

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0046b.4
(1706 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
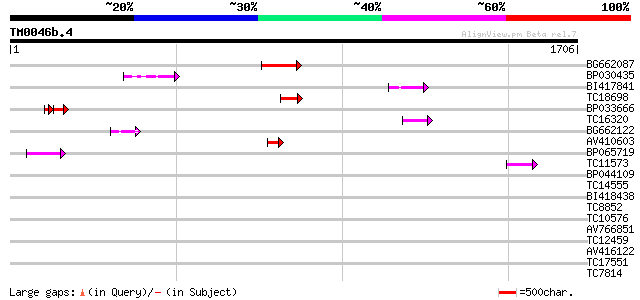
Score E
Sequences producing significant alignments: (bits) Value
BG662087 197 1e-50
BP030435 126 3e-29
BI417841 82 5e-16
TC18698 81 1e-15
BP033666 48 5e-12
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 62 1e-09
BG662122 59 8e-09
AV410603 53 4e-07
BP065719 47 3e-05
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 45 1e-04
BP044109 37 0.026
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 35 0.076
BI418438 35 0.099
TC8852 similar to UP|O24046 (O24046) 2-dehydro-3-deoxyphosphohep... 34 0.17
TC10576 similar to UP|Q93ZQ3 (Q93ZQ3) AT3g63200/F16M2_50, partia... 34 0.22
AV766851 33 0.29
TC12459 33 0.29
AV416122 33 0.38
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 33 0.38
TC7814 similar to UP|Q9LKJ6 (Q9LKJ6) Water-selective transport i... 28 9.3
>BG662087
Length = 373
Score = 197 bits (501), Expect = 1e-50
Identities = 90/120 (75%), Positives = 104/120 (86%)
Frame = +1
Query: 758 NGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEE 817
+GKWRM DYT LNK CPKDSYPLP++DKLVDGAS NELLSLMDAYSGY+QI MHPSDE+
Sbjct: 13 SGKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDED 192
Query: 818 STTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDH 877
T FMT + NYCY+T+PFGLKNAGATYQ LMD++F VGRNMEVY+D+MIVKSA ++H
Sbjct: 193 KTAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>BP030435
Length = 533
Score = 126 bits (316), Expect = 3e-29
Identities = 73/169 (43%), Positives = 97/169 (57%)
Frame = +1
Query: 341 CEYHKALGHTTDNCWNLRREIDRLIKAGHLANFVKDTAAPEVAKITQGDKGKGKEIVEEL 400
C YH+A+GH T+ C L+RE+ +LI G + +I G+ K +
Sbjct: 103 CVYHQAMGHITEECRTLQREVGKLIATG------------KPTRIQGGNAPVPKGV---- 234
Query: 401 GDPVGECSSIAGGFGGGAISSKARKRYVAAVHSVQESNEGECWVNHSPIIFTPQDFAHVI 460
+IAGGFGGG I+S ARKRY AV++V E G +H I F+ DF +
Sbjct: 235 -------HTIAGGFGGGGITSAARKRYARAVNTVTEVLFG---FSHPVITFSSDDFIRIK 384
Query: 461 PHDNDPIVVTIRVNNYVTKKVFLDQGSSADIIYGDAFDRLGLKESDLKP 509
PH ++PIV+ +RVN ++V LDQGSSADIIYG FDRLGL E+DL P
Sbjct: 385 PHLDNPIVILLRVNQLNVQRVLLDQGSSADIIYGGVFDRLGLNEADLTP 531
>BI417841
Length = 617
Score = 82.4 bits (202), Expect = 5e-16
Identities = 47/125 (37%), Positives = 70/125 (55%), Gaps = 4/125 (3%)
Frame = +1
Query: 1140 LSVDGSSN----NTGSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGLRLAIEL 1195
L DGSS + G+GA + E K+ + + + + +NNQ+EY LI GL+ A E
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKVYLREGVGNQ---TNNQAEYRGLILGLKHAHEQ 312
Query: 1196 GVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPRGQNERAD 1255
G Q + +KGDSQLV KQV+G ++ ++P ++ + L + + I HVPR N AD
Sbjct: 313 GYQHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEAD 492
Query: 1256 VLAKL 1260
V A L
Sbjct: 493 VQANL 507
>TC18698
Length = 808
Score = 81.3 bits (199), Expect = 1e-15
