

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0042.13
(250 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
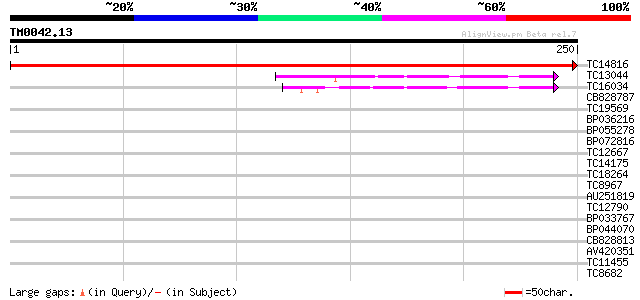
Score E
Sequences producing significant alignments: (bits) Value
TC14816 496 e-141
TC13044 similar to UP|FKB7_WHEAT (Q43207) 70 kDa peptidylprolyl ... 48 1e-06
TC16034 homologue to UP|FKB2_VICFA (Q41649) FK506-binding protei... 40 3e-04
CB828787 39 0.001
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 38 0.001
BP036216 38 0.002
BP055278 36 0.005
BP072816 36 0.005
TC12667 similar to PIR|D96525|D96525 protein T1N15.19 [imported]... 36 0.005
TC14175 similar to UP|Q9SII5 (Q9SII5) At2g17230 protein (At2g172... 36 0.007
TC18264 similar to UP|Q9SP93 (Q9SP93) Thiol protease, partial (4%) 35 0.009
TC8967 similar to UP|FXGA_HUMAN (P55316) Forkhead box protein G1... 35 0.012
AU251819 35 0.012
TC12790 similar to PIR|H96696|H96696 protein F1N21.16 [imported]... 35 0.012
BP033767 35 0.012
BP044070 35 0.016
CB828813 34 0.021
AV420351 34 0.021
TC11455 similar to UP|Q9ZUS8 (Q9ZUS8) At2g37380 protein, partial... 34 0.021
TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate... 34 0.021
>TC14816
Length = 952
Score = 496 bits (1276), Expect = e-141
Identities = 249/250 (99%), Positives = 249/250 (99%)
Frame = +2
Query: 1 MATFFGSPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPI 60
MATFFGSPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPI
Sbjct: 29 MATFFGSPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPI 208
Query: 61 KPVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDV 120
KPVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDV
Sbjct: 209 KPVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDV 388
Query: 121 EKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALV 180
EKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALV
Sbjct: 389 EKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALV 568
Query: 181 MGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIV 240
MGSRPY KGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIV
Sbjct: 569 MGSRPYRKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIV 748
Query: 241 EVDKVSIAPA 250
EVDKVSIAPA
Sbjct: 749 EVDKVSIAPA 778
>TC13044 similar to UP|FKB7_WHEAT (Q43207) 70 kDa peptidylprolyl isomerase
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase) , partial (26%)
Length = 531
Score = 48.1 bits (113), Expect = 1e-06
Identities = 43/126 (34%), Positives = 61/126 (48%), Gaps = 1/126 (0%)
Frame = +2
Query: 118 RDVEKEEEVTLPNGIRYCELKVGGG-ASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKP 176
+D EE +G+R LK G G +P GD V + G + G F ++ + D P
Sbjct: 86 KDKVGEEREIGNSGLRKKLLKEGQGWETPEVGDEVQVHYTGTLLD-GSKFDSSRDRDA-P 259
Query: 177 LALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATL 236
+ +G KG EGI KTMK G +PP+L + E+ GS IPP ATL
