

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0047c.1
(619 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
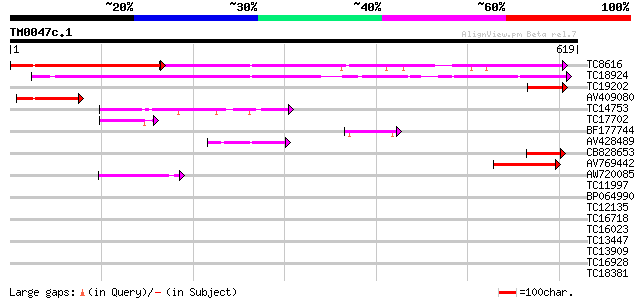
Score E
Sequences producing significant alignments: (bits) Value
TC8616 UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like pro... 293 e-135
TC18924 homologue to UP|Q94FN3 (Q94FN3) Phosphatidylinositol tra... 406 e-114
TC19202 similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (5%) 88 4e-18
AV409080 73 1e-13
TC14753 similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, partial... 59 2e-09
TC17702 weakly similar to GB|AAN73306.1|25141223|BT002309 At3g51... 51 5e-07
BF177744 49 2e-06
AV428489 47 9e-06
CB828653 46 2e-05
AV769442 45 3e-05
AW720085 41 5e-04
TC11997 similar to UP|O48940 (O48940) Polyphosphoinositide bindi... 38 0.004
BP064990 37 0.009
TC12135 similar to GB|AAP37803.1|30725562|BT008444 At4g35750 {Ar... 37 0.012
TC16718 32 0.23
TC16023 32 0.30
TC13447 32 0.30
TC13909 weakly similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, ... 32 0.30
TC16928 UP|Q94FN0 (Q94FN0) Phosphatidylinositol transfer-like pr... 30 1.1
TC18381 similar to UP|Q9ASR8 (Q9ASR8) AT5g59960/mmn10_180, parti... 30 1.5
>TC8616 UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like protein III,
complete
Length = 2302
Score = 293 bits (750), Expect(2) = e-135
Identities = 172/465 (36%), Positives = 270/465 (57%), Gaps = 19/465 (4%)
Frame = +3
Query: 164 IERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGL 223
+ERLGK P++LM++TT+DRY++YHVQEFE++ KFPAC++AAKR I S+TTILDVQG+
Sbjct: 603 LERLGKVDPNKLMQVTTMDRYVRYHVQEFEKSFAIKFPACTMAAKRHIDSSTTILDVQGV 782
Query: 224 GMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKI 283
G+KNFT++A L+ + K+D + YPETL QM+I+NAG GF ++LW + F DPKT +KI
Sbjct: 783 GLKNFTKSARELITRLQKVDGDNYPETLCQMFIINAGPGF-RLLWNTVKSFLDPKTTSKI 959
Query: 284 QILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATL 343
+L +K K LEVID+S+LP+ LGG+CTC +GGCL++ KGPW +P+I+K+V + E
Sbjct: 960 HVLGNKYHSKWLEVIDASELPECLGGACTCEDQGGCLRSDKGPWKNPEILKMVLNGEPRR 1139
Query: 344 VRQLSRASNEQQNFDSF---QVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAP 400
RQ+ + N + ++ + ++KG SD+STAESGS+ D +SP ++ +HLRL P
Sbjct: 1140 ARQVVKVLNSEGKVIAYAKPRYPTVKG--SDTSTAESGSEAEDIASPKAMKNYSHLRLTP 1313
Query: 401 VDEEVRVPD----LNGYYSCDDSAPTTEKVVE---NGQFHLSQKQSLQTNDMENVTHRTI 453
V EE ++ N + D+ P +K V+ Q L + + Q T RT
Sbjct: 1314 VREEAKIVGKSSYTNNFSGYDEYVPMVDKPVDAVLKKQASLQRSYTSQGAPSRPATQRT- 1490
Query: 454 SEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWR---TQNNVHPS 510
EG L I +F+ + T R + + ++ H
Sbjct: 1491 PEGIQARILVAI-----------------TAFLLTIFTIFRQVACRVTKKLPAISSNHDQ 1619
Query: 511 NTTEHNINN------HSAAVEAAFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKM 564
+T+E + S++ A E + + +RL LE+ + L +KP MP EKE++
Sbjct: 1620 STSEPTFDTTVVEVIPSSSTPAHTEENLLPSMLKRLGELEEKVDTLQSKPSEMPYEKEEL 1799
Query: 565 LMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSNCR 609