Identities = 39/66 (59%), Positives = 49/66 (74%)
Frame = -2
Query: 816 EESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARAS 875
++ TT N+ NY Y+ MP GLKN TYQRLMDKIF KQ+ +N+EVYV+DMIVKS++
Sbjct: 807 KKKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE* 628
Query: 876 DHGGDL 881
H GDL
Sbjct: 627 FHRGDL 610
>BP033666
Length = 342
Score = 47.8 bits (112), Expect(2) = 5e-12
Identities = 19/26 (73%), Positives = 23/26 (88%)
Frame = -2
Query: 105 TFKGMAMQWFIRQPPFSIDNFTDLST 130
+FKG+AM WFI+QPP+SI NFTD ST
Sbjct: 341 SFKGVAMTWFIKQPPYSISNFTDFST 264
Score = 41.6 bits (96), Expect(2) = 5e-12
Identities = 24/52 (46%), Positives = 32/52 (61%), Gaps = 6/52 (11%)
Frame = -1
Query: 132 FLTQFSANKTTKATMFDLISIHQQPGEKLKTYMA------RFSKMAVQLEDE 177
FLT FSAN+T KAT DL +I QQ GE+++ Y + R +KM V + E
Sbjct: 261 FLTXFSANQTQKATSADLFNICQQVGERVRPYFSLYCF*ERLNKMRVLFKFE 106
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 61.6 bits (148), Expect = 1e-09
Identities = 32/91 (35%), Positives = 49/91 (53%)
Frame = +2
Query: 1182 YEALIAGLRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEI 1241
Y LI GL+ AI+ G + + +KGDS LV QV+G +++K+ ++ + L +
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 1242 KIEHVPRGQNERADVLAKLASTGRLGNYQTV 1272
KI H+PR N ADV A + R G + V
Sbjct: 182 KINHIPREYNSEADVQANFGISLRAGQVEEV 274
>BG662122
Length = 386
Score = 58.5 bits (140), Expect = 8e-09
Identities = 35/94 (37%), Positives = 55/94 (58%), Gaps = 4/94 (4%)
Frame = +2
Query: 304 KLNAPLSTILRAVGQTNVV----QYPPPPRRPPANVDTNRWCEYHKALGHTTDNCWNLRR 359
+LNA L+ IL+ V ++V Q PPPRR +DT + CEY +++ H D+C+ L+R
Sbjct: 98 ELNAHLTDILQDVKAAHMVGKSGQS*PPPRR---GIDTTK*CEYRRSVVHDIDDCFTLKR 268
Query: 360 EIDRLIKAGHLANFVKDTAAPEVAKITQGDKGKG 393
EI++LIK G L + + + QGD+ +G
Sbjct: 269 EIEKLIKMGRLKQYDRGSR-------QQGDRQQG 349
>AV410603
Length = 162
Score = 53.1 bits (126), Expect = 4e-07
Identities = 22/49 (44%), Positives = 33/49 (66%)
Frame = +1
Query: 776 KDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTN 824
KDS+P+P VD+L+D G++ S +D SGY+QI++ P D T F T+
Sbjct: 10 KDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTH 156
>BP065719
Length = 567
Score = 47.0 bits (110), Expect = 3e-05
Identities = 29/120 (24%), Positives = 53/120 (44%), Gaps = 2/120 (1%)
Frame = +3
Query: 50 FVPAITRVDIPKHLQTMALDAFSGES--DPMEHLRYFNTKMVIGGATDAVKCRLLPSTFK 107
F+ + ++P+ + FSG+S +EH+ + + + +K + PS+
Sbjct: 135 FIEEVLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLT 314
Query: 108 GMAMQWFIRQPPFSIDNFTDLSTKFLTQFSANKTTKATMFDLISIHQQPGEKLKTYMARF 167
A WF P S+ + L F QF + K + DL S+ ++P E + Y+ RF
Sbjct: 315 KNAFTWFTTLAPRSVHTWAQLERIFHEQFFRGE-CKVSXKDLASVKRKPAESIDDYLNRF 491
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 44.7 bits (104), Expect = 1e-04
Identities = 32/93 (34%), Positives = 43/93 (45%)
Frame = +1
Query: 1496 NTIHK*CYPRILYRNGNRDAICLCRTSTDQWAG*IG*QSYPEGFKEETG*SKRALGGGTT 1555
N+IH+ L +GN D + + S++ WA Q EG KE *S R + T
