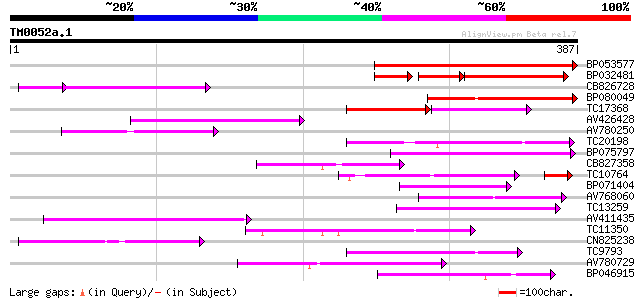
BP053577 212 1e-55
BP032481 97 2e-38
CB826728 98 2e-25
BP080049 92 1e-19
TC17368 59 1e-18
AV426428 83 9e-17
AV780250 80 6e-16
TC20198 similar to UP|Q9ZPU0 (Q9ZPU0) At2g13980 protein, partial... 78 2e-15
BP075797 70 5e-13
CB827358 70 6e-13
TC10764 65 3e-12
BP071404 66 9e-12
AV768060 66 9e-12
TC13259 66 1e-11
AV411435 64 3e-11
TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3,... 64 4e-11
CN825238 60 5e-10
TC9793 60 5e-10
AV780729 55 2e-08
BP046915 51 3e-07
>BP053577
Length = 463
Score = 212 bits (539), Expect = 1e-55
Identities = 104/138 (75%), Positives = 117/138 (84%)
Frame = +3
Query: 250 GGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDC 309
GG AGLGMVARN +GE MA+ACS PV ISSPLLAEA+ +RWTMQLA DLGF I LETDC
Sbjct: 3 GGIAGLGMVARNCDGEVMASACSSPVSISSPLLAEALGLRWTMQLATDLGFRRITLETDC 182
Query: 310 LNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARNANTYANM 369
L LFN WR+ EE RSYLS +I DCR+LISAF++VS+ FVRRTGN VADFLARN++TYANM
Sbjct: 183 LQLFNTWRSQEEGRSYLSSIISDCRILISAFDYVSVSFVRRTGNTVADFLARNSDTYANM 362
Query: 370 VWVEEVPLAVLPLFDADV 387
VWVEEVPL V+ L DADV
Sbjct: 363 VWVEEVPLTVVSLIDADV 416
>BP032481
Length = 447
Score = 97.1 bits (240), Expect(3) = 2e-38
Identities = 45/71 (63%), Positives = 55/71 (77%)
Frame = -1
Query: 311 NLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARNANTYANMV 370
NLF +W+ E + SYLS +++DCRLL +AF+ VSL FVRRTGN VA F ARNA TYAN+V
Sbjct: 261 NLFKSWKAKEVE*SYLSSILQDCRLLFTAFDVVSLSFVRRTGNTVAGFFARNAETYANLV 82
Query: 371 WVEEVPLAVLP 381
W EEVP+A P
Sbjct: 81 WPEEVPVAAFP 49
Score = 54.7 bits (130), Expect(3) = 2e-38
Identities = 26/31 (83%), Positives = 27/31 (86%)
Frame = -2
Query: 280 PLLAEAMAMRWTMQLAIDLGFH*IVLETDCL 310
PLLAEAM+MRW MQLAIDLGF I LETDCL
Sbjct: 353 PLLAEAMSMRWIMQLAIDLGFRRICLETDCL 261
Score = 44.7 bits (104), Expect(3) = 2e-38
Identities = 18/26 (69%), Positives = 21/26 (80%)
Frame = -3
Query: 250 GGTAGLGMVARNSEGEAMAAACSFPV 275
GG G+GMVARNSEG+ MA CSFP+
Sbjct: 442 GGLGGMGMVARNSEGDVMATTCSFPI 365
>CB826728
Length = 509
Score = 98.2 bits (243), Expect(2) = 2e-25
Identities = 46/101 (45%), Positives = 60/101 (58%)
Frame = +1
Query: 37 ILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCW 96
+ FWP SS+G Y+SK+GY F+R PS+S A WK W+ S+LPRCKE W
Sbjct: 205 VFFWPGSSDGWYSSKSGYEFIRLENKSLLPSTSPAAAVPSLIWKTVWSASSLPRCKEVMW 384
Query: 97 RAISGYLPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLC 137
RA +GYLP+R L RR +DVDP E+E H+ + C
Sbjct: 385 RACAGYLPVRSALKRRGLDVDPSSPWSGLEDEAEVHVLLTC 507
Score = 34.3 bits (77), Expect(2) = 2e-25
Identities = 14/33 (42%), Positives = 19/33 (57%)
Frame = +3
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILF 39
W +L+ VFSPST + L++ L L D LF
Sbjct: 114 WNENLVSMVFSPSTTAVFLAVSLPLQSYEDCLF 212
