

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0041.7
(1610 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
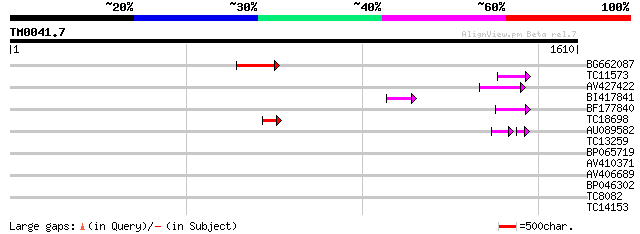
Score E
Sequences producing significant alignments: (bits) Value
BG662087 115 4e-26
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 72 9e-13
AV427422 61 1e-09
BI417841 59 8e-09
BF177840 54 2e-07
TC18698 51 2e-06
AU089582 35 7e-05
TC13259 39 0.006
BP065719 35 0.093
AV410371 31 1.8
AV406689 30 3.0
BP046302 30 3.0
TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-... 30 3.0
TC14153 weakly similar to UP|P93333 (P93333) PR10-1 protein (Cla... 28 8.7
>BG662087
Length = 373
Score = 115 bits (289), Expect = 4e-26
Identities = 58/122 (47%), Positives = 81/122 (65%)
Frame = +1
Query: 645 NGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVS 704
+GK R+ +D+ DLN A PKD Y +P + +VD A+ +E LSL+D YSGY+QI + D
Sbjct: 13 SGKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDED 192
Query: 705 KTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRD 764
KTAF A Y + +PFGLKNAGATYQ +M+ +F D + M+VY+D+++VKS R
Sbjct: 193 KTAFMT--ARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRA 366
Query: 765 EH 766
H
Sbjct: 367 NH 372
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 71.6 bits (174), Expect = 9e-13
Identities = 33/96 (34%), Positives = 52/96 (53%)
Frame = +2
Query: 1384 DQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPKSWHE 1443
+ G F ++ F + GI++ S + Q NGQ EAANK+++ IK+ + W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 1444 SLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEV 1479
L VLW+Y P+ + TPF L YG + +LP E+
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEI 295
>AV427422
Length = 417
Score = 61.2 bits (147), Expect = 1e-09
Identities = 33/133 (24%), Positives = 66/133 (48%), Gaps = 1/133 (0%)
Frame = +1
Query: 1333 SRQHKYIIVAIDYFIKWVEAIPLQS-VTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVG 1391
S+ ++ ++V +D K+ +PL+ T + + + +V G+P S+ +D+ +F+
Sbjct: 19 SKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMS 198
Query: 1392 RKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPKSWHESLSQVLWA 1451
+ G KL S Y+ +++GQ E N+ L + ++ + +PKSW + +
Sbjct: 199 NFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYW 378
Query: 1452 YRNSPREATGTTP 1464
Y S +TG TP
Sbjct: 379 YNTSYHVSTGQTP 417
>BI417841
Length = 617
Score = 58.5 bits (140), Expect = 8e-09
Identities = 29/85 (34%), Positives = 50/85 (58%)
Frame = +1
Query: 1071 SNNESEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLL 1130
+NN++EY L+ GL+ G +++ VKGDS+LV KQ+ +K + N+A +A L
Sbjct: 253 TNNQAEYRGLILGLKHAHEQGYQHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELK 432
Query: 1131 ANFDQAGVSYIPRVSNQEANELAQI 1155
+ F ++++PR N EA+ A +
Sbjct: 433 SKFQSFDINHVPRQYNSEADVQANL 507
>BF177840
Length = 410
Score = 53.9 bits (128), Expect = 2e-07
Identities = 29/100 (29%), Positives = 46/100 (46%)
Frame = +2
Query: 1380 SLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPK 1439
S+ +D+ T F+ G KLL S + Q +GQ E NK L +L++ + R K
Sbjct: 14 SIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLK 193
Query: 1440 SWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEV 1479
W L + +AY T +PF + YG + P ++
Sbjct: 194 MWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>TC18698
Length = 808
Score = 50.8 bits (120), Expect = 2e-06
Identities = 25/56 (44%), Positives = 35/56 (61%)
Frame = -2
Query: 717 YEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRK 772
Y + VMP GLKN TYQR+M+ IFH I ++VY++D++VKS H L +
Sbjct: 771 YYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*FHRGDLSR 604