L ++ R+ ++E +L TK+ L+ A+ +Q E+ ++ ++ R
Sbjct: 1800 LNAAVCRVDALEAELIATKKALYEALMRQEELLAYIDRQAEAKLR 1934
Score = 206 bits (523), Expect(2) = e-135
Identities = 105/171 (61%), Positives = 131/171 (76%), Gaps = 1/171 (0%)
Frame = +2
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRG-KRKIDYRV 59
+EG +DE RERRSD ENSEDERR+ +IG+L+KKA++AS+KF HSL+K+ +RK D RV
Sbjct: 113 FEGFSGSDEKRERRSDFENSEDERRT-RIGSLKKKALSASTKFKHSLRKKSSRRKSDGRV 289
Query: 60 PAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEE 119
+VSIEDVRD E V RQ LI LLP DDY+ +LRFLKAR F+IEK MW E
Sbjct: 290 SSVSIEDVRDVEELQAVDAFRQSLIMDELLPQAFDDYHMMLRFLKARKFDIEKAKHMWAE 469
Query: 120 MLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKA 170
ML WRKE+G DTI+++FEF+EL+EV++YYP G+HGVDKEGRPVYI K+
Sbjct: 470 MLQWRKEFGADTIMQDFEFQELDEVVRYYPHGHHGVDKEGRPVYIRETWKS 622
>TC18924 homologue to UP|Q94FN3 (Q94FN3) Phosphatidylinositol transfer-like
protein II, complete
Length = 1976
Score = 406 bits (1043), Expect = e-114
Identities = 234/591 (39%), Positives = 343/591 (57%), Gaps = 2/591 (0%)
Frame = +3
Query: 25 RSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETVVHELRQRLI 84
+S K G+L+KKAM+ S + R R+ +V +V IEDVRD + V E RQ LI
Sbjct: 78 KSDKAGSLKKKAMSLRSSLS-----RKSRRSSSKVMSVEIEDVRDAEDLKAVDEFRQALI 242
Query: 85 ERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEV 144
LLP +HDDY+ LLRFLKAR F+ EK+ QMW +ML WRKE+G DTI+E+F+F+E++EV
Sbjct: 243 LDELLPEKHDDYHQLLRFLKARKFDTEKSKQMWSDMLQWRKEFGADTIVEDFDFKEIDEV 422
Query: 145 LQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACS 204
++YYP G+HGVDK+GRPVYIE +G+ ++LM++TT+DRY+KYHV+EFER KF ACS
Sbjct: 423 VKYYPHGHHGVDKDGRPVYIENIGQVDATKLMQVTTMDRYIKYHVKEFERTFDLKFAACS 602
Query: 205 IAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFK 264
IAAK+ I +TTILDVQG+G+KNF + A L+ + KID + YPETL++M+I+NAGSGF
Sbjct: 603 IAAKKHIDQSTTILDVQGVGLKNFNKHARELITRLQKIDGDNYPETLNRMFIINAGSGF- 779
Query: 265 KMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSK 324
+MLW + F DPKT +KI +L +K KLLEVID+SQLP+FLGG+CTC +GGC+++ K
Sbjct: 780 RMLWSTVKSFLDPKTTSKIHVLGNKYQSKLLEVIDASQLPEFLGGTCTCADQGGCMRSDK 959
Query: 325 GPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCSDSSTAESGSDINDY 384
GPW DP+++++V + E H +C E
Sbjct: 960 GPWKDPELVRMVQNGE----------------------HKCSRKCESPVVEEKKIP---- 1061
Query: 385 SSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTT--EKVVENGQFHLSQKQSLQT 442
T+ ++ +L+ V EV Y +A T +K+ EN QF +S K ++QT
Sbjct: 1062EETTKMGANFTSQLSSVFGEVPATKACNYEDFVPAADETVWKKIEENEQFQMS-KAAVQT 1238
Query: 443 NDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWR 502
+M + +I EK N T V+A F+ +T +R R +
Sbjct: 1239FNMADSC------------------KIHEKVNSQIFTGVMA-FVMGNVTMVRMTRNMPKK 1361
Query: 503 TQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKE 562
+ SN+ N S + + +R+ +LE+ +N K MP EKE
Sbjct: 1362LTDANFYSNSVYSGGQNPSDQTNPSISAQEFMTVMKRMAKLEEKMGNMNYKTC-MPPEKE 1538
Query: 563 KMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSNCRVSWS 613
