

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0036c.6
(70 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
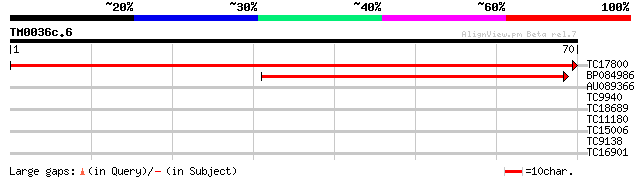
TC17800 similar to UP|KIWI_ARATH (O65154) RNA polymerase II tran... 145 1e-36
BP084986 41 4e-05
AU089366 25 3.7
TC9940 weakly similar to UP|Q9LHB1 (Q9LHB1) Transcription factor... 25 3.7
TC18689 24 6.2
TC11180 24 6.2
TC15006 weakly similar to UP|P93333 (P93333) PR10-1 protein (Cla... 23 8.2
TC9138 similar to UP|Q8VX01 (Q8VX01) Casein kinase II alpha subu... 23 8.2
TC16901 similar to GB|BAB03049.1|9294683|AP001305 syringomycin b... 23 8.2
>TC17800 similar to UP|KIWI_ARATH (O65154) RNA polymerase II transcriptional
coactivator KIWI, partial (60%)
Length = 592
Score = 145 bits (367), Expect = 1e-36
Identities = 70/70 (100%), Positives = 70/70 (100%)
Frame = +2
Query: 1 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI
Sbjct: 149 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 328
Query: 61 EEIDKAVNES 70
EEIDKAVNES
Sbjct: 329 EEIDKAVNES 358
>BP084986
Length = 515
Score = 41.2 bits (95), Expect = 4e-05
Identities = 16/38 (42%), Positives = 25/38 (65%)
Frame = -1
Query: 32 FYVKDGKQMPGKKGISLTMDQWNVLRNHIEEIDKAVNE 69
+Y KDGK +P KGISLT +QW+ + + +KA+ +
Sbjct: 515 YYRKDGKDLPTSKGISLTEEQWSTFKKSVPAKEKAIEK 402
>AU089366
Length = 717
Score = 24.6 bits (52), Expect = 3.7
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = +1
Query: 44 KGISLTMDQWNVLRNHIEEIDKAVNE 69
K I Q +LR H++ +D AVN+
Sbjct: 211 KNIDGLASQLKILRRHVDVLDSAVNK 288
>TC9940 weakly similar to UP|Q9LHB1 (Q9LHB1) Transcription factor X1-like
protein, partial (18%)
Length = 577
Score = 24.6 bits (52), Expect = 3.7
Identities = 11/44 (25%), Positives = 23/44 (52%)
Frame = +3
Query: 14 RRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLR 57
R ++ N GR + Y K+G++ K+G+ + + QW + +
Sbjct: 255 REINEYNPSGRYITSELWNY-KEGRRATSKEGVQVLLKQWKLYK 383
>TC18689
Length = 397
Score = 23.9 bits (50), Expect = 6.2
Identities = 8/21 (38%), Positives = 13/21 (61%)
Frame = +1
Query: 49 TMDQWNVLRNHIEEIDKAVNE 69
+M W +L+ H EI + +NE
Sbjct: 175 SMRIWEILKTHWNEISRMINE 237
>TC11180
Length = 640
Score = 23.9 bits (50), Expect = 6.2
Identities = 9/14 (64%), Positives = 12/14 (85%)
Frame = +1
Query: 54 NVLRNHIEEIDKAV 67
N+LRN +EEID A+
Sbjct: 16 NILRNKLEEIDVAI 57
>TC15006 weakly similar to UP|P93333 (P93333) PR10-1 protein (Class 10 PR
protein), partial (75%)
Length = 1047
Score = 23.5 bits (49), Expect = 8.2
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = -1
Query: 29 IREFYVKDGKQMPGKKGISLTMDQWNVLRNHIEEI 63
I+ ++ DG PG T+D +N LR + EI
Sbjct: 258 IKHGHLLDGSSSPGSFDEFNTLDGFNNLREDVVEI 154
>TC9138 similar to UP|Q8VX01 (Q8VX01) Casein kinase II alpha subunit
precursor , partial (24%)
Length = 693
Score = 23.5 bits (49), Expect = 8.2
Identities = 11/34 (32%), Positives = 19/34 (55%)
Frame = -1
Query: 27 VDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
+++ EFY ++ KG+ + QW VL +HI
Sbjct: 315 LNVFEFYYFLHFELD*SKGVPWLL*QWGVLGDHI 214
>TC16901 similar to GB|BAB03049.1|9294683|AP001305 syringomycin
biosynthesis enzyme-like protein {Arabidopsis
thaliana;}, partial (16%)
Length = 533
Score = 23.5 bits (49), Expect = 8.2
Identities = 10/34 (29%), Positives = 17/34 (49%)
Frame = +3
Query: 25 IVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRN 58
I+ D ++ +P +KG L +D W VL +
Sbjct: 12 IIYDCLNILEEESVAIPWRKGDVLLLDNWAVLHS 113
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.136 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,126,150
Number of Sequences: 28460
Number of extensions: 10631
Number of successful extensions: 47
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 47
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 47
length of query: 70
length of database: 4,897,600
effective HSP length: 46
effective length of query: 24
effective length of database: 3,588,440
effective search space: 86122560
effective search space used: 86122560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0036c.6


