

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
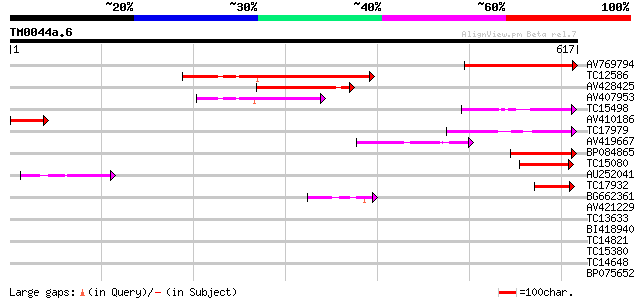
Query= TM0044a.6
(617 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV769794 252 1e-67
TC12586 weakly similar to PIR|C86410|C86410 protein F3M18.18 [im... 180 5e-46
AV428425 108 3e-24
AV407953 107 6e-24
TC15498 weakly similar to PIR|C86410|C86410 protein F3M18.18 [im... 85 3e-17
AV410186 83 1e-16
TC17979 83 1e-16
AV419667 81 4e-16
BP084865 74 7e-14
TC15080 73 2e-13
AU252041 68 4e-12
TC17932 54 7e-08
BG662361 49 2e-06
AV421229 31 0.67
TC13633 30 1.5
BI418940 30 1.5
TC14821 homologue to UP|AAR12195 (AAR12195) Molecular chaperone ... 29 1.9
TC15380 similar to UP|Q8H5P5 (Q8H5P5) OJ1003_C06.27 protein, par... 28 3.3
TC14648 28 3.3
BP075652 28 4.3
>AV769794
Length = 559
Score = 252 bits (643), Expect = 1e-67
Identities = 122/122 (100%), Positives = 122/122 (100%)
Frame = -2
Query: 496 HLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNS 555
HLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNS
Sbjct: 558 HLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNS 379
Query: 556 GIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALLD 615
GIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALLD
Sbjct: 378 GIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALLD 199
Query: 616 YW 617
YW
Sbjct: 198 YW 193
>TC12586 weakly similar to PIR|C86410|C86410 protein F3M18.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(17%)
Length = 622
Score = 180 bits (457), Expect = 5e-46
Identities = 99/214 (46%), Positives = 136/214 (63%), Gaps = 5/214 (2%)
Frame = +1
Query: 189 VGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNR 248
+GG D+I KQ++SS + P ++R+ L+D+ D F Q A DGK E F
Sbjct: 4 LGGTDIIYAKQQYSSPVQPVEVRKLLKDMADNNFLGQAG----QHNAHDGKSNTKEKF-- 165
Query: 249 VMQSNTMSFTSISETSSKDG----LTIICSKRGGDMLKH-SHSSWLQTVPSNPEAILFKF 303
M N + F I S + + +C ++GG++ K +H+ W QTV S P+ I F
Sbjct: 166 -MMDNGLDFMGIQPRSYYEAEVQDIKFMCKRKGGNLKKSMTHNDWCQTVLSQPDLIAMSF 342
Query: 304 VPISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRR 363
+PI+SLL G+ GSGYL+HAINLYLRYKPP E+L FLEFQ+PRQWAP+F EL L +R+
Sbjct: 343 IPITSLLGGVNGSGYLTHAINLYLRYKPPIEELHQFLEFQLPRQWAPVFGELAL-GPERK 519
Query: 364 KSSSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLR 397
++ SLQF+FLGPKL+V++T V +KPV GLR
Sbjct: 520 SQNTASLQFSFLGPKLYVNTTPVDVGKKPVTGLR 621
>AV428425
Length = 422
Score = 108 bits (270), Expect = 3e-24
Identities = 55/107 (51%), Positives = 72/107 (66%)
Frame = +3
Query: 269 LTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLR 328
+T+I +RGGD L+ SH+ W++TV P+ I KF PI+SLL G+PG +L+ AI LYL
Sbjct: 117 VTVIFRRRGGDDLEQSHTKWVETVKLAPDIINMKFTPIASLLEGVPGVKHLARAIELYLE 296
