

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0037a.10
(168 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
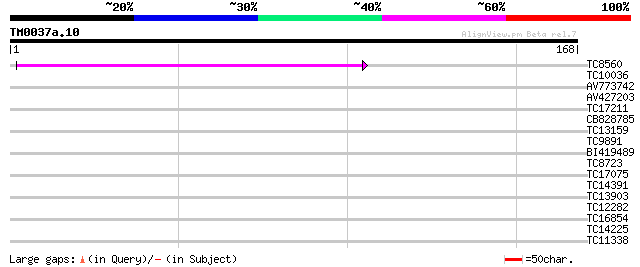
Score E
Sequences producing significant alignments: (bits) Value
TC8560 53 2e-08
TC10036 weakly similar to UP|Q9LFB5 (Q9LFB5) Anthranilate N-benz... 27 2.4
AV773742 27 2.4
AV427203 27 2.4
TC17211 similar to UP|PRT2_SEPOF (P80002) Spermatid-specific pro... 26 3.1
CB828785 26 3.1
TC13159 26 3.1
TC9891 similar to UP|XTH9_ARATH (Q8LDW9) Xyloglucan endotransglu... 26 4.0
BI419489 26 4.0
TC8723 homologue to UP|TBB1_LUPAL (P37392) Tubulin beta-1 chain,... 26 4.0
TC17075 similar to UP|Q7Y0H8 (Q7Y0H8) Cinnamoyl CoA reductase, p... 25 5.3
TC14391 similar to UP|Q93YG9 (Q93YG9) Insulin degrading enzyme, ... 25 5.3
TC13903 similar to UP|Q9FJD6 (Q9FJD6) Respiratory burst oxidase ... 25 5.3
TC12282 similar to UP|Q8S996 (Q8S996) Glucosyltransferase-13 (Fr... 25 6.9
TC16854 25 9.0
TC14225 similar to UP|Q40478 (Q40478) EREBP-4, partial (5%) 25 9.0
TC11338 similar to UP|Q8LDT8 (Q8LDT8) GDSL-motif lipase/hydrolas... 25 9.0
>TC8560
Length = 1044
Score = 53.1 bits (126), Expect = 2e-08
Identities = 31/104 (29%), Positives = 52/104 (49%)
Frame = +2
Query: 3 IVSRKFAVEFAHDLGLQWKLVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQGMNWF 62
+V R+F +F L +LVDP G+ VS+ + +EP + E+R + +QG
Sbjct: 173 VVPRQFLNDFEESLSKYVELVDPVGNKFCVSFYFDRDEPRFGANISELRTVHQIQGSVII 352
Query: 63 LLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSHTSFH 106
Y G S FD+ I + ++ E+ Y R+ + + SH +FH
Sbjct: 353 CFIYLGDSRFDIPISDLNLFEIEYRKRTAHVAANVASSSHLNFH 484
>TC10036 weakly similar to UP|Q9LFB5 (Q9LFB5) Anthranilate
N-benzoyltransferase-like protein (AT5g01210/F7J8_190),
partial (32%)
Length = 502
Score = 26.6 bits (57), Expect = 2.4
Identities = 17/50 (34%), Positives = 23/50 (46%)
Frame = +3
Query: 90 SLPPSITPTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLSPNQVSSS 139
+L PSI+P+T S C S+S + +P TT SP S S
Sbjct: 330 TLKPSISPSTPSFRPVSSTCILVSRSFSPTTCRFPTPATTSPSPPSRSPS 479
>AV773742
Length = 345
Score = 26.6 bits (57), Expect = 2.4
Identities = 13/46 (28%), Positives = 26/46 (56%)
Frame = +3
Query: 50 IRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSI 95
I+ +Y +++F + G F +F YN SI+ ++ +LPP++
Sbjct: 108 IKSYYD--NLSYFKARRDGIITFLIFKYNSSIQCAK*ISENLPPNM 239
>AV427203
Length = 426
Score = 26.6 bits (57), Expect = 2.4
Identities = 17/61 (27%), Positives = 30/61 (48%), Gaps = 2/61 (3%)
Frame = -2
Query: 47 LLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSL--PPSITPTTHSHTS 104
