

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0041.8
(562 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
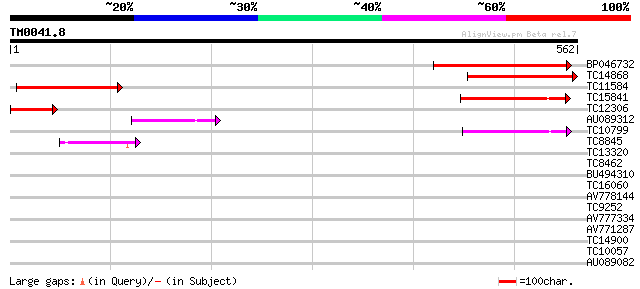
BP046732 274 3e-74
TC14868 similar to UP|Q8H1Z0 (Q8H1Z0) Cuticle protein, partial (... 235 1e-62
TC11584 similar to UP|O23679 (O23679) CER1 protein, partial (30%) 105 3e-23
TC15841 similar to UP|O80938 (O80938) CER1-like protein, partial... 97 7e-21
TC12306 weakly similar to UP|AAR90847 (AAR90847) Glossy1 protein... 91 6e-19
AU089312 77 1e-14
TC10799 weakly similar to UP|O23678 (O23678) CER1-like protein, ... 65 3e-11
TC8845 similar to UP|Q8VWZ8 (Q8VWZ8) Sterol 4-alpha-methyl-oxida... 52 3e-07
TC13320 similar to UP|O80456 (O80456) At2g23390 protein (At2g233... 30 1.0
TC8462 similar to UP|PSAN_ARATH (P49107) Photosystem I reaction ... 30 1.4
BU494310 30 1.4
TC16060 weakly similar to UP|Q9SHK4 (Q9SHK4) F12K11.6, partial (8%) 29 1.8
AV778144 29 2.3
TC9252 similar to UP|Q93X62 (Q93X62) Beta-oxyacyl-[acyl-carrier ... 29 2.3
AV777334 29 2.3
AV771287 28 3.9
TC14900 similar to GB|AAL36081.1|17380636|AY063731 AT3g49800/T16... 28 5.1
TC10057 similar to UP|Q41124 (Q41124) Chloroplast RNA binding pr... 28 5.1
AU089082 27 6.7
>BP046732
Length = 495
Score = 274 bits (700), Expect = 3e-74
Identities = 127/137 (92%), Positives = 132/137 (95%)
Frame = -1
Query: 421 IQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFR 480
IQKEAP EYQSYLVQVTKYQAAQ CKTWIVGKWITPREQSWAPRGTHFHQFVVPPIL FR
Sbjct: 495 IQKEAPLEYQSYLVQVTKYQAAQTCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILSFR 316
Query: 481 RDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRI 540
R+CTYG LAAMRLP+DVEGLG CEYTMERGVVHACHAGGVVHSLEGW+HHEVGAIDV+RI
Sbjct: 315 RNCTYGGLAAMRLPQDVEGLGCCEYTMERGVVHACHAGGVVHSLEGWSHHEVGAIDVNRI 136
Query: 541 DVVWKAALKHGLRPVSS 557
D+VWKAALKHGLRPVSS
Sbjct: 135 DLVWKAALKHGLRPVSS 85
>TC14868 similar to UP|Q8H1Z0 (Q8H1Z0) Cuticle protein, partial (16%)
Length = 570
Score = 235 bits (600), Expect = 1e-62
Identities = 109/109 (100%), Positives = 109/109 (100%)
Frame = +3
Query: 454 ITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVH 513
ITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVH
Sbjct: 3 ITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVH 182
Query: 514 ACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWKAALKHGLRPVSSGSTGK 562
ACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWKAALKHGLRPVSSGSTGK
Sbjct: 183 ACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWKAALKHGLRPVSSGSTGK 329
>TC11584 similar to UP|O23679 (O23679) CER1 protein, partial (30%)
Length = 617
Score = 105 bits (261), Expect = 3e-23
Identities = 46/106 (43%), Positives = 66/106 (61%)
Frame = +1
Query: 7 NRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHV 66
N RI+ +G++F Q+D+E DWD+ ++ ++ +A Y LP LP W GV++A++LH
Sbjct: 271 NTRIVDKGIEFDQVDRERDWDDQILFNGLLYYLACYTLPGASRLPLWRTDGVLIAILLHA 450
Query: 67 GISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEH 112
G E LYYW+HR H YL++ YHS HHSS V E +T EH
Sbjct: 451 GPVEFLYYWLHRALHHHYLYSRYHSHHHSSVVTEPITSVIHPFAEH 588
>TC15841 similar to UP|O80938 (O80938) CER1-like protein, partial (14%)
Length = 636
Score = 97.1 bits (240), Expect = 7e-21
Identities = 48/109 (44%), Positives = 66/109 (60%)
Frame = +1
Query: 448 WIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTM 507
W+VG +T EQ AP GT F + P +R+DC+Y AM++P VE + SCE +
Sbjct: 22 WLVGDGLTEEEQVNAPXGTIFIPYSQFPPKKYRKDCSYHCTPAMQIPSSVENIHSCEDWL 201