Sbjct: 260 FSFTLGQGQVIKGWDEGI----KTMKKGENALFTIPPELAYGES----GSPPTIPPNATL 415
Query: 237 EYIVEV 242
++ VE+
Sbjct: 416 QFDVEL 433
>TC16034 homologue to UP|FKB2_VICFA (Q41649) FK506-binding protein 2
precursor (Peptidyl-prolyl cis-trans isomerase)
(PPIase) (Rotamase) (15 kDa FKBP) (FKBP-15) , partial
(91%)
Length = 592
Score = 40.4 bits (93), Expect = 3e-04
Identities = 42/127 (33%), Positives = 63/127 (49%), Gaps = 5/127 (3%)
Frame = +3
Query: 121 EKEEEVT-LPNGIRY----CELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKK 175
+K +VT L G++Y CE++ GD V + GK+ + G VF ++FE +
Sbjct: 90 KKTADVTELQIGVKYKPTSCEVQA------HKGDKVKVHYRGKL-TDGTVFDSSFERNS- 245
Query: 176 PLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLAT 235
P+ +GS KG +G L M G KRK+ +P +LG+ E GS IP AT
Sbjct: 246 PIDFELGSGQVIKGWDQG----LLGMCLGEKRKLKIPAKLGYGEQ----GSPPTIPGGAT 401
Query: 236 LEYIVEV 242
L + E+
Sbjct: 402 LVFDTEL 422
>CB828787
Length = 343
Score = 38.5 bits (88), Expect = 0.001
Identities = 19/49 (38%), Positives = 32/49 (64%)
Frame = +1
Query: 197 VLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
+++ M+ GGKR V+VPP+ G+ + G + +IPP AT E +E+ +V
Sbjct: 202 IVEGMRVGGKRTVLVPPENGYGKRGMN-----EIPPGATFEMNIELLQV 333
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 38.1 bits (87), Expect = 0.001
Identities = 24/73 (32%), Positives = 32/73 (42%), Gaps = 2/73 (2%)
Frame = +1
Query: 7 SPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQ--GTAKAQPQNPIKPVT 64
SPP S T+ H +S P P P ++ PTA +S S K+ P P P
Sbjct: 232 SPPNPSTSTTQQPKHSPTSPPASPSPSPPSTKPPTAMASPSTSPLSATKSHPTPPAAPSA 411
Query: 65 SSTKVDSTDWIAT 77
SST +T + T
Sbjct: 412 SSTPPPTTTSLKT 450
Score = 33.5 bits (75), Expect = 0.036
Identities = 21/66 (31%), Positives = 36/66 (53%), Gaps = 2/66 (3%)
Frame = +1
Query: 5 FGSPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQS-PTASSSQSVQGTAKAQPQ-NPIKP 62
+ PP S T + +S+ TPPP P + +++ PTA S+ + T+ A + +P P
Sbjct: 67 YSPPPPPSQQPTHSPSTSPTSTTTPPPSPTKVTENPPTAPSTSTKSPTSSASEEPSPPNP 246
Query: 63 VTSSTK 68
TS+T+
Sbjct: 247 STSTTQ 264
Score = 30.0 bits (66), Expect = 0.39
Identities = 20/54 (37%), Positives = 25/54 (46%), Gaps = 6/54 (11%)
Frame = +1
Query: 21 HIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQP------QNPIKPVTSSTK 68
H +SSS + PP SQ PT S S S T P +NP ++STK
Sbjct: 40 HTSSSSSSSYSPPPPPSQQPTHSPSTSPTSTTTPPPSPTKVTENPPTAPSTSTK 201
Score = 26.2 bits (56), Expect = 5.7
Identities = 9/15 (60%), Positives = 10/15 (66%)
Frame = +2
Query: 20 HHIASSSPTPPPIPP 34
HH+ SS PPP PP
Sbjct: 20 HHVGCSSTLPPPPPP 64
>BP036216
Length = 574
Score = 37.7 bits (86), Expect = 0.002
Identities = 38/123 (30%), Positives = 55/123 (43%), Gaps = 1/123 (0%)
Frame = +1
Query: 121 EKEEEVTLPNGIRYCELKVGGGA-SPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLAL 179
EKE + G++ +K G G +P GD V + G + + + G P