>BP080049
Length = 356
Score = 92.4 bits (228), Expect = 1e-19
Identities = 49/102 (48%), Positives = 64/102 (62%)
Frame = -1
Query: 286 MAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSL 345
+++RW + LA +LGF *IV ETDCL+L+ AW+ S L LI DCR L+S F+ L
Sbjct: 356 LSLRWALGLASELGFR*IVFETDCLHLYEAWKR-RGGCSVLDTLILDCRNLLSYFDVFQL 180
Query: 346 CFVRRTGNAVADFLARNANTYANMVWVEEVPLAVLPLFDADV 387
FVRRT N V D LA+ N+VWVEE+P + L +DV
Sbjct: 179 SFVRRTSNCVVDALAKLCFQLGNVVWVEELPHELYGLVQSDV 54
>TC17368
Length = 563
Score = 58.5 bits (140), Expect(2) = 1e-18
Identities = 26/57 (45%), Positives = 38/57 (66%)
Frame = +1
Query: 231 WRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMA 287
W+ P G+I NF ASF+ G AG G++ARN G+ +A A + V +SSP++A+A A
Sbjct: 55 WKPPAQGSINVNFGASFKAGVGAGFGLIARNGSGQVLAIATNLIVDVSSPVVAQAYA 225
Score = 50.8 bits (120), Expect(2) = 1e-18
Identities = 26/68 (38%), Positives = 37/68 (54%)
Frame = +3
Query: 289 RWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSLCFV 348
RW ++L +DLGF ++ E DC L AW + YLS +++D L F S +V
Sbjct: 231 RWAIELTMDLGFSGVIFENDC*VLHCAWISKLNSPPYLSFIVQDYWFLSFGFNSCSFVYV 410
Query: 349 RRTGNAVA 356
+R NAVA
Sbjct: 411 KRQCNAVA 434
>AV426428
Length = 413
Score = 82.8 bits (203), Expect = 9e-17
Identities = 43/119 (36%), Positives = 64/119 (53%)
Frame = +3
Query: 83 WATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPAAKQ 142
W ALPRC ET WRA G LP++ L R + DP C C +TA+H + CP
Sbjct: 57 WKAEALPRCCETGWRACLGVLPVKASLHARGLAEDPICPRCLAAPKTADHAVLFCP*VHP 236
Query: 143 V*YASSLSLRVDSFIAFADFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSV 201
+ +ASSL R++ +F A ++ D + TI+YAIW A+N++ F+ T ++
Sbjct: 237 IWFASSLGFRLNQECKMHEFVADFMQVAD*DTCGEFLTILYAIWTAQNELCFRYTTVTL 413
>AV780250
Length = 494
Score = 80.1 bits (196), Expect = 6e-16
Identities = 43/107 (40%), Positives = 54/107 (50%)
Frame = +3
Query: 36 DILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETC 95
D+L WP +SNG YTSK+GY +R PSSSTA+ SP WK LPRC+E
Sbjct: 174 DVLAWPHTSNGEYTSKSGYAVVRNKVVHALPSSSTALQVSPVVWK----APTLPRCRELI 341
Query: 96 WRAISGYLPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPAAKQ 142
RA LP R L R +DVD C C E+ + + P +
Sbjct: 342 GRAFLKILPTRSALRARGMDVDELCPSCETHRESPAQVLLFVPVVSK 482
>TC20198 similar to UP|Q9ZPU0 (Q9ZPU0) At2g13980 protein, partial (13%)
Length = 630
Score = 78.2 bits (191), Expect = 2e-15
Identities = 51/157 (32%), Positives = 77/157 (48%), Gaps = 2/157 (1%)
Frame = +3
Query: 231 WRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRW 290
W P G K NFD S AGL +V R+ G + + I + + ++ R
Sbjct: 93 WPPPPDGVFKLNFDTSVLASS*AGLSLVVRDCAG*EVFS-------IQN*VFGASLISRD 251
Query: 291 --TMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSLCFV 348
+ + +DL ++L+TDC NLF W+ SY + R LL + F FV L FV