Query: 329 YKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQFNFL 375
YKPP EDLQYFL+FQI R WAP L QR++ SLQF+ +
Sbjct: 297 YKPPIEDLQYFLDFQITRVWAPEQNNL-----QRKEPVCPSLQFSLM 422
>AV407953
Length = 416
Score = 107 bits (267), Expect = 6e-24
Identities = 61/144 (42%), Positives = 86/144 (59%), Gaps = 4/144 (2%)
Frame = +1
Query: 204 KIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVMQSNTMSFTSISET 263
++ P D+++ L+++ D F D Q A + + F ++ ++F +IS +
Sbjct: 1 QLQPADVQKRLKEMADRRFMDANG----QYSVASDQVFPNDKFG--VREQRLTFANISPS 162
Query: 264 SS---KDGLTIICSKRGG-DMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYL 319
SS K+ + I +RGG D SH WLQTV P+ I F+PI+SLL G+PGSG+L
Sbjct: 163 SSYSHKEDIVSIYKRRGGSDDRNLSHHEWLQTVQVEPDVISMSFIPITSLLNGVPGSGFL 342
Query: 320 SHAINLYLRYKPPAEDLQYFLEFQ 343
SHAINLYLRYKPP E+L FLEFQ
Sbjct: 343 SHAINLYLRYKPPIEELHQFLEFQ 414
>TC15498 weakly similar to PIR|C86410|C86410 protein F3M18.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(7%)
Length = 515
Score = 85.1 bits (209), Expect = 3e-17
Identities = 48/125 (38%), Positives = 74/125 (58%)
Frame = +3
Query: 492 KNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPA 551
+NVLHL+LLF+ +P C+IR++ W P ++ ++ +++ S T++ TS K+ +
Sbjct: 18 QNVLHLKLLFSKVPGCTIRRSVWDHNPSTPATAAQRSDGASS-SLTKK--TSDDKKEDSS 188
Query: 552 MLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKF 611
+ GKL K V+ E+ RGP D PGHWLVT AKL E GKI L++K+
Sbjct: 189 --------------IHIGKLAKIVDMTEMFRGPQDIPGHWLVTGAKLGVEKGKIVLRIKY 326
Query: 612 ALLDY 616
+LL+Y
Sbjct: 327 SLLNY 341
>AV410186
Length = 433
Score = 83.2 bits (204), Expect = 1e-16
Identities = 42/42 (100%), Positives = 42/42 (100%)
Frame = +2
Query: 1 MAVEEEGIEMVALESLGKGFDLASDFRLRFAKGIREGRGSSS 42
MAVEEEGIEMVALESLGKGFDLASDFRLRFAKGIREGRGSSS
Sbjct: 308 MAVEEEGIEMVALESLGKGFDLASDFRLRFAKGIREGRGSSS 433
>TC17979
Length = 777
Score = 83.2 bits (204), Expect = 1e-16
Identities = 48/141 (34%), Positives = 74/141 (52%)
Frame = +3
Query: 476 VYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFS 535
V +VTGAQL +NVL+++LL++ +P C+IR++ W P + + N S
Sbjct: 3 VCVVTGAQLGVWDFGSRNVLYMKLLYSRLPGCTIRRSLWDHIPNQPLPTVSVGNTSN--- 173
Query: 536 FTQQSATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTA 595
P ++ P +L+KYV+ +E+ +GP D PGHWLVT
Sbjct: 174 ----------PDESSISFRENTTP--------GNELVKYVDLSEMSKGPQDPPGHWLVTG 299
Query: 596 AKLVTEGGKIGLQVKFALLDY 616
KL E GKI L++K++LL+Y
Sbjct: 300 GKLGVEKGKIVLRLKYSLLNY 362
>AV419667
Length = 406
Score = 81.3 bits (199), Expect = 4e-16
Identities = 54/127 (42%), Positives = 74/127 (57%)
Frame = +1
Query: 378 KLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETSTPSMWRGS 437
+L ++ QV +KPV GLRLYLEG+K + LA+H+ HLSSLP L E + G+