Sbjct: 16 NSIHQQTNSGFLRWHGNSDEVLISEASSN*WADRGCEQGDFEGHKEAAI*S*RKMD**TP 195
Query: 1556 RRSLGIQHHGAVKHKGNPVSSNLWN*RHAPSGN 1588
+ I HH +KHK N +LW AP GN
Sbjct: 196 DSFVVI*HHATIKHKRNTFLIDLWGRYDAPCGN 294
>BP044109
Length = 497
Score = 37.0 bits (84), Expect = 0.026
Identities = 23/89 (25%), Positives = 31/89 (33%)
Frame = +2
Query: 243 SQDKSKAGKETRIAQTPRVPRPGPYQNSKPGFQNTWHRNPQGPPAATAGQTGASPAPVPL 302
S + K T QTP P P P + KP PP+ A + SP P
Sbjct: 155 SHASNATSKSTPRTQTPTTPPPSPTSSPKP------------PPSTFATKPSTSPPETPS 298
Query: 303 TKLNAPLSTILRAVGQTNVVQYPPPPRRP 331
+ + P T+ + P PP P
Sbjct: 299 SSSHGPAPTLPSPPSSSTPTSTPSPPSPP 385
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 35.4 bits (80), Expect = 0.076
Identities = 29/97 (29%), Positives = 41/97 (41%), Gaps = 7/97 (7%)
Frame = +2
Query: 243 SQDKSKAGKETRIAQTPRVPRPG-------PYQNSKPGFQNTWHRNPQGPPAATAGQTGA 295
+Q S + + TP P+P P Q+S P Q++ PPA ++ +
Sbjct: 182 AQSPSNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPVS 361
Query: 296 SPAPVPLTKLNAPLSTILRAVGQTNVVQYPPPPRRPP 332
SP PV T AP ST A + + PPP PP
Sbjct: 362 SPPPVQSTPPPAPASTPPPA----SPPPFSPPPATPP 460
>BI418438
Length = 431
Score = 35.0 bits (79), Expect = 0.099
Identities = 28/82 (34%), Positives = 34/82 (41%)
Frame = +1
Query: 253 TRIAQTPRVPRPGPYQNSKPGFQNTWHRNPQGPPAATAGQTGASPAPVPLTKLNAPLSTI 312
T I P P P P Q S P PPA+ + A+P P P + L PLS+
Sbjct: 232 TTIPPPPPSPTPPPRQKSS---------KPARPPASPSN---ANP-PSPTSLLTQPLSS- 369
Query: 313 LRAVGQTNVVQYPPPPRRPPAN 334
+N PPPP PP N
Sbjct: 370 -----SSNPPSPPPPPTSPPPN 420
>TC8852 similar to UP|O24046 (O24046) 2-dehydro-3-deoxyphosphoheptonate
aldolase precursor , partial (31%)
Length = 808
Score = 34.3 bits (77), Expect = 0.17
Identities = 18/63 (28%), Positives = 30/63 (47%)
Frame = +1
Query: 262 PRPGPYQNSKPGFQNTWHRNPQGPPAATAGQTGASPAPVPLTKLNAPLSTILRAVGQTNV 321
P PGP S P + + P PP T+ + +P P P +++ P++ LRA + +
Sbjct: 226 PNPGPIPPSSPSTPPSLQKTPSSPP--TSRRFHRNPPPPPPSEMPGPVNGPLRAGNRRRL 399
Query: 322 VQY 324
Y
Sbjct: 400 CSY 408
>TC10576 similar to UP|Q93ZQ3 (Q93ZQ3) AT3g63200/F16M2_50, partial (39%)
Length = 751
Score = 33.9 bits (76), Expect = 0.22
Identities = 29/108 (26%), Positives = 41/108 (37%), Gaps = 22/108 (20%)
Frame = -3
Query: 249 AGKETRIAQTPRVPRPGP-----YQNSKPGF-----------------QNTWHRNPQGPP 286
A ++T TP + P P + N PGF Q+ R Q P
Sbjct: 479 ASRQTVTTPTPSLSAPPPPLSLQFPNQTPGF*RSQYDSSRKRSLTSGQQSRKLRRAQCQP 300
Query: 287 AATAGQTGASPAPVPLTKLNAPLSTILRAVGQTNVVQYPPPPRRPPAN 334
G SP+P ++ P + +R+ G+ Q PPPR PP N
Sbjct: 299 PTAWSTCGGSPSP---SRSRMPSHSTIRSQGKE---QGDPPPRSPPGN 174
>AV766851
Length = 445
Score = 33.5 bits (75), Expect = 0.29
Identities = 28/114 (24%), Positives = 46/114 (39%), Gaps = 6/114 (5%)
Frame = +2
Query: 221 EQDDQKKQEREEGRKQSQSGGVSQDKSKAGKETRIAQTPRVPRPGPYQNSKPGFQNTWH- 279
++ + K++ERE+G +Q+ G S + ++ A Y+ + P + H
Sbjct: 71 KERENKEREREDGPRQTDRYGHSTGPPEGQMQSATAGCG-------YRRALPAYPLDIHP 229