+ML ++ R ++E +L TK+ L ++ +Q E++ +E +K +W+
Sbjct: 1539EMLNAAISRADALEQELMSTKKALEDSLAKQEELSAYIEKKKKKKKLFAWA 1691
>TC19202 similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (5%)
Length = 526
Score = 88.2 bits (217), Expect = 4e-18
Identities = 44/44 (100%), Positives = 44/44 (100%)
Frame = +3
Query: 566 MDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSNCR 609
MDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSNCR
Sbjct: 3 MDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSNCR 134
>AV409080
Length = 423
Score = 73.2 bits (178), Expect = 1e-13
Identities = 44/74 (59%), Positives = 53/74 (71%), Gaps = 1/74 (1%)
Frame = +2
Query: 8 DEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSL-KKRGKRKIDYRVPAVSIED 66
DE R+RRSD E SEDERR+ +IG+L+KKA+NASSKF HSL KK +RK R +VSIED
Sbjct: 203 DERRDRRSDYEVSEDERRT-RIGSLKKKAINASSKFRHSLRKKSSRRKNASRSNSVSIED 379
Query: 67 VRDEGEETVVHELR 80
+RD E V R
Sbjct: 380 IRDVKELQAVDAFR 421
>TC14753 similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, partial (57%)
Length = 1472
Score = 59.3 bits (142), Expect = 2e-09
Identities = 58/223 (26%), Positives = 101/223 (45%), Gaps = 12/223 (5%)
Frame = +1
Query: 99 LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKE 158
LL+FL+ARDF ++ M + + WRKE+G D +++E E ++V+ + QG+ DK
Sbjct: 91 LLKFLRARDFKVKDAFTMIKNTVRWRKEFGIDALVDEDLGSEWDKVV--FTQGH---DKL 255
Query: 159 GRPVYIERLGKAHPSRLMRITTID-----RYLKYHVQEFERALQEKFPACSIAAKRRIFS 213
G V G+ L + T D +++K+ +Q E+++++ + + A I
Sbjct: 256 GHAVCYNVFGEFEDKELYQKTFSDDEKRQKFIKWRIQVLEKSVRKLDFSPTPTAISTIVQ 435
Query: 214 TTTILDVQGLG---MKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAG---SGFKKML 267
+ + GLG ++ T A LL + YPE + + +N F +M+
Sbjct: 436 VNDLKNSPGLGKWELRQATNQALQLL-------QDNYPEFVAKQVFINVPWWYLAFSRMI 594
Query: 268 WPAAQKFCDPKTIAKIQIL-DSKSLYKLLEVIDSSQLPDFLGG 309
P F +T +K SKS L + I +P GG
Sbjct: 595 SP----FLTQRTKSKFVFAGPSKSAETLFKYIAPELVPVQYGG 711
>TC17702 weakly similar to GB|AAN73306.1|25141223|BT002309
At3g51670/T18N14_50 {Arabidopsis thaliana;}, partial
(20%)
Length = 574
Score = 51.2 bits (121), Expect = 5e-07
Identities = 31/68 (45%), Positives = 41/68 (59%), Gaps = 4/68 (5%)
Frame = +2
Query: 99 LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTIL-EEFEFEELEEVL---QYYPQGYHG 154
LL+FL+ARDF + + M + L WRKE+G D I+ EE F+ELE V+ Q Y
Sbjct: 377 LLKFLRARDFRVCDALNMLLKCLAWRKEFGADNIVDEELGFKELEGVVACTQVY------ 538
Query: 155 VDKEGRPV 162
D+EG PV
Sbjct: 539 -DREGHPV 559
>BF177744
Length = 251
Score = 49.3 bits (116), Expect = 2e-06
Identities = 30/73 (41%), Positives = 39/73 (53%), Gaps = 11/73 (15%)
Frame = +1
Query: 366 KGRC-----SDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSC---- 416
K RC SD+STAESGS+ D +SP +S +HLRL PV EE +V + Y
Sbjct: 31 KPRCPMIKGSDTSTAESGSETEDIASPKTMKSYSHLRLTPVREEAKVVGKSNYAGSGNLA 210
Query: 417 --DDSAPTTEKVV 427
D+ P +K V
Sbjct: 211 GYDEYIPMVDKAV 249
>AV428489
Length = 386
Score = 47.0 bits (110), Expect = 9e-06