Sbjct: 46 RLTLNYLQVDVGKKPVTGLRLYLEGKKNNCLAIHLQHLSSLPKTFQLMDEPN------GN 207
Query: 438 DDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSWPKNVLHL 497
+ S ++ E V+WK FS+VCTA V+ D NS +VTGA+ S K VL L
Sbjct: 208 VPHSSERKYYEKVQWKSFSHVCTAPVESDGE--NS-----VVTGARF*VGESGLKKVLFL 366
Query: 498 RLLFTNI 504
RL F+ +
Sbjct: 367 RLHFSKV 387
>BP084865
Length = 484
Score = 73.9 bits (180), Expect = 7e-14
Identities = 31/71 (43%), Positives = 47/71 (65%)
Frame = -3
Query: 546 PKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKI 605
PK + S YP GPP V+ LL++V+ AE++RGP TPG+W+V+ A+ + GKI
Sbjct: 392 PKGGEVTIGSAAYPTGPPSRVQTPSLLRFVDTAEIIRGPEHTPGYWVVSGARFSVQNGKI 213
Query: 606 GLQVKFALLDY 616
L VK++LL++
Sbjct: 212 YLSVKYSLLNF 180
>TC15080
Length = 716
Score = 72.8 bits (177), Expect = 2e-13
Identities = 30/59 (50%), Positives = 45/59 (75%)
Frame = +1
Query: 555 SGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFAL 613
S +YP GPPV ++ KLL++V+ E+ RGP D+PG+W+V+ A+L E GKI L+VK++L
Sbjct: 1 SALYPGGPPVXAQSTKLLRFVDTTEMTRGPQDSPGYWVVSGARLYVEKGKISLKVKYSL 177
>AU252041
Length = 326
Score = 68.2 bits (165), Expect = 4e-12
Identities = 38/104 (36%), Positives = 61/104 (58%)
Frame = +3
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
++++LG+GFD+ SD RL + KG + RLV DE++ RD+ I + GV + V
Sbjct: 45 SIQALGRGFDVTSDIRLLYCKG--------APGSRLVRFDEEHTRDLAI--SHGVVVPNV 194
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFN 115
S +I C G K+ V F++M++ N++S + G+IP G FN
Sbjct: 195 SGDIVCSLGKSALEKTPVCSFHEMAKHFNERSGITGQIPLGSFN 326
>TC17932
Length = 572
Score = 53.9 bits (128), Expect = 7e-08
Identities = 21/43 (48%), Positives = 34/43 (78%)
Frame = +1
Query: 572 LKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALL 614
LK+V+ E+ RGP + PG+W+V+ A+LV E G++ L+VK++LL
Sbjct: 1 LKFVDTTEMSRGPQENPGYWVVSGARLVVEKGRVSLRVKYSLL 129
>BG662361
Length = 330
Score = 49.3 bits (116), Expect = 2e-06
Identities = 34/84 (40%), Positives = 44/84 (51%), Gaps = 8/84 (9%)
Frame = +1
Query: 325 LYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSST 384
LY KPP EDLQYFL+FQI R WAP +++ + K + LS+ P + + T
Sbjct: 115 LYEPDKPPIEDLQYFLDFQITRVWAPE------QNNLQTKGACLSI------PSVQLDGT 258
Query: 385 --------QVLSEQKPVVGLRLYL 400
QV +KPV GLRL L
Sbjct: 259 *AFCSVPDQVTVGRKPVTGLRLSL 330
>AV421229
Length = 262
Score = 30.8 bits (68), Expect = 0.67
Identities = 16/45 (35%), Positives = 24/45 (52%), Gaps = 4/45 (8%)
Frame = +3
Query: 550 PAMLNSGIYPDGPPVPVRAGKLLKYV----EAAEVVRGPHDTPGH 590
P++ N I P+ P +R +L+K EA +RGP +TP H
Sbjct: 39 PSLTNLRIRPNNPEHHLRPSRLIKLPCGGPEANRSIRGPRETPPH 173
>TC13633
Length = 706
Score = 29.6 bits (65), Expect = 1.5
Identities = 13/48 (27%), Positives = 24/48 (49%)
Frame = +2
Query: 457 NVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNI 504
