

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0051b.7
(189 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
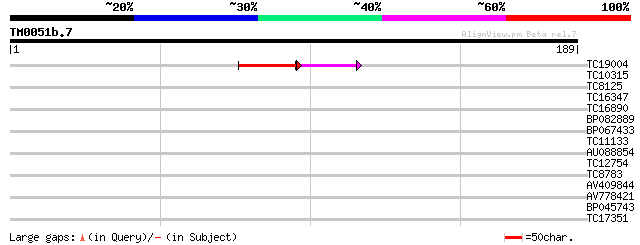
Score E
Sequences producing significant alignments: (bits) Value
TC19004 32 7e-05
TC10315 similar to UP|P93166 (P93166) SCOF-1, partial (42%) 28 0.99
TC8125 28 0.99
TC16347 similar to GB|AAC96325.1|4028592|AF104902 ZIC2 protein {... 27 1.7
TC16890 27 2.9
BP082889 26 3.8
BP067433 26 3.8
TC11133 26 3.8
AU088854 26 4.9
TC12754 similar to UP|Q9FHM5 (Q9FHM5) Similarity to DNA-binding ... 25 6.4
TC8783 similar to UP|Q8L759 (Q8L759) Expressed protein (At5g5765... 25 6.4
AV409844 25 6.4
AV778421 25 8.4
BP045743 25 8.4
TC17351 weakly similar to UP|Q8L7T3 (Q8L7T3) Ubiquitin-conjugati... 25 8.4
>TC19004
Length = 445
Score = 32.3 bits (72), Expect(2) = 7e-05
Identities = 15/21 (71%), Positives = 15/21 (71%)
Frame = -3
Query: 77 VRSDWAPNRNRPIQLIWSRFF 97
VRSD APNRNRP Q WS F
Sbjct: 395 VRSDQAPNRNRPNQPKWSSVF 333
Score = 28.9 bits (63), Expect(2) = 7e-05
Identities = 12/22 (54%), Positives = 13/22 (58%)
Frame = -2
Query: 96 FFPSLNPPSDPSEPSGPVFGRG 117
F P LNPP DP+EP P G
Sbjct: 336 FSPLLNPPPDPTEPRRPFLVAG 271
Score = 28.1 bits (61), Expect = 0.99
Identities = 12/28 (42%), Positives = 14/28 (49%)
Frame = -1
Query: 94 SRFFPSLNPPSDPSEPSGPVFGRGPVGF 121
+RFF P + P FG GPVGF
Sbjct: 343 ARFFSITQPATRPDRTEKAFFGSGPVGF 260
>TC10315 similar to UP|P93166 (P93166) SCOF-1, partial (42%)
Length = 592
Score = 28.1 bits (61), Expect = 0.99
Identities = 15/32 (46%), Positives = 20/32 (61%)
Frame = -2
Query: 40 SMEEEGKKLRRCSGGVARVVAAQRMAAQVMRR 71
S+EE + CSGGVA VVAA A+ + +R
Sbjct: 384 SLEEPEGAVAGCSGGVAVVVAAPPRASMMRQR 289
>TC8125
Length = 1373
Score = 28.1 bits (61), Expect = 0.99
Identities = 26/93 (27%), Positives = 38/93 (39%), Gaps = 3/93 (3%)
Frame = -2
Query: 6 LDRFRVGRVTGGSGRFHRNRLRQNEDEQAISDQRSMEEEGKKLRRCSGGV---ARVVAAQ 62
L R R ++ G R H N R + +Q+ + LRR SG V +V +
Sbjct: 529 LPRHRRHQLADGGFRPHENLHRPEASKNRHLEQQLRRLSRQTLRRRSGVVPVRQKVRVSL 350
Query: 63 RMAAQVMRRRVVAVVRSDWAPNRNRPIQLIWSR 95
R A + VVAV R + P +W+R
Sbjct: 349 RTAERSRSHGVVAVARRRSLNRQPSPPVAVWTR 251
>TC16347 similar to GB|AAC96325.1|4028592|AF104902 ZIC2 protein {Homo
sapiens;} , partial (5%)
Length = 599
Score = 27.3 bits (59), Expect = 1.7
Identities = 12/26 (46%), Positives = 14/26 (53%)
Frame = -3
Query: 87 RPIQLIWSRFFPSLNPPSDPSEPSGP 112
R I ++ PSL PP PSEP P
Sbjct: 375 RMISMLPPPAMPSLTPPPQPSEPPSP 298
>TC16890
Length = 965
Score = 26.6 bits (57), Expect = 2.9
Identities = 17/57 (29%), Positives = 27/57 (46%)
Frame = +3
Query: 46 KKLRRCSGGVARVVAAQRMAAQVMRRRVVAVVRSDWAPNRNRPIQLIWSRFFPSLNP 102
+K+RRC G + + +R + R R+ S W + +P L+ F SLNP
Sbjct: 231 RKMRRCQGLLFPISRPRRKRLRGRRWRLERRFSSSWVGLKKKPNVLL--PFVRSLNP 395
>BP082889
Length = 347
Score = 26.2 bits (56), Expect = 3.8
Identities = 10/12 (83%), Positives = 12/12 (99%)
Frame = -2
Query: 122 VQEVMVKENVHI 133
VQE+MVKENVH+
Sbjct: 256 VQEIMVKENVHL 221
>BP067433
Length = 421
Score = 26.2 bits (56), Expect = 3.8