Sbjct: 175 EKEXGI---QGLKKKLIKEGEGXDTPDAGDEVHVHYTGTLLDGTKFDSSRDRGT--PFNF 339
Query: 180 VMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYI 239
+G KG EGI KTMK G +P +L + E+G S IPP ATL++
Sbjct: 340 TLGQGQVIKGWDEGI----KTMKKGENALFTIPAELAYGESG----SPPTIPPNATLQFD 495
Query: 240 VEV 242
VE+
Sbjct: 496 VEL 504
>BP055278
Length = 449
Score = 36.2 bits (82), Expect = 0.005
Identities = 13/31 (41%), Positives = 23/31 (73%)
Frame = -2
Query: 193 GIEYVLKTMKAGGKRKVIVPPQLGFRENGAD 223
GI++ +++MK GG R+VI+PP LG++ +
Sbjct: 445 GIDFHVRSMKVGGIRRVIIPPSLGYQNTSQE 353
>BP072816
Length = 492
Score = 36.2 bits (82), Expect = 0.005
Identities = 22/72 (30%), Positives = 34/72 (46%)
Frame = +1
Query: 7 SPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSS 66
+PP+ P +S +SSS +P P PP +P+ S S + TA P P +S
Sbjct: 82 NPPLSHQPC--HSPSSSSSSSSPLPSPPTTLHAPSTSPSSAPSATAAPPPSTPTPAANTS 255
Query: 67 TKVDSTDWIATS 78
+K ++ TS
Sbjct: 256 SKPSTSSKPTTS 291
Score = 26.2 bits (56), Expect = 5.7
Identities = 22/82 (26%), Positives = 33/82 (39%), Gaps = 14/82 (17%)
Frame = +1
Query: 13 LPLTRNSHHIASSSPT--------PPPIPPQQSQSPTASSSQSVQGTAKAQP------QN 58
LP + H S+SP+ PPP P + + ++ S S + T A P
Sbjct: 145 LPSPPTTLHAPSTSPSSAPSATAAPPPSTPTPAANTSSKPSTSSKPTTSAAPPASSLLST 324
Query: 59 PIKPVTSSTKVDSTDWIATSLT 80
P KP + ST+ A + T
Sbjct: 325 PPKPAGPPSNPSSTNSPAPAST 390
>TC12667 similar to PIR|D96525|D96525 protein T1N15.19 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(10%)
Length = 498
Score = 36.2 bits (82), Expect = 0.005
Identities = 21/69 (30%), Positives = 31/69 (44%), Gaps = 14/69 (20%)
Frame = +2
Query: 18 NSHHIASS----SPTPPPIPPQQSQSPTASSSQSVQG----------TAKAQPQNPIKPV 63
+SHH S+ SP PPP PP++S P + S+ G T K NP+ P+
Sbjct: 212 HSHHSTSTAALASPPPPPPPPKKSHPPPPPPTNSIPGPNGSPSSTA*TPKVTSPNPVLPM 391
Query: 64 TSSTKVDST 72
T ++
Sbjct: 392 ALFTSTSTS 418
>TC14175 similar to UP|Q9SII5 (Q9SII5) At2g17230 protein
(At2g17230/T23A1.9), partial (82%)
Length = 1447
Score = 35.8 bits (81), Expect = 0.007
Identities = 16/42 (38%), Positives = 21/42 (49%)
Frame = +2
Query: 18 NSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNP 59
N HH+ ++ P PPP PP + S+ Q K QPQ P
Sbjct: 68 NHHHVTATQPPPPPPPPLLRHN*WRGSNTEPQPPFKIQPQAP 193
Score = 32.0 bits (71), Expect = 0.10
Identities = 22/78 (28%), Positives = 36/78 (45%), Gaps = 8/78 (10%)
Frame = +1
Query: 21 HIASSSPTPPPIPPQ---QSQSPTASSSQSVQGTAKAQPQNPIKPVTSS-----TKVDST 72
H SSSP+ PPQ + + T+++ Q+ ++ + P P P S + ST
Sbjct: 88 HSTSSSPSSSSSPPQLMERFKH*TSTTLQNSTPSSHSHPSEPSPPQNGSKAPQTSLSSST 267
Query: 73 DWIATSLTRRFGLGAGLA 90
W+ +SL R +G A
Sbjct: 268 TWVLSSLPRSTSTSSGTA 321
>TC18264 similar to UP|Q9SP93 (Q9SP93) Thiol protease, partial (4%)
Length = 570
Score = 35.4 bits (80), Expect = 0.009
Identities = 26/67 (38%), Positives = 29/67 (42%), Gaps = 10/67 (14%)
Frame = -1
Query: 27 PTPPPIPPQQSQSPTASSSQSVQGTAKAQP---------QNPIKPVTSSTKVDSTDWIAT 77