N+C H PN++ Y+ + L+ WP+ +HL+ L +N+
Sbjct: 386 NICIVNQGHTPNYIRVGASEYLQSS--LIILLPWPRENIHLQ*LKSNV 523
>BI418940
Length = 550
Score = 29.6 bits (65), Expect = 1.5
Identities = 21/59 (35%), Positives = 28/59 (46%), Gaps = 2/59 (3%)
Frame = +3
Query: 515 AAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNSGIYPDGPPVPV--RAGKL 571
A P + SS L S F T +A+ PP APA + P PP+P+ +GKL
Sbjct: 243 ATTPTPTNPSSSL*EKSKHFCSTTSAASYPPPTPAPATTQT---PTTPPLPLSENSGKL 410
>TC14821 homologue to UP|AAR12195 (AAR12195) Molecular chaperone Hsp90-1,
partial (29%)
Length = 612
Score = 29.3 bits (64), Expect = 1.9
Identities = 25/90 (27%), Positives = 40/90 (43%), Gaps = 6/90 (6%)
Frame = -2
Query: 213 HLEDLGDFLFSDVR---SPPLLQRKAADGKQ---KVPEVFNRVMQSNTMSFTSISETSSK 266
H ED D F VR ++R + K+ K+ FNR + ++ M F I +T +
Sbjct: 440 HDEDTADIQFDIVRLLLCVKEIKRCSLGYKEDSLKLQLTFNREVLNSQMLFPIIGKTLVE 261
Query: 267 DGLTIICSKRGGDMLKHSHSSWLQTVPSNP 296
+ I C ++L+ H +WL V P
Sbjct: 260 SSILIFC-----NLLRLPHPNWLLFVHQRP 186
>TC15380 similar to UP|Q8H5P5 (Q8H5P5) OJ1003_C06.27 protein, partial (28%)
Length = 1306
Score = 28.5 bits (62), Expect = 3.3
Identities = 18/56 (32%), Positives = 25/56 (44%), Gaps = 4/56 (7%)
Frame = +2
Query: 516 AAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQA----PAMLNSGIYPDGPPVPVR 567
AAP A ++ T+ + F T +T+ PP + PA S P PP P R
Sbjct: 158 AAPTAPSTAASSTSSTARFPQTPNPSTASPPPTSPWTPPATSGSASLPPPPPPPPR 325
>TC14648
Length = 669
Score = 28.5 bits (62), Expect = 3.3
Identities = 25/93 (26%), Positives = 38/93 (39%)
Frame = -2
Query: 276 RGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPAED 335
RGG SH W+ VPS EA L + + SL+ + L H ++L L + +D
Sbjct: 269 RGGVEQVLSHFHWVILVPSLVEAQLLPHILVPSLVE----AQLLPHILSLLLHWVSSPQD 102
Query: 336 LQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSL 368
P++ P P + S+SL
Sbjct: 101 PSL---LSYPQEEDPSLLSYPQEENNSLLSTSL 12
>BP075652
Length = 363
Score = 28.1 bits (61), Expect = 4.3
Identities = 13/38 (34%), Positives = 23/38 (60%)
Frame = +1
Query: 253 NTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQ 290
N + T++ + S+DG T RGG +++ +H+S LQ
Sbjct: 127 NLIYSTTLPSSPSRDGCT*SSGGRGGRIIREAHTSNLQ 240
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,539,415
Number of Sequences: 28460
Number of extensions: 181541
Number of successful extensions: 958
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 943
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 949
length of query: 617
length of database: 4,897,600
effective HSP length: 96
effective length of query: 521
effective length of database: 2,165,440
effective search space: 1128194240
effective search space used: 1128194240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0044a.6