Sbjct: 252 G*SYSVLVDLFLTSVILKTDCYNLFVQWKKKHVSASYFGVYCRTFFLLCACFSFVDLSFV 431
Query: 349 RRTGNAVADFLARNANTYANMVWVEEVPLAVLPLFDA 385
RR GN VA+ LA+ A + VW+E+ P+ ++ L +
Sbjct: 432 RR-GNKVANDLAKLAFSSHGYVWIEDSPIGLVALLSS 539
>BP075797
Length = 504
Score = 70.5 bits (171), Expect = 5e-13
Identities = 46/126 (36%), Positives = 65/126 (51%)
Frame = -2
Query: 261 NSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIE 320
N EG+A+ A+ + P +AEA+++R T+ L D F I LE+DCL L +AW
Sbjct: 464 NHEGKAIIASSFHLLVAYEPAIAEALSLRRTITLPQDENFRIIELESDCLQLVHAWDRSV 285
Query: 321 EDRSYLSILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARNANTYANMVWVEEVPLAVL 380
YL+ +++DC+ L S + SL R N VAD LA+ A Y + WV P A
Sbjct: 284 PHPHYLASIVQDCKDLSSFSDSCSLKHSSRKINEVADHLAKLAFVYQDHFWVGIDPPATS 105
Query: 381 PLFDAD 386
L D
Sbjct: 104 LLLQPD 87
>CB827358
Length = 453
Score = 70.1 bits (170), Expect = 6e-13
Identities = 38/105 (36%), Positives = 57/105 (54%), Gaps = 4/105 (3%)
Frame = +2
Query: 169 QGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTGVLRRVAAMN----SASQKSDGGPQT 224
+ DS+ +++ Q A+WEA N++ FQ+ F ++R +M+ S S GP
Sbjct: 146 EADSDVVASCQMASCALWEAHNKLIFQEVPFRCEAAIQRATSMSQVDVSTVDPSANGPV- 322
Query: 225 GSRPARWRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAA 269
A WRRP G K NFD S++ +G+GMVARN +G +AA
Sbjct: 323 --HDAIWRRPPRGVFKVNFDGSWKSDSVSGIGMVARNHDGLILAA 451
Score = 35.4 bits (80), Expect = 0.016
Identities = 18/42 (42%), Positives = 23/42 (53%)
Frame = +3
Query: 85 TSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
T ALPRCK+ W + R L+R+ V+ DP C LC E
Sbjct: 3 TKALPRCKDLAW-----CVCCR*SLWRKGVNSDPSCPLCGNE 113
>TC10764
Length = 555
Score = 65.5 bits (158), Expect(2) = 3e-12
Identities = 42/126 (33%), Positives = 58/126 (45%), Gaps = 2/126 (1%)
Frame = +1
Query: 225 GSRPAR--WRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLL 282
GS+PA W P FD S G G++A + EGE +AA +FP+ + PL+
Sbjct: 34 GSKPATFGWENPN*------FDVSVVEHLDVGFGLLAGDHEGEVLAATSAFPLSVLPPLI 195
Query: 283 AEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEF 342
AEA W + + F E L L+ W SY+ L+ DCRLL F+F
Sbjct: 196 AEAFC--WALSSDMTSCFGSFCFEIGYLQLYETWMKT*PGSSYVCTLVNDCRLLACYFDF 369
Query: 343 VSLCFV 348
+ FV
Sbjct: 370 IEPSFV 387
Score = 22.3 bits (46), Expect(2) = 3e-12
Identities = 9/19 (47%), Positives = 12/19 (62%)
Frame = +3
Query: 366 YANMVWVEEVPLAVLPLFD 384
+ N+VW+EEV L L D
Sbjct: 447 FQNLVWIEEVRLHPLTRHD 503
>BP071404
Length = 394
Score = 66.2 bits (160), Expect = 9e-12
Identities = 33/76 (43%), Positives = 44/76 (57%)
Frame = -2
Query: 267 MAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYL 326
+ AA + P PL+AEA M+W+M+L LGF +VLETDCL L W +SY
Sbjct: 387 LVAASYYAGPSLDPLVAEAACMKWSMELCRMLGFQSLVLETDCLLLHQIWTARTLPKSYA 208
Query: 327 SILIRDCRLLISAFEF 342