Identities = 29/90 (32%), Positives = 43/90 (47%)
Frame = +1
Query: 217 ILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCD 276
+LD+ GL + LL A++ +D YPE YIVNA F W +
Sbjct: 22 VLDMSGLKFSALNQL--RLLTAISTVDDLNYPEKTDTYYIVNAPYVF-SACWKVVKPLLQ 192
Query: 277 PKTIAKIQILDSKSLYKLLEVIDSSQLPDF 306
+T KIQ+L +LL+++D + LP F
Sbjct: 193 ERTRRKIQVLQGSGKEELLKIMDYASLPHF 282
>CB828653
Length = 524
Score = 45.8 bits (107), Expect = 2e-05
Identities = 19/42 (45%), Positives = 34/42 (80%)
Frame = +1
Query: 565 LMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKS 606
L +S+ RIKS+E+DL+KTK+ L + +Q+E+A+ +E+L++S
Sbjct: 1 LQESLSRIKSIEYDLQKTKKALLATASKQVELAESLESLKES 126
>AV769442
Length = 559
Score = 45.4 bits (106), Expect = 3e-05
Identities = 24/73 (32%), Positives = 46/73 (62%)
Frame = -2
Query: 529 ERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHS 588
E + + ++RL LE+ + LN+KP MP EKE++L ++ R+ ++E +L TK+ L+
Sbjct: 492 EAELLSSASKRLCELEEKVDILNSKPNIMPYEKEELLNAAVYRVDALEAELIATKKALYE 313
Query: 589 AVKQQLEIADLVE 601
A+ +Q E+ ++
Sbjct: 312 ALIRQEELLAYID 274
>AW720085
Length = 524
Score = 41.2 bits (95), Expect = 5e-04
Identities = 29/95 (30%), Positives = 47/95 (48%), Gaps = 2/95 (2%)
Frame = +1
Query: 98 TLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEE--LEEVLQYYPQGYHGV 155
TL+RFLKAR++N K +M + L WR + D+IL + E V G G
Sbjct: 232 TLMRFLKAREWNASKAHKMLVDCLNWRVQSEIDSILSKPIIPEDLYRAVRDSQLIGLSGF 411
Query: 156 DKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQ 190
+EG PV+ +G + + ++ Y++ H+Q
Sbjct: 412 SREGLPVFAIGVGLSTFDK----ASVHYYVQSHIQ 504
>TC11997 similar to UP|O48940 (O48940) Polyphosphoinositide binding protein
Ssh2p, partial (26%)
Length = 492
Score = 38.1 bits (87), Expect = 0.004
Identities = 24/58 (41%), Positives = 33/58 (56%), Gaps = 1/58 (1%)
Frame = +2
Query: 253 QMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYK-LLEVIDSSQLPDFLGG 309
+++IV+A F K+ W F D T KI +D++ L LLE ID SQLP+ GG
Sbjct: 2 KLFIVHAPYIFMKV-WKLVYPFIDDHTKKKIVFVDNQKLKSTLLEEIDESQLPEIYGG 172
>BP064990
Length = 470
Score = 37.0 bits (84), Expect = 0.009
Identities = 20/50 (40%), Positives = 31/50 (62%)
Frame = +1
Query: 29 IGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETVVHE 78
+ + +KKA+NAS+ +SL ++G+R +V +V IEDV D E V E
Sbjct: 322 VDSFKKKAINASNMLRNSLTRKGRR--SSKVMSVEIEDVHDAEELKAVEE 465
>TC12135 similar to GB|AAP37803.1|30725562|BT008444 At4g35750 {Arabidopsis
thaliana;}, partial (78%)
Length = 562
Score = 36.6 bits (83), Expect = 0.012
Identities = 21/68 (30%), Positives = 41/68 (59%)
Frame = +3
Query: 133 LEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEF 192
+ +FE EEL + L+ + G DK+GR + + +GK P+RL+ + + +YL+ + F
Sbjct: 87 ISQFEQEELLDKLEVFK--IKGRDKQGRKI-LRIIGKFFPARLVSVDVLKKYLEERI--F 251
Query: 193 ERALQEKF 200
+ +++KF
Sbjct: 252 PKLVKKKF 275
>TC16718
Length = 464
Score = 32.3 bits (72), Expect = 0.23
Identities = 18/66 (27%), Positives = 31/66 (46%), Gaps = 4/66 (6%)
Frame = -2
