

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0046a.5
(327 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
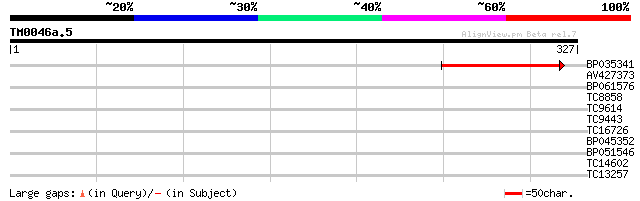
Score E
Sequences producing significant alignments: (bits) Value
BP035341 91 3e-19
AV427373 35 0.013
BP061576 34 0.029
TC8858 homologue to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 32 0.19
TC9614 similar to UP|O22799 (O22799) At2g33490 protein, partial ... 28 2.1
TC9443 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting p... 28 2.7
TC16726 similar to UP|GS11_ARATH (Q9LMP7) Golgi SNARE 11 protein... 28 2.7
BP045352 28 2.7
BP051546 27 4.7
TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinas... 27 6.1
TC13257 26 8.0
>BP035341
Length = 523
Score = 90.5 bits (223), Expect = 3e-19
Identities = 40/72 (55%), Positives = 54/72 (74%), Gaps = 1/72 (1%)
Frame = -3
Query: 250 SENGSGKTM-YFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPS 308
+ NGS +M YFL E P++ +SE EL+ S GD+IV+RKVT GW+EGEC G+AGWFP
Sbjct: 458 AHNGSTDSMGYFLGEVLFPYHAESEVELNLSAGDYIVIRKVTNNGWAEGECKGRAGWFPF 279
Query: 309 AYVEKRQRVPSS 320
Y+E+R+RV +S
Sbjct: 278 GYIERRERVLAS 243
>AV427373
Length = 393
Score = 35.4 bits (80), Expect = 0.013
Identities = 21/71 (29%), Positives = 32/71 (44%)
Frame = +3
Query: 75 TAIGYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKEHEELNRLLASQV 134
T G H+E TK+ +D K +NN + + A S L A +LLA Q
Sbjct: 189 TGTGEDHVENATKVKDDGAKTNLDNNFEKLKRAAVSTLAAAAVKA--------KLLADQE 344
Query: 135 LDPLRQMINGV 145
D +RQ+ + +
Sbjct: 345 EDEIRQLTSSL 377
>BP061576
Length = 528
Score = 34.3 bits (77), Expect = 0.029
Identities = 24/69 (34%), Positives = 35/69 (49%), Gaps = 1/69 (1%)
Frame = -2
Query: 165 ETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKE-AAAALASVDAQQQ 223
E R EI K VRES ++ A+ AAE + ELKA A K+ AL+ Q +
Sbjct: 509 ELARYEIDKYDAAVRESAYLKECAEREAAERKETELKAIRAAXEKDKLEGALSGSVLQYR 330
Query: 224 RLTFQRLVA 232
+ T+ +V+
Sbjct: 329 KFTWDEIVS 303
>TC8858 homologue to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (65%)
Length = 1272
Score = 31.6 bits (70), Expect = 0.19
Identities = 25/83 (30%), Positives = 41/83 (49%), Gaps = 2/83 (2%)
Frame = +1
Query: 162 QEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKE--AAAALASVD 219
+EA L+ E K+++++ E E++ +L AEA M +LKAN A E ALA D
Sbjct: 607 REAGELKTERQKKKLQIEEL---ERIVRLKNAEADMFQLKANEAKREAERLQRIALAKSD 777
Query: 220 AQQQRLTFQRLVAMMVSDRQKKE 242
++ T L + +K+
Sbjct: 778 KSEEEYTSNYLKQKLSEAEAEKQ 846
>TC9614 similar to UP|O22799 (O22799) At2g33490 protein, partial (12%)
Length = 673
Score = 28.1 bits (61), Expect = 2.1
Identities = 21/78 (26%), Positives = 36/78 (45%), Gaps = 3/78 (3%)
Frame = +2
Query: 49 QLEKLYKATR---TMRDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDNIL 105
QLE+L +ATR MRD ++ AA + Y+ E+ + + A N+ +
Sbjct: 329 QLEELAQATRDMQDMRDCYDTLLSAAAATASSAYEFAESLRDMGSCLLEKTALNDHEEET 508
Query: 106 AKAASVLGDARKHVEKEH 123
K +LG + ++K H
Sbjct: 509 GKVLLMLGKIQFKLQKTH 562
>TC9443 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (41%)
Length = 1025
Score = 27.7 bits (60), Expect = 2.7
Identities = 22/67 (32%), Positives = 33/67 (48%), Gaps = 2/67 (2%)
Frame = +1
Query: 162 QEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKE--AAAALASVD 219
+E L+ E K++ +V E E++ +L AEA M +LKAN A E A A D
Sbjct: 232 REVAELKMERQKKKSQVEEL---EKIVRLKNAEADMFQLKANEAKREAERLQRIAFAKPD 402
Query: 220 AQQQRLT 226
++ T
Sbjct: 403 KSEEEFT 423
>TC16726 similar to UP|GS11_ARATH (Q9LMP7) Golgi SNARE 11 protein (AtGOS11)
(Golgi SNAP receptor complex member 1-1), partial (80%)
Length = 655
Score = 27.7 bits (60), Expect = 2.7
Identities = 20/60 (33%), Positives = 29/60 (48%)
Frame = +2
Query: 2 DALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMR 61
DALRKQA L Q+ ++ + K S S +D D E I Q L++L + M+
Sbjct: 140 DALRKQARNLEAQLDERMSSFRKLVSASVSAKTDAAENDL-ESWIEQLLKQLQQVNSQMQ 316
>BP045352
Length = 434
Score = 27.7 bits (60), Expect = 2.7
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = -3
Query: 90 EDCCKYGAENNIDNILAKAASVLGDARKHVEKEHEELNR 128
E + AE N + ++ + + RKH+E+ HEEL +
Sbjct: 291 EPAARLKAEENANLARVRSDKEIRELRKHLERAHEELRQ 175
>BP051546
Length = 574
Score = 26.9 bits (58), Expect = 4.7
Identities = 11/32 (34%), Positives = 17/32 (52%)
Frame = -2
Query: 92 CCKYGAENNIDNILAKAASVLGDARKHVEKEH 123
CC G +NNID+ AK + H +++H
Sbjct: 510 CCAPGCKNNIDHPRAKPLKDFRTLQTHYKRKH 415
>TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1256
Score = 26.6 bits (57), Expect = 6.1
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = +2
Query: 255 GKTMYFLAEATHPFYGDSEKEL 276
G +Y + T PFYGDS E+
Sbjct: 629 GVILYIMLSGTPPFYGDSAAEI 694
>TC13257
Length = 544
Score = 26.2 bits (56), Expect = 8.0
Identities = 13/27 (48%), Positives = 16/27 (59%)
Frame = +1
Query: 241 KESAPPVVVSENGSGKTMYFLAEATHP 267
K PP S NG GKT+ +A+A HP
Sbjct: 55 KLKVPP---SSNGDGKTVAVVADAFHP 126
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.127 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,778,160
Number of Sequences: 28460
Number of extensions: 37065
Number of successful extensions: 168
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 168
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 168
length of query: 327
length of database: 4,897,600
effective HSP length: 91
effective length of query: 236
effective length of database: 2,307,740
effective search space: 544626640
effective search space used: 544626640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0046a.5


