

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
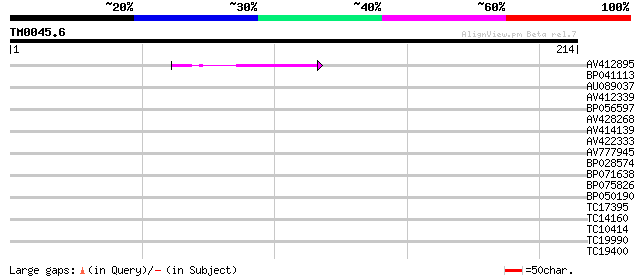
Query= TM0045.6
(214 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV412895 39 9e-04
BP041113 35 0.008
AU089037 35 0.010
AV412339 34 0.022
BP056597 34 0.022
AV428268 30 0.32
AV414139 30 0.41
AV422333 29 0.54
AV777945 28 0.92
BP028574 28 0.92
BP071638 28 1.6
BP075826 28 1.6
BP050190 26 4.6
TC17395 similar to UP|Q9LN83 (Q9LN83) T12C24.17, partial (37%) 26 6.0
TC14160 homologue to GB|BAB90396.1|20161472|AP003570 ADP-ribosyl... 26 6.0
TC10414 similar to GB|AAP13363.1|30023660|BT006255 At5g55710 {Ar... 26 6.0
TC19990 weakly similar to UP|AAQ88099 (AAQ88099) NADPH-dependent... 25 7.8
TC19400 25 7.8
>AV412895
Length = 405
Score = 38.5 bits (88), Expect = 9e-04
Identities = 23/57 (40%), Positives = 29/57 (50%)
Frame = +1
Query: 62 DTYFEHPDFEPLVEGPEALAQVLGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWK 118
D Y E PD P+ F+ +W AP N+KAF WR LL +I +R NL K
Sbjct: 43 DQYLEEPD--PV------------FN*IWMVPAPLNIKAFVWRVLLGRIQTRDNLLK 171
>BP041113
Length = 503
Score = 35.4 bits (80), Expect = 0.008
Identities = 18/33 (54%), Positives = 23/33 (69%)
Frame = +1
Query: 86 FSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWK 118
F+ +W APS VKAFAWR LL ++P+ NL K
Sbjct: 322 FATIWRTVAPS-VKAFAWRCLLGRLPTYDNLIK 417
>AU089037
Length = 245
Score = 35.0 bits (79), Expect = 0.010
Identities = 14/31 (45%), Positives = 20/31 (64%)
Frame = -1
Query: 86 FSHVWHFAAPSNVKAFAWRFLLVKIPSRLNL 116
F +W APSN +FAWR +L ++ +R NL
Sbjct: 200 FQKLWKVPAPSNAVSFAWRLILDRVQTRGNL 108
>AV412339
Length = 420
Score = 33.9 bits (76), Expect = 0.022
Identities = 17/52 (32%), Positives = 29/52 (55%)
Frame = +3
Query: 117 WKLGATRGLISFGYNRSKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYF 168
W G+ L FGY+ S D++L+V+I+LC N+IT+ + V ++
Sbjct: 252 WDSGS-HTLYGFGYDPSNDDYLLVVIRLCEFDPGRTNQITVFPGKTNVPSFY 404
>BP056597
Length = 414
Score = 33.9 bits (76), Expect = 0.022
Identities = 26/112 (23%), Positives = 48/112 (42%), Gaps = 16/112 (14%)
Frame = +3
Query: 21 EVMAIAHGYQN---FQNMWRNIGAYED----GSVRDMITLAKYNGGDLDTY--------- 64
+++ + H +N + W+N + ED G ++ +++ G DT+
Sbjct: 48 KIVEMGHWLENKWVWDFSWKNPISGEDVLKLGVMKQIVSSFSLINGKKDTWVWILEGDGQ 227
Query: 65 FEHPDFEPLVEGPEALAQVLGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNL 116
+ L+ G + + FS +W APSN A WR L +I ++ NL
Sbjct: 228 YSVKSAYDLLSGLDTTIGISVFSKLWKACAPSNAVALGWRVFLDRIQTKDNL 383
>AV428268
Length = 429
Score = 30.0 bits (66), Expect = 0.32
Identities = 11/35 (31%), Positives = 21/35 (59%)
Frame = +3
Query: 86 FSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLG 120
F +W PSN A +W+ L+ ++ +++NL + G
Sbjct: 69 FKLLWSVKVPSNAIALSWKVLINRVQTKVNLNRRG 173
>AV414139
Length = 309
Score = 29.6 bits (65), Expect = 0.41
Identities = 12/22 (54%), Positives = 13/22 (58%)
Frame = -2
Query: 86 FSHVWHFAAPSNVKAFAWRFLL 107
+ VW APSN AF WR LL
Sbjct: 68 YEKVWVGIAPSNANAFVWRLLL 3
>AV422333
Length = 482
Score = 29.3 bits (64), Expect = 0.54
Identities = 11/20 (55%), Positives = 18/20 (90%)
Frame = +2
Query: 125 LISFGYNRSKDEFLVVLIKL 144
L FGY++SKD++L+V+I+L
Sbjct: 338 LSGFGYDQSKDDYLLVIIRL 397
>AV777945
Length = 279
Score = 28.5 bits (62), Expect = 0.92
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +1
Query: 86 FSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWK 118
F +W APSN A AW L+ K+ ++ L K