LLEIRKF +L +N ++ ++ ++ + ++ CY ++ L PP H S
Sbjct: 413 LLEIRKFQNLNHLNLYI--GHNKTSTEIP*WEKKLQHFCYWSQKLENPPPDLTLNHQANS 240
Query: 105 F 105
F
Sbjct: 239 F 237
>TC17211 similar to UP|PRT2_SEPOF (P80002) Spermatid-specific protein T2
[Contains: Sperm protamine SP2], partial (31%)
Length = 540
Score = 26.2 bits (56), Expect = 3.1
Identities = 11/16 (68%), Positives = 11/16 (68%)
Frame = +1
Query: 87 LTRSLPPSITPTTHSH 102
L RSLPP I THSH
Sbjct: 100 LRRSLPPQILTRTHSH 147
>CB828785
Length = 382
Score = 26.2 bits (56), Expect = 3.1
Identities = 10/30 (33%), Positives = 15/30 (49%)
Frame = +2
Query: 93 PSITPTTHSHTSFHGECTTGSKSVTEEWIF 122
P+ TP T+S FH S S+ W++
Sbjct: 11 PTSTPKTYSTLCFHNCKIPSSSSIPSNWVY 100
>TC13159
Length = 439
Score = 26.2 bits (56), Expect = 3.1
Identities = 15/44 (34%), Positives = 22/44 (49%)
Frame = +1
Query: 96 TPTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLSPNQVSSS 139
TPT HS +SF+G K+ T F+ + L +QV +S
Sbjct: 247 TPTVHSRSSFNG------KAATNHMSFFTFSDSRELKKDQVPNS 360
>TC9891 similar to UP|XTH9_ARATH (Q8LDW9) Xyloglucan
endotransglucosylase/hydrolase protein 9 precursor
(At-XTH9) (XTH-9) , partial (41%)
Length = 600
Score = 25.8 bits (55), Expect = 4.0
Identities = 12/26 (46%), Positives = 14/26 (53%), Gaps = 1/26 (3%)
Frame = -1
Query: 78 NDSIEEVCYLT-RSLPPSITPTTHSH 102
N+S CY + S PPS P TH H
Sbjct: 273 NNSYPHCCYTSSHSPPPSPAPDTHKH 196
>BI419489
Length = 479
Score = 25.8 bits (55), Expect = 4.0
Identities = 15/43 (34%), Positives = 20/43 (45%), Gaps = 3/43 (6%)
Frame = +2
Query: 78 NDSIEEVCYLTRSLPPSITPTT---HSHTSFHGECTTGSKSVT 117
N YLT S PP+ TP S T+ +CT S++ T
Sbjct: 92 NQPPNPTAYLTASPPPTATPAAL*KTSSTTPTAKCTVPSRTTT 220
>TC8723 homologue to UP|TBB1_LUPAL (P37392) Tubulin beta-1 chain, complete
Length = 1689
Score = 25.8 bits (55), Expect = 4.0
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = -3
Query: 88 TRSLPPSITPTTHSHTSFHGECTTGS 113
T++ PP + P HSHT H + G+
Sbjct: 1462 TKNPPPELHPHLHSHTPHHLQLHPGT 1385
>TC17075 similar to UP|Q7Y0H8 (Q7Y0H8) Cinnamoyl CoA reductase, partial (7%)
Length = 534
Score = 25.4 bits (54), Expect = 5.3
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = -2
Query: 4 VSRKFAVEFAHDLGLQWKLVDPFGS 28
V+R+ V+F H+ G KLVD G+
Sbjct: 173 VNREIRVQFRHNAGKTRKLVDTIGA 99
>TC14391 similar to UP|Q93YG9 (Q93YG9) Insulin degrading enzyme, partial
(10%)
Length = 782
Score = 25.4 bits (54), Expect = 5.3
Identities = 12/34 (35%), Positives = 19/34 (55%), Gaps = 2/34 (5%)
Frame = +3
Query: 27 GSVHLVSYESNSEEPYL--FEGLLEIRKFYSLQG 58