Query: 458 SVNNLFRII----KEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTT 513
S+ N F I K +K+ ++Y+ + A + +RSL F W +HP+ T
Sbjct: 433 SLGNYFDITSRQGKNYQQKSYYMYINEIYAWY*AHAPNPLRSLCFISWIINTYIHPTRTQ 254
Query: 514 EHNINN 519
+N+ N
Sbjct: 253 HNNVAN 236
>TC16023
Length = 894
Score = 32.0 bits (71), Expect = 0.30
Identities = 15/30 (50%), Positives = 22/30 (73%)
Frame = -3
Query: 487 ERLITFIRSLRFEFWRTQNNVHPSNTTEHN 516
E++IT R+ R WR QN++HPS+TT H+
Sbjct: 682 EQVITRRRNTR*R-WRFQNSIHPSSTTPHH 596
>TC13447
Length = 554
Score = 32.0 bits (71), Expect = 0.30
Identities = 32/134 (23%), Positives = 60/134 (43%)
Frame = +2
Query: 94 DDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYH 153
DD + FLK R F++++ + + + WR+++ + EE + Y +
Sbjct: 173 DDEDMIQWFLKDRKFSVDEAVSKLTKAIKWRQDFEVSKLTEEAVKDAARSGKAYV---HD 343
Query: 154 GVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFS 213
+D GRPV + K P + + +R + V E+AL K P + K ++
Sbjct: 344 FLDINGRPVLVVVASKHFP-KAQDPSDDERLCVFLV---EKAL-SKLP----SGKEQVLG 496
Query: 214 TTTILDVQGLGMKN 227
I+D++G G +N
Sbjct: 497 ---IVDLRGFGTEN 529
>TC13909 weakly similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, partial
(20%)
Length = 399
Score = 32.0 bits (71), Expect = 0.30
Identities = 12/29 (41%), Positives = 20/29 (68%)
Frame = +3
Query: 99 LLRFLKARDFNIEKTIQMWEEMLLWRKEY 127
LL+FL+AR+F ++ + M L WRK++
Sbjct: 306 LLKFLRAREFKVKDALLMINNTLRWRKDF 392
>TC16928 UP|Q94FN0 (Q94FN0) Phosphatidylinositol transfer-like protein IV,
partial (12%)
Length = 442
Score = 30.0 bits (66), Expect = 1.1
Identities = 12/20 (60%), Positives = 19/20 (95%)
Frame = -2
Query: 249 ETLHQMYIVNAGSGFKKMLW 268
+TL++M+I+NAGSGF ++LW
Sbjct: 99 QTLNRMFIINAGSGF-RILW 43
Score = 27.3 bits (59), Expect = 7.4
Identities = 13/40 (32%), Positives = 20/40 (49%)
Frame = -3
Query: 224 GMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGF 263
G+K+F + A L+ + K+D + YPE I G F
Sbjct: 278 GLKSFNKHARELVTRIQKVDGDNYPEVCFSTLIKLWGQEF 159
>TC18381 similar to UP|Q9ASR8 (Q9ASR8) AT5g59960/mmn10_180, partial (53%)
Length = 758
Score = 29.6 bits (65), Expect = 1.5
Identities = 19/70 (27%), Positives = 34/70 (48%)
Frame = -2
Query: 469 IVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAF 528
I E+ +YV +++ S +E+ E + H NTT+H HS ++
Sbjct: 679 ICERGEQIYVKQMIDSRVEQSCQG*VCHSQELVQNNQQAHHRNTTDH----HSHTLQ--- 521
Query: 529 ERDHILPCAQ 538
++ H+LPC+Q
Sbjct: 520 KKSHLLPCSQ 491
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,666,893
Number of Sequences: 28460
Number of extensions: 117382
Number of successful extensions: 669
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 657
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 660
length of query: 619
length of database: 4,897,600
effective HSP length: 96
effective length of query: 523
effective length of database: 2,165,440
effective search space: 1132525120
effective search space used: 1132525120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0047c.1