>AU089582
Length = 383
Score = 35.4 bits (80), Expect(2) = 7e-05
Identities = 18/63 (28%), Positives = 33/63 (51%)
Frame = +2
Query: 1369 NHIVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILIS 1428
+ IV G+P S+ +D+G F +F + G +L S ++ Q +GQ E +IL
Sbjct: 38 DEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSERTIQILED 217
Query: 1429 LIK 1431
+++
Sbjct: 218 MLR 226
Score = 29.3 bits (64), Expect(2) = 7e-05
Identities = 14/37 (37%), Positives = 19/37 (50%)
Frame = +3
Query: 1440 SWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLP 1476
SW + LS + +AY NS R + PF YG+ P
Sbjct: 252 SWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSP 362
>TC13259
Length = 506
Score = 38.9 bits (89), Expect = 0.006
Identities = 29/82 (35%), Positives = 44/82 (53%)
Frame = +1
Query: 1075 SEYEALVSGLEILLALGAKNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFD 1134
SE AL GL++ L LG VEV+ D+++VV K +S LA + + F
Sbjct: 109 SEAMALRWGLQLSLELGFFEVEVEIDNQVVVNSWKGTKKHVSY-LANVIADVVRISQCFR 285
Query: 1135 QAGVSYIPRVSNQEANELAQIA 1156
++Y+PR+S+ A+ LAQ A
Sbjct: 286 FFSLAYVPRLSHLAADYLAQFA 351
>BP065719
Length = 567
Score = 35.0 bits (79), Expect = 0.093
Identities = 25/49 (51%), Positives = 27/49 (55%)
Frame = +1
Query: 1 *V*RIWLASKERMPNLLMTT*IDSGC*NLDPLLMSQNMNWLFWPQVV*S 49
*V R WL SKE +LLM T* GC NLD M N + L VV*S
Sbjct: 421 *VXRTWLVSKESQQSLLMIT*TGLGCLNLDASPMCLNTS*LS*XLVV*S 567
>AV410371
Length = 347
Score = 30.8 bits (68), Expect = 1.8
Identities = 17/51 (33%), Positives = 26/51 (50%), Gaps = 3/51 (5%)
Frame = +3
Query: 321 NPTQEKVKVAARDEDEDMLEDFDESDGEDLNI---ICIDDKAIFQRPSEHM 368
NP + +D+DEDM+ED D+ ED + DD+ IF+ +M
Sbjct: 195 NPNGSNFDNSCQDDDEDMVEDDDDDADEDPGYDTDLYYDDEDIFEDDYSNM 347
>AV406689
Length = 418
Score = 30.0 bits (66), Expect = 3.0
Identities = 17/36 (47%), Positives = 23/36 (63%), Gaps = 3/36 (8%)
Frame = +3
Query: 110 STRTTRSPPKS---TLMTSIWLSSSQETLMCARH*S 142
S+RTTRSPP+S T++W S+S T A +*S
Sbjct: 180 SSRTTRSPPRS*FNLFPTALWFSASPPTSTAA*N*S 287
>BP046302
Length = 435
Score = 30.0 bits (66), Expect = 3.0
Identities = 14/80 (17%), Positives = 35/80 (43%)
Frame = +3
Query: 194 LNRRKAKNTVSITTFLATGPHNVFVSGTLFKEHWILVGSSTAKRLKAKQRSTQIHSRKRS 253
+ + + + V++ + G H + G+ ++ HW +G ++ + +H S
Sbjct: 150 VTNQASNSDVNLCKRVTYGTHQLTDIGSTYRSHWRAIGGIECPGVRWSSNAIFLHPMVES 329
Query: 254 TPKLEKLLSSSYLEAKMEKR 273
P E ++ +E +M+ R
Sbjct: 330 LPMKELVVVVGRMEVRMKVR 389
>TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(54%)
Length = 1913
Score = 30.0 bits (66), Expect = 3.0
Identities = 16/46 (34%), Positives = 23/46 (49%), Gaps = 9/46 (19%)
Frame = +1
Query: 1492 IPSEVYWNMMLD---------ELVNLDEERVLALDVLTRQKDRIAK 1528
+P E+YW MML EL+ D +D T+ KD+I+K
Sbjct: 97 LPPEIYWKMMLPSTPMPKAVIELLRGDVSIETCVDTYTQSKDKISK 234
>TC14153 weakly similar to UP|P93333 (P93333) PR10-1 protein (Class 10 PR
protein), partial (97%)
Length = 754
Score = 28.5 bits (62), Expect = 8.7
Identities = 10/21 (47%), Positives = 17/21 (80%)
Frame = +3
Query: 369 KSHLKPLFVWAKVEDKGVNKI 389
KSHL+P+++W ++ED+ N I
Sbjct: 387 KSHLRPIWLWVQMEDQLQNTI 449
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,689,942
Number of Sequences: 28460
Number of extensions: 361690
Number of successful extensions: 2149
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 2122
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2146
length of query: 1610
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1507
effective length of database: 1,966,220
effective search space: 2963093540
effective search space used: 2963093540
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0041.7