Query: 280 -----RNPQGPPAATAGQTGASPAPVPLTKLNAPLSTILRAVGQTNVVQYPPPP 328
R+ PP G P+P P L PLS + R G+ PP P
Sbjct: 230 RTMPSRSATVPPPRGPGTHQPPPSPPPPPLLLEPLSAVERDTGEQGSR*RPPAP 391
>TC12459
Length = 797
Score = 33.5 bits (75), Expect = 0.29
Identities = 23/82 (28%), Positives = 38/82 (46%), Gaps = 6/82 (7%)
Frame = -2
Query: 258 TPRVPRPGPYQNSKPGFQNTWHRNPQGPPAATAGQTGASP------APVPLTKLNAPLST 311
+P++ + P + GF+ R PP+ T TG P A P T +P++
Sbjct: 364 SPQISKTLPNASKHLGFRCAKRRRSTTPPSPTT-LTGNGPR*TRCLADTPATPRTSPIA- 191
Query: 312 ILRAVGQTNVVQYPPPPRRPPA 333
+ + T+V+ PPPP PP+
Sbjct: 190 -VSSTTTTSVLPSPPPPPPPPS 128
>AV416122
Length = 414
Score = 33.1 bits (74), Expect = 0.38
Identities = 20/68 (29%), Positives = 32/68 (46%), Gaps = 1/68 (1%)
Frame = +1
Query: 261 VPRPGPYQNSKPGFQNTWHRNPQGPPAATAGQTGAS-PAPVPLTKLNAPLSTILRAVGQT 319
+P P ++NS T +NP PP+AT + P P P + T+++ T
Sbjct: 43 LPYPHSWRNS------TTTKNPHSPPSATPFLIPKNNPPPKPQSSPQPQNLTLIQTTTTT 204
Query: 320 NVVQYPPP 327
+Q+PPP
Sbjct: 205 TTLQFPPP 228
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 33.1 bits (74), Expect = 0.38
Identities = 29/80 (36%), Positives = 34/80 (42%), Gaps = 3/80 (3%)
Frame = +3
Query: 255 IAQTPRVPRPGPYQNSKPGFQNTWHRN---PQGPPAATAGQTGASPAPVPLTKLNAPLST 311
I TP P P P N HRN P PP+ T + +SPA PL L +P S
Sbjct: 60 IQTTPPPPPPPPSPN---------HRNNLPPPFPPSLTTSFSTSSPASHPLPSLPSPSSA 212
Query: 312 ILRAVGQTNVVQYPPPPRRP 331
AVG T+ PP P
Sbjct: 213 ---AVGPTS----STPPTSP 251
Score = 28.5 bits (62), Expect = 9.3
Identities = 21/78 (26%), Positives = 31/78 (38%), Gaps = 5/78 (6%)
Frame = +3
Query: 259 PRVPRP-----GPYQNSKPGFQNTWHRNPQGPPAATAGQTGASPAPVPLTKLNAPLSTIL 313
P +P P GP ++ P +P AAT+ T SP+P P + P S+
Sbjct: 186 PSLPSPSSAAVGPTSSTPP-------TSPTSAAAATSSATPPSPSPPPTSASPPPPSSTP 344
Query: 314 RAVGQTNVVQYPPPPRRP 331
++ P P RP
Sbjct: 345 NGTPPSSSPATTPSPSRP 398
>TC7814 similar to UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic
membrane protein 1, partial (37%)
Length = 660
Score = 28.5 bits (62), Expect = 9.3
Identities = 27/88 (30%), Positives = 39/88 (43%), Gaps = 4/88 (4%)
Frame = +2
Query: 249 AGKETRIAQTPRVP-RPGPYQNSKPGFQNTWHRNPQGPPAATAG--QTGASPAPVPLT-K 304
AG++ R A +P VP R P Q P G P ++ + +SPAPVP +
Sbjct: 368 AGRKCRSATSPSVPLRRKPTQT------------P*GXPXLSSSPPSSSSSPAPVPESPT 511
Query: 305 LNAPLSTILRAVGQTNVVQYPPPPRRPP 332
+N+P++ V PPP PP
Sbjct: 512 INSPMT-----------VPPPPPXSSPP 562
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,867,219
Number of Sequences: 28460
Number of extensions: 324540
Number of successful extensions: 2566
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 2440
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2538
length of query: 1706
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1603
effective length of database: 1,966,220
effective search space: 3151850660
effective search space used: 3151850660
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0046b.4