++ DCR+L S F F
Sbjct: 207 GSIMEDCRMLASGFLF 160
>AV768060
Length = 416
Score = 66.2 bits (160), Expect = 9e-12
Identities = 36/101 (35%), Positives = 53/101 (51%)
Frame = -3
Query: 280 PLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISA 339
PL+AEAM RW + LA + G +V +DC L A+ E S I++ DC L+++
Sbjct: 300 PLIAEAMCFRWALNLAANWGLDSLVFSSDCQQLVKAFHCPSEFPSLAHIML-DCTQLVNS 124
Query: 340 FEFVSLCFVRRTGNAVADFLARNANTYANMVWVEEVPLAVL 380
F S CF R N VA LA ++ Y + W + P+ +L
Sbjct: 123 FRNFSFCFEYRQTNRVAHALAAESHLYKDCEWWGDPPVGIL 1
>TC13259
Length = 506
Score = 65.9 bits (159), Expect = 1e-11
Identities = 39/112 (34%), Positives = 63/112 (55%)
Frame = +1
Query: 265 EAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRS 324
+ +A A S S +EAMA+RW +QL+++LGF + +E D + N+W+ ++ S
Sbjct: 55 QVLATASSKWPRTVSVATSEAMALRWGLQLSLELGFFEVEVEIDNQVVVNSWKGTKKHVS 234
Query: 325 YLSILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARNANTYANMVWVEEVP 376
YL+ +I D + F F SL +V R + AD+LA+ A + V +EE P
Sbjct: 235 YLANVIADVVRISQCFRFFSLAYVPRLSHLAADYLAQFALSDYCFVGIEEHP 390
>AV411435
Length = 426
Score = 64.3 bits (155), Expect = 3e-11
Identities = 37/142 (26%), Positives = 62/142 (43%)
Frame = +3
Query: 24 ILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFW 83
I+ P S+ G D LFWP +G Y+ K+GY ++ + +S++ S W W
Sbjct: 3 IMQTPFSITGSEDRLFWPFIPSGEYSVKSGYRAVKATQFNLSNLASSSSHPSSFLWHTIW 182
Query: 84 ATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGENETAEHLFMLCPAAKQV 143
+ + WRA + +P++ L RR++ D C +C+ E H + C + V
Sbjct: 183 GAQVPKKVRSFLWRAANNAIPVKRNLKRRNMGRDDFCPICNKGPEDINHALLACEWTRAV 362
Query: 144 *YASSLSLRVDSFIAFADFWAW 165
+ L+ V F AW
Sbjct: 363 WFGCPLNFVVHEVHGML-FGAW 425
>TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3, P1 clone:
MIG10, partial (18%)
Length = 1118
Score = 63.9 bits (154), Expect = 4e-11
Identities = 52/179 (29%), Positives = 86/179 (47%), Gaps = 22/179 (12%)
Frame = -3
Query: 162 FWAWVVEQGD---------SEFISTVQTIIYAIWEARNQVQFQQRTFSVTGVLRRVAAMN 212
F W VE+ E ++T+ ++AIW+ RN+V FQ + + ++ M
Sbjct: 1065 FDRWFVERSQFILNHSLNHEEDLATLSFTLWAIWKNRNRVIFQLEEPNPVNTIFQIRHMQ 886
Query: 213 ------SASQKSDGGPQ------TGSRPARWRRPQPGTIKCNFDASFRGGG-TAGLGMVA 259
S SQ + + T +R WR P G K N DAS++ + +G+V
Sbjct: 885 QDANLMSTSQSNKQTTRERNINVTSARDRSWRPPPLGVYKLNSDASWKAPSPSCTVGVVI 706
Query: 260 RNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRT 318
R++ G ++ S P+ SP+++EA+A+R + LA LG I+ E+DC L A R+
Sbjct: 705 RDNLGLLISGTASRPLA-PSPIVSEALALREGLILARSLGLDRILSESDCQPLIEACRS 532
>CN825238
Length = 692
Score = 60.5 bits (145), Expect = 5e-10
Identities = 39/128 (30%), Positives = 59/128 (45%), Gaps = 1/128 (0%)
Frame = +3
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W LI+ +F TA +I SIPL R+ W S NG ++ ++ Y ++ ++