P PPP PP P + SQ V G + P +NP K TS TK I T
Sbjct: 510 PPPPPPPPPPPSPPHSPPSQIVSGPGSSPPMKPPKAPFLKNPKKSRTSITKSSKR*LINT 331
Query: 78 S-LTRRF 83
S TRRF
Sbjct: 330 SRATRRF 310
>TC8967 similar to UP|FXGA_HUMAN (P55316) Forkhead box protein G1A
(Forkhead-related protein FKHL2) (Transcription factor
BF-2) (Brain factor 2) (BF2) (HFK2), partial (6%)
Length = 1032
Score = 35.0 bits (79), Expect = 0.012
Identities = 21/47 (44%), Positives = 27/47 (56%), Gaps = 1/47 (2%)
Frame = +2
Query: 27 PTPPPIPPQQSQSPTASSSQSVQGTAKAQPQN-PIKPVTSSTKVDST 72
P PPP PP +SPTAS + S T+KA+ + P T S+K ST
Sbjct: 146 PPPPPPPPPPRRSPTASCA-SPPSTSKARSTT*LVSPATPSSKPSST 283
>AU251819
Length = 368
Score = 35.0 bits (79), Expect = 0.012
Identities = 29/72 (40%), Positives = 39/72 (53%), Gaps = 5/72 (6%)
Frame = +1
Query: 7 SPPVFSLPLTRNSHHI--ASSSPTPPPIPPQQSQSPTASSS--QSVQGTAK-AQPQNPIK 61
SPP SL + ++SH I + SS + PP P S S + SSS SV G+ K Q + P+
Sbjct: 106 SPP--SLKINKDSHSIKKSPSSSSSPPPPSLSSSSSSISSSLVNSVAGSNKPPQQRQPVI 279
Query: 62 PVTSSTKVDSTD 73
T S +V TD
Sbjct: 280 IYTHSPRVIHTD 315
>TC12790 similar to PIR|H96696|H96696 protein F1N21.16 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 727
Score = 35.0 bits (79), Expect = 0.012
Identities = 18/45 (40%), Positives = 26/45 (57%)
Frame = -1
Query: 7 SPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGT 51
S P +LP + ++SP PP PP++S S TA SS+ V+ T
Sbjct: 271 SAPHLALPAPTQTTP*ETNSPDAPPAPPRRSHSSTAPSSRRVKPT 137
>BP033767
Length = 541
Score = 35.0 bits (79), Expect = 0.012
Identities = 22/59 (37%), Positives = 26/59 (43%)
Frame = +1
Query: 8 PPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSS 66
PP S P T HH+ PPP PP SP +SS +P NP P +SS
Sbjct: 280 PPHLSSPHTHLHHHLL----LPPPSPPSPPTSPPSSS---------LKPPNPNPPTSSS 417
Score = 25.8 bits (55), Expect = 7.4
Identities = 18/71 (25%), Positives = 27/71 (37%)
Frame = +3
Query: 7 SPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSS 66
S P+ + + S P+ PP PP S SP+ P NP P +S+
Sbjct: 117 SEPIINRRILHQPFTQGSLPPSQPPNPPPPSPSPS--------------PPNPKYPFSST 254
Query: 67 TKVDSTDWIAT 77
+T +T
Sbjct: 255 PTTTTTQNAST 287
>BP044070
Length = 289
Score = 34.7 bits (78), Expect = 0.016
Identities = 28/77 (36%), Positives = 32/77 (41%), Gaps = 6/77 (7%)
Frame = +3
Query: 2 ATFFGSPPVFSLPLTRNSHHIASSSPTPPPI---PPQQSQSPTASSS---QSVQGTAKAQ 55
AT SPP NS A SSP PPP P S PT S S S + T A
Sbjct: 66 ATSAPSPP-------SNSPPTAVSSPPPPPTRPSAPMASPPPTVSPSPRCSSTRATTXAS 224
Query: 56 PQNPIKPVTSSTKVDST 72
P +P P +S+ T
Sbjct: 225 PTSPSPPTPASSSPPPT 275
>CB828813
Length = 465
Score = 34.3 bits (77), Expect = 0.021
Identities = 21/57 (36%), Positives = 27/57 (46%), Gaps = 5/57 (8%)
Frame = +1
Query: 8 PPVFSLPLTRNSHHIASSSP-----TPPPIPPQQSQSPTASSSQSVQGTAKAQPQNP 59
PP + P+ H I S++P TPPP PP +PT +SS S P NP
Sbjct: 154 PPNLTPPILTKPHPIGSTTPFSALTTPPPPPP--PPTPTRTSSTSKPPIPTPPPPNP 318