Query: 508 ERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWKAALKHGLRPVS 556
R V+ A G+VHSLEGWT HE G + ID VWK+ L+HG +P++
Sbjct: 202 PRRVMSAWRIAGIVHSLEGWTEHECG-YTMHNIDKVWKSTLQHGFKPLT 345
>TC12306 weakly similar to UP|AAR90847 (AAR90847) Glossy1 protein, partial
(8%)
Length = 586
Score = 90.5 bits (223), Expect = 6e-19
Identities = 44/47 (93%), Positives = 45/47 (95%)
Frame = +2
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFL 47
MLFLTRNRRILQ+GVDFKQIDKEWD DNFLILQTIIASMAYYM PFL
Sbjct: 446 MLFLTRNRRILQEGVDFKQIDKEWD*DNFLILQTIIASMAYYMPPFL 586
>AU089312
Length = 272
Score = 76.6 bits (187), Expect = 1e-14
Identities = 38/89 (42%), Positives = 52/89 (57%)
Frame = +3
Query: 121 IPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHS 180
IP+L S + + GYV D + +GHCN +VP LF FP +KYL+YTP++HS
Sbjct: 9 IPLLTLVFTRTSSVAAMVGYVTYIDVMNNMGHCNFXIVPKWLFDVFPPLKYLMYTPSFHS 188
Query: 181 LHHTDNKDTNFCLFMPLFDALGHTLNTKS 209
LHHT TN+ LFMP + + T + S
Sbjct: 189 LHHT-XFXTNYSLFMPXYXYIYXTXDKAS 272
>TC10799 weakly similar to UP|O23678 (O23678) CER1-like protein, partial
(13%)
Length = 605
Score = 65.1 bits (157), Expect = 3e-11
Identities = 37/108 (34%), Positives = 56/108 (51%)
Frame = +2
Query: 450 VGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMER 509
VG + E+ A G+ F F P R+DC Y AM P + + SCE + R
Sbjct: 2 VGDQLNEAERQKAAHGSLFIPFSRFPPKKLRKDCFYLSTPAMVTPPSLVNVHSCENWLPR 181
Query: 510 GVVHACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWKAALKHGLRPVSS 557
V+ A G++H+LE W +E G + + I+ VW+A+L HG RP+ +
Sbjct: 182 RVMSAWRIAGILHALEDWNINECGDV-MFSIEKVWQASLYHGFRPLKT 322
>TC8845 similar to UP|Q8VWZ8 (Q8VWZ8) Sterol 4-alpha-methyl-oxidase,
partial (97%)
Length = 1159
Score = 52.0 bits (123), Expect = 3e-07
Identities = 26/82 (31%), Positives = 44/82 (52%), Gaps = 2/82 (2%)
Frame = +3
Query: 50 LPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATI 109
LPSWN+ V+ ++ + + + ++YW HR H +L+ + HS+HH P LT +A
Sbjct: 429 LPSWNI--VLTQIIFYFILEDFIFYWGHRILHTKWLYKHVHSVHHEYATPFGLTSEYAHP 602
Query: 110 VEHAFI--SVVIGIPILGASVM 129
E F+ + + G I G +M
Sbjct: 603 AEILFLGFATIFGPAITGPHLM 668
>TC13320 similar to UP|O80456 (O80456) At2g23390 protein
(At2g23390/F26B6.4), partial (19%)
Length = 565
Score = 30.0 bits (66), Expect = 1.0
Identities = 31/119 (26%), Positives = 49/119 (41%), Gaps = 2/119 (1%)
Frame = -2
Query: 339 NKNEALNGGGKLFVDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIA 398
++++ L G ++ + H L +R T I D IPK V + + +A
Sbjct: 327 SEHDTLLKGTRITQEFHNKLHLR-----RFTKQKIFDSIPKFV-----------INKVMA 196
Query: 399 LYLCQKKVKVL--MLTLSADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWIT 455
C +K+ L M TLS+ F + E + ++ K Q TWIVG W T
Sbjct: 195 AICCHRKISTLNLMFTLSSS-FYCFKVEFNSCFNCLVITCFKMQ------TWIVGLWPT 40
>TC8462 similar to UP|PSAN_ARATH (P49107) Photosystem I reaction centre
subunit N, chloroplast precursor (PSI-N), partial (80%)
Length = 841
Score = 29.6 bits (65), Expect = 1.4
Identities = 19/46 (41%), Positives = 23/46 (49%)
Frame = -2
Query: 148 RCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNKDTNFCL 193
R LGH N V H KTF YL + P++ S +D T FCL
Sbjct: 753 RTLGHENDAVFNHHFQKTFEPHLYLSF-PSHSSSKSSDMNGTFFCL 619
>BU494310
Length = 402
Score = 29.6 bits (65), Expect = 1.4
Identities = 12/42 (28%), Positives = 20/42 (47%)
Frame = -3
Query: 420 RIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSW 461
R QK +PQ+ + + + ++ Q W+ G W RE W
Sbjct: 226 RFQKVSPQQEEVFWLHEQRHDNEQAFHQWLWGGWCRNREPPW 101