Sbjct: 321 WNVQLIDQLFPKETAAVIKSIPLGWRTTRNSCLWRWSRNGAFSVRSAYYNIQVAKEMQT- 497
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
SSS A F WK W T + K +R +P R+ L RR + V C C+ +
Sbjct: 498 SSSAASSFP---WKKLWRTPVPNKMKHFAYRVARNIIPCRENLQRRGIAVQNECPTCNHQ 668
Query: 127 -NETAEHL 133
E+ HL
Sbjct: 669 IPESMSHL 692
>TC9793
Length = 568
Score = 60.5 bits (145), Expect = 5e-10
Identities = 34/120 (28%), Positives = 58/120 (48%)
Frame = +2
Query: 231 WRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRW 290
W P G IK N DA + G + G +VAR+ + AA P LAEA+A+RW
Sbjct: 62 WEPPPEGIIKVNIDAGWSGSCSTGFSLVARSHNAVMLVAATHLEETQIEPTLAEALALRW 241
Query: 291 TMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSLCFVRR 350
+ + ++ I++E+ L + N + R + +++ D R L + F F++ + R
Sbjct: 242 CLSMIKEMEMERIIVESYSLVVLNTLHQ-KTKRPDIELVVYDSRELAAEFNFINFKHIGR 418
>AV780729
Length = 475
Score = 55.5 bits (132), Expect = 2e-08
Identities = 44/146 (30%), Positives = 67/146 (45%), Gaps = 3/146 (2%)
Frame = +3
Query: 156 FIAFADFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTG---VLRRVAAMN 212
F + +FW ++ GD++ I I+ ++ Q Q + F G L +VAA
Sbjct: 39 FSS*TEFWVAMIHDGDADLIGADTDIVLHFMRSKEQRQV*AQGFLSFGSGAALTKVAA-T 215
Query: 213 SASQKSDGGPQTGSRPARWRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACS 272
SA + T + P + F G + GMVARN E E +A+ +
Sbjct: 216 SARGLA*TESLTHGSARDVEKAYPRCVHG*F*CYI*GWKSFRYGMVARNYECEILASVTA 395
Query: 273 FPVPISSPLLAEAMAMRWTMQLAIDL 298
+PV SPLLAEA RW ++LA++L
Sbjct: 396 YPVTSLSPLLAEAGCFRWALRLAMEL 473
>BP046915
Length = 401
Score = 51.2 bits (121), Expect = 3e-07
Identities = 40/124 (32%), Positives = 59/124 (47%), Gaps = 3/124 (2%)
Frame = -2
Query: 252 TAGLGMVARNSEGEAMAAACSFPV-PISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCL 310
TA LGMVAR+S+G A+ A F + LA+++ + W M LAIDLG ++ +
Sbjct: 364 TASLGMVARDSDGAALGAVACFQCRQLCLW*LAKSLGLCWAMSLAIDLGLINLLRDRLFA 185
Query: 311 NLFNAWRTIEEDR--SYLSILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARNANTYAN 368
N + + S ++ DC ++ F + R+G VADFL N Y
Sbjct: 184 NSSTLGNFFDPGSHWPWWSRIVIDCL*VLMLFLY---RLFXRSGTCVADFLTMNIFIYP* 14
Query: 369 MVWV 372
VWV
Sbjct: 13 SVWV 2
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.138 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,696,487
Number of Sequences: 28460
Number of extensions: 112352
Number of successful extensions: 879
Number of sequences better than 10.0: 87
Number of HSP's better than 10.0 without gapping: 860
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 863
length of query: 387
length of database: 4,897,600
effective HSP length: 92
effective length of query: 295
effective length of database: 2,279,280
effective search space: 672387600
effective search space used: 672387600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0052a.1