GS+H Y++ + EP+L + + RK SL G
Sbjct: 279 GSLHSSEYKAEASEPHLAKIDNIFSFRKSQSLYG 380
>TC13903 similar to UP|Q9FJD6 (Q9FJD6) Respiratory burst oxidase protein,
partial (7%)
Length = 558
Score = 25.4 bits (54), Expect = 5.3
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = +1
Query: 59 MNWFLLKYQGCSNFDLFIYNDSIEEV 84
+N+FLL ++ + LF YN S+E +
Sbjct: 253 LNYFLLLHETAKDRRLFFYNHSLENL 330
>TC12282 similar to UP|Q8S996 (Q8S996) Glucosyltransferase-13 (Fragment),
partial (7%)
Length = 511
Score = 25.0 bits (53), Expect = 6.9
Identities = 24/88 (27%), Positives = 34/88 (38%), Gaps = 16/88 (18%)
Frame = -1
Query: 33 SYESNSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNF-----------DLFIYNDSI 81
SY+S Y F + R +Y N F LK Q ++F LF + +
Sbjct: 460 SYDSRFGSVYFFS*SITHRNYYRSFSQN*FQLKNQLENDF*TCYQLLGATHKLFHFRARL 281
Query: 82 E-----EVCYLTRSLPPSITPTTHSHTS 104
E E C L+ + PS+ T H S
Sbjct: 280 EIAFLDEPCSLSAAAAPSLNSLTLLHIS 197
>TC16854
Length = 568
Score = 24.6 bits (52), Expect = 9.0
Identities = 10/23 (43%), Positives = 12/23 (51%)
Frame = -3
Query: 7 KFAVEFAHDLGLQWKLVDPFGSV 29
K F + LGL W LV P S+
Sbjct: 104 KILPPFTYSLGLSWVLVSPVDSI 36
>TC14225 similar to UP|Q40478 (Q40478) EREBP-4, partial (5%)
Length = 604
Score = 24.6 bits (52), Expect = 9.0
Identities = 16/51 (31%), Positives = 20/51 (38%), Gaps = 9/51 (17%)
Frame = -1
Query: 98 TTHSHTSFHGECTTGSKSV---------TEEWIFYPCFFTTRLSPNQVSSS 139
T+ S FH E G +E F+ CF T LS +SSS
Sbjct: 157 TSVSQNPFHNEGVKGQTVAAETETPLLGSEHSSFFTCFLTVTLSSTHLSSS 5
>TC11338 similar to UP|Q8LDT8 (Q8LDT8) GDSL-motif lipase/hydrolase-like
protein, partial (6%)
Length = 600
Score = 24.6 bits (52), Expect = 9.0
Identities = 16/58 (27%), Positives = 26/58 (44%), Gaps = 2/58 (3%)
Frame = +2
Query: 74 LFIYNDSIEEVCYLTRSLPPSITPTTHSHTSFHGECTTGSKSV--TEEWIFYPCFFTT 129
+FI + SI+ L + S+T T+ HGEC + E++F+ F T
Sbjct: 23 IFINSTSIDRENSLVSGI--SVTDAACCRTTLHGECIPDERPCYNRNEYVFWDDFHPT 190
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.136 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,711,112
Number of Sequences: 28460
Number of extensions: 60736
Number of successful extensions: 402
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 402
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 402
length of query: 168
length of database: 4,897,600
effective HSP length: 84
effective length of query: 84
effective length of database: 2,506,960
effective search space: 210584640
effective search space used: 210584640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0037a.10