>AV420351
Length = 188
Score = 34.3 bits (77), Expect = 0.021
Identities = 15/44 (34%), Positives = 22/44 (49%)
Frame = +3
Query: 20 HHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPV 63
H I P PPP PPQ +P++ ++S T P P+ P+
Sbjct: 54 HFIPPPPPPPPPQPPQPYSAPSSPLNRSTTRTLLPSPLLPLNPL 185
>TC11455 similar to UP|Q9ZUS8 (Q9ZUS8) At2g37380 protein, partial (10%)
Length = 575
Score = 34.3 bits (77), Expect = 0.021
Identities = 19/62 (30%), Positives = 29/62 (46%), Gaps = 5/62 (8%)
Frame = +2
Query: 24 SSSPTPPPIPPQQS-----QSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDSTDWIATS 78
SSSP PPP PP ++P A++ S + + P P S S+D+ +T+
Sbjct: 284 SSSPPPPPHPPPHPPPPPLETPLAAAHPSTATSPSSPTATPPAPAPSQRTTSSSDFTSTT 463
Query: 79 LT 80
T
Sbjct: 464 AT 469
Score = 26.9 bits (58), Expect = 3.3
Identities = 17/63 (26%), Positives = 24/63 (37%), Gaps = 3/63 (4%)
Frame = +2
Query: 8 PPVFSLPLTRNSHHIASSSPTP---PPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVT 64
PP PL A+S +P PP P ++ ++S S T NP
Sbjct: 329 PPPLETPLAAAHPSTATSPSSPTATPPAPAPSQRTTSSSDFTSTTATTTTHYSNPTSRKL 508
Query: 65 SST 67
+ST
Sbjct: 509 TST 517
>TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate
dioxygenase (4HPPD) (HPD) (HPPDase) , partial (29%)
Length = 576
Score = 34.3 bits (77), Expect = 0.021
Identities = 23/70 (32%), Positives = 31/70 (43%), Gaps = 9/70 (12%)
Frame = +1
Query: 7 SPPVFSLPLTRNSH---HIASSSPTPPPIP------PQQSQSPTASSSQSVQGTAKAQPQ 57
+PP S P T S H SP+PPP P P S SP ++S S +K+
Sbjct: 367 TPPTSSAPATSASSSPPHTLPKSPSPPPPPSPLSPPPPASPSPPPTASPSAPSPSKSPTP 546
Query: 58 NPIKPVTSST 67
+ P S+T
Sbjct: 547 SKPSPPQSTT 576
Score = 29.3 bits (64), Expect = 0.67
Identities = 22/68 (32%), Positives = 28/68 (40%), Gaps = 8/68 (11%)
Frame = +1
Query: 7 SPPVFSLPLTRNSHHIASSSPTPP--------PIPPQQSQSPTASSSQSVQGTAKAQPQN 58
SPP +L +S S+S +PP P PP SP +S S TA +
Sbjct: 346 SPPETTLTPPTSSAPATSASSSPPHTLPKSPSPPPPPSPLSPPPPASPSPPPTASPSAPS 525
Query: 59 PIKPVTSS 66
P K T S
Sbjct: 526 PSKSPTPS 549
Score = 26.6 bits (57), Expect = 4.3
Identities = 18/48 (37%), Positives = 24/48 (49%)
Frame = +1
Query: 24 SSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDS 71
SSS P P P + SP AS+ +S Q + P+ + P TSS S
Sbjct: 262 SSSGAPTP-PTPHAASPGASACRSSQNQT-SPPETTLTPPTSSAPATS 399
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,202,746
Number of Sequences: 28460
Number of extensions: 72095
Number of successful extensions: 2452
Number of sequences better than 10.0: 497
Number of HSP's better than 10.0 without gapping: 1728
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2166
length of query: 250
length of database: 4,897,600
effective HSP length: 88
effective length of query: 162
effective length of database: 2,393,120
effective search space: 387685440
effective search space used: 387685440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0042.13