Identities = 10/12 (83%), Positives = 12/12 (99%)
Frame = -2
Query: 122 VQEVMVKENVHI 133
VQE+MVKENVH+
Sbjct: 360 VQEIMVKENVHL 325
>TC11133
Length = 598
Score = 26.2 bits (56), Expect = 3.8
Identities = 10/12 (83%), Positives = 12/12 (99%)
Frame = +2
Query: 122 VQEVMVKENVHI 133
VQE+MVKENVH+
Sbjct: 26 VQEIMVKENVHL 61
>AU088854
Length = 675
Score = 25.8 bits (55), Expect = 4.9
Identities = 14/37 (37%), Positives = 17/37 (45%), Gaps = 1/37 (2%)
Frame = +3
Query: 71 RRVVAVVRSDWAPN-RNRPIQLIWSRFFPSLNPPSDP 106
+R V DW R+ P W R PSL+ PS P
Sbjct: 78 KRSVRTEAKDWKSRPRSHPQPWPWFRSLPSLSLPSQP 188
>TC12754 similar to UP|Q9FHM5 (Q9FHM5) Similarity to DNA-binding protein,
partial (24%)
Length = 932
Score = 25.4 bits (54), Expect = 6.4
Identities = 14/42 (33%), Positives = 18/42 (42%), Gaps = 3/42 (7%)
Frame = +1
Query: 79 SDWAPNRNRPIQLIWSRF---FPSLNPPSDPSEPSGPVFGRG 117
S W R RP++ I F F S PP+ P G + G
Sbjct: 613 SAWKRGRGRPVESIKKSFKLDFESPGPPAAPGPGEGIAYSIG 738
>TC8783 similar to UP|Q8L759 (Q8L759) Expressed protein (At5g57655),
partial (49%)
Length = 1243
Score = 25.4 bits (54), Expect = 6.4
Identities = 12/38 (31%), Positives = 18/38 (46%)
Frame = -2
Query: 66 AQVMRRRVVAVVRSDWAPNRNRPIQLIWSRFFPSLNPP 103
A +R R + + + WA N++ L F LNPP
Sbjct: 504 AAFLRPRAIVSIPTMWAMNKSSTSVLSLRNFASKLNPP 391
>AV409844
Length = 419
Score = 25.4 bits (54), Expect = 6.4
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = +2
Query: 2 RCVGLDRFRVGRVTGGSG 19
+CVGL R RVG G SG
Sbjct: 134 KCVGLQRHRVGH*WGSSG 187
>AV778421
Length = 211
Score = 25.0 bits (53), Expect = 8.4
Identities = 8/19 (42%), Positives = 12/19 (63%)
Frame = +1
Query: 93 WSRFFPSLNPPSDPSEPSG 111
WS +P++ P SD P+G
Sbjct: 40 WSSLYPTIRPHSDTEYPTG 96
>BP045743
Length = 512
Score = 25.0 bits (53), Expect = 8.4
Identities = 14/43 (32%), Positives = 23/43 (52%), Gaps = 2/43 (4%)
Frame = +2
Query: 64 MAAQVMRRRV--VAVVRSDWAPNRNRPIQLIWSRFFPSLNPPS 104
MAA+ RR + + W P + P +L+ S +FPS + P+
Sbjct: 371 MAAKASSRRFH*CLLHGNSWKPPLSSPPKLVSSPYFPSQHSPN 499
>TC17351 weakly similar to UP|Q8L7T3 (Q8L7T3) Ubiquitin-conjugating enzyme
E2 (Ubiquitin-protein ligase) (Ubiquitin carrier
protein) (At1g50490) , partial (33%)
Length = 461
Score = 25.0 bits (53), Expect = 8.4
Identities = 16/46 (34%), Positives = 22/46 (47%)
Frame = -1
Query: 42 EEEGKKLRRCSGGVARVVAAQRMAAQVMRRRVVAVVRSDWAPNRNR 87
E G + RC+GGVA ++A + R + RS PN NR
Sbjct: 194 ETAGVRRLRCAGGVAGELSAVGLG------RTSMIARSRRRPNPNR 75
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.143 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,154,060
Number of Sequences: 28460
Number of extensions: 41406
Number of successful extensions: 375
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 372
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 374
length of query: 189
length of database: 4,897,600
effective HSP length: 85
effective length of query: 104
effective length of database: 2,478,500
effective search space: 257764000
effective search space used: 257764000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0051b.7