>TC16060 weakly similar to UP|Q9SHK4 (Q9SHK4) F12K11.6, partial (8%)
Length = 835
Score = 29.3 bits (64), Expect = 1.8
Identities = 19/78 (24%), Positives = 36/78 (45%), Gaps = 9/78 (11%)
Frame = +2
Query: 315 NKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKH---------PNLRVRVVHG 365
+++ E A K+GVK + + + K + GKL + ++ P + ++ +H
Sbjct: 215 DREAEVKAFDATKLGVKGLVESGVTKIPRIFNSGKLDITENAPTDSMLNVPIIDLKDIHN 394
Query: 366 NTLTAAVIIDEIPKDVEE 383
N A +ID+I EE
Sbjct: 395 NPALRAAVIDQIRSACEE 448
>AV778144
Length = 533
Score = 28.9 bits (63), Expect = 2.3
Identities = 12/25 (48%), Positives = 13/25 (52%)
Frame = +3
Query: 160 HGLFKTFPFMKYLLYTPTYHSLHHT 184
H K PF L Y P YH LH+T
Sbjct: 81 HNFRKRIPFKSKLEYPPGYHHLHNT 155
>TC9252 similar to UP|Q93X62 (Q93X62) Beta-oxyacyl-[acyl-carrier protein]
reductase , partial (28%)
Length = 593
Score = 28.9 bits (63), Expect = 2.3
Identities = 18/46 (39%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Frame = +3
Query: 380 DVEEVFLTGATSKLGRAIALYLCQKKVKVLM-LTLSADRFKRIQKE 424
D V +TGA+ +GRAIAL L + KVL+ S++ + + KE
Sbjct: 390 DAPVVVVTGASRGIGRAIALSLGKAGCKVLVNYARSSEEAEEVSKE 527
>AV777334
Length = 592
Score = 28.9 bits (63), Expect = 2.3
Identities = 28/129 (21%), Positives = 53/129 (40%), Gaps = 3/129 (2%)
Frame = -2
Query: 29 FLILQTII--ASMAYYMLPFLQNLPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLF 86
FLI++ I A ++ + + S + G ++ ++ S Y K DY
Sbjct: 393 FLIIKQSINLAFSFFFFILYHYFFDSVSTFGAVLDMITDRSFSLYSMYACKPKECADYCC 214
Query: 87 TNYHSLHHS-SPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFD 145
N+ L H + + L+ G + ++ F+ + M Y +V+G+V F+
Sbjct: 213 FNFFILFHKCTSMIFLLSSGSSVVIIFNFLD*EL--------CMSYFGLKVVHGFVSLFE 58
Query: 146 FLRCLGHCN 154
C G+CN
Sbjct: 57 LFCCFGYCN 31
>AV771287
Length = 525
Score = 28.1 bits (61), Expect = 3.9
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = +2
Query: 53 WNVKGVIVALVLHVGISEPLYYWVHRKFHG 82
W + V + L++ S PLY W+H FHG
Sbjct: 398 WA*QKVQESTSLYLLHSSPLYAWLHESFHG 487
>TC14900 similar to GB|AAL36081.1|17380636|AY063731 AT3g49800/T16K5_150
{Arabidopsis thaliana;}, partial (5%)
Length = 534
Score = 27.7 bits (60), Expect = 5.1
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = -1
Query: 71 PLYYWVHRKFHGDYLFTNYHSL 92
PL+Y + FH D F NYH L
Sbjct: 66 PLHYQYEQMFHWDQPFQNYHPL 1
>TC10057 similar to UP|Q41124 (Q41124) Chloroplast RNA binding protein
precursor, partial (49%)
Length = 598
Score = 27.7 bits (60), Expect = 5.1
Identities = 17/47 (36%), Positives = 23/47 (48%)
Frame = -3
Query: 56 KGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESL 102
+G V LVL IS P + FH D+ Y+ H+SS +SL
Sbjct: 578 RGTRVLLVLVCQISSPAKFGPRTLFHQDFR*QYYNLQHNSSSQMQSL 438
>AU089082
Length = 300
Score = 27.3 bits (59), Expect = 6.7
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = +3
Query: 81 HGDYLFTNYHSLHHSSPV 98
H FT+YH LHH +P+
Sbjct: 120 HSHSNFTHYHCLHHQTPI 173
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,915,774
Number of Sequences: 28460
Number of extensions: 198674
Number of successful extensions: 1377
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 1364
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1373
length of query: 562
length of database: 4,897,600
effective HSP length: 95
effective length of query: 467
effective length of database: 2,193,900
effective search space: 1024551300
effective search space used: 1024551300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0041.8