Sbjct: 151 FDFLWASKAPSNSLALAWNVLIKKLQTKDELRK 249
>BP028574
Length = 454
Score = 28.5 bits (62), Expect = 0.92
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +3
Query: 86 FSHVWHFAAPSNVKAFAWRFLLVKIPSRLNL 116
F +W APSN A AWR L ++P L
Sbjct: 360 FRKLWQSKAPSNSIALAWRVGLNRVPCMAEL 452
>BP071638
Length = 406
Score = 27.7 bits (60), Expect = 1.6
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = -3
Query: 87 SHVWHFAAPSNVKAFAWRFLL 107
++V F + SN+K F WRFLL
Sbjct: 149 AYVHSFFSSSNIKRFEWRFLL 87
>BP075826
Length = 327
Score = 27.7 bits (60), Expect = 1.6
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = -2
Query: 87 SHVWHFAAPSNVKAFAWRFLL 107
++V F + SN+K F WRFLL
Sbjct: 107 AYVHSFFSSSNIKRFEWRFLL 45
>BP050190
Length = 564
Score = 26.2 bits (56), Expect = 4.6
Identities = 16/60 (26%), Positives = 24/60 (39%)
Frame = +1
Query: 33 QNMWRNIGAYEDGSVRDMITLAKYNGGDLDTYFEHPDFEPLVEGPEALAQVLGFSHVWHF 92
QN + + +DG D +TLA +G P+ P LA L F W++
Sbjct: 292 QNRYGVVSGLKDGGYSDDMTLAAISGAQKRLITS----PPVAVFPHPLASDLNFGRYWNY 459
>TC17395 similar to UP|Q9LN83 (Q9LN83) T12C24.17, partial (37%)
Length = 570
Score = 25.8 bits (55), Expect = 6.0
Identities = 13/40 (32%), Positives = 22/40 (54%)
Frame = +2
Query: 100 AFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSKDEFLV 139
+F WRF+ ++P + +G +SFG++ S FLV
Sbjct: 137 SFLWRFIPGRLPKHIFSAAVGVLLSYLSFGFS-SNLHFLV 253
>TC14160 homologue to GB|BAB90396.1|20161472|AP003570 ADP-ribosylation factor
{Oryza sativa (japonica cultivar-group);}, partial (48%)
Length = 1105
Score = 25.8 bits (55), Expect = 6.0
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = +3
Query: 100 AFAWRFLLVKIPSRLNLWKL 119
+ WRFLL KIP R L+ L
Sbjct: 1029 SIGWRFLLCKIP*RKTLYSL 1088
>TC10414 similar to GB|AAP13363.1|30023660|BT006255 At5g55710 {Arabidopsis
thaliana;}, partial (34%)
Length = 504
Score = 25.8 bits (55), Expect = 6.0
Identities = 12/40 (30%), Positives = 20/40 (50%)
Frame = -3
Query: 116 LWKLGATRGLISFGYNRSKDEFLVVLIKLCNLLHLAKNKI 155
+W +G ++ FGY D +VV++ C LL K +
Sbjct: 310 VWGVGGV-AVVGFGYYEPLDVVVVVVVGFCELLWKKKRGV 194
>TC19990 weakly similar to UP|AAQ88099 (AAQ88099) NADPH-dependent cinnamyl
alcohol dehydrogenase, partial (26%)
Length = 421
Score = 25.4 bits (54), Expect = 7.8
Identities = 21/53 (39%), Positives = 27/53 (50%)
Frame = +1
Query: 105 FLLVKIPSRLNLWKLGATRGLISFGYNRSKDEFLVVLIKLCNLLHLAKNKITI 157
FLL K SR LWK A R L +F S+DE + I L+H+ K I +
Sbjct: 205 FLL*K*VSR-KLWKASAKRKLSTFNL*YSRDE*IHDNIS**LLIHVTKLSIIL 360
>TC19400
Length = 519
Score = 25.4 bits (54), Expect = 7.8
Identities = 17/42 (40%), Positives = 25/42 (59%), Gaps = 1/42 (2%)
Frame = -3
Query: 125 LISFGYNRSKDEFLVVLIKLCNLLHLAK-NKITIISLRNYVR 165
LIS YN+ + L+ + C LL LAK N++ +SLR + R
Sbjct: 430 LISINYNKYGTQMLLFFVHAC-LL*LAKPNQLLHVSLRFFGR 308
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.144 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,050,811
Number of Sequences: 28460
Number of extensions: 56456
Number of successful extensions: 352
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 351
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 352
length of query: 214
length of database: 4,897,600
effective HSP length: 86
effective length of query: 128
effective length of database: 2,450,040
effective search space: 313605120
effective search space used: 313605120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0045.6


