

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0036b.1
(122 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
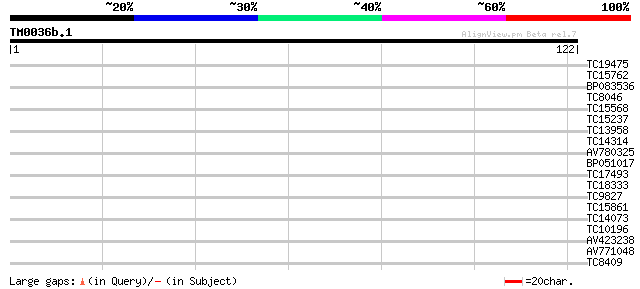
Score E
Sequences producing significant alignments: (bits) Value
TC19475 30 0.12
TC15762 similar to GB|BAA25822.1|3088342|AB007158 ribosomal prot... 29 0.26
BP083536 28 0.34
TC8046 similar to UP|Q7XJC2 (Q7XJC2) Adenine phosphoribosyltrans... 28 0.34
TC15568 27 0.98
TC15237 similar to UP|AAQ24632 (AAQ24632) GPRP, partial (64%) 27 0.98
TC13958 similar to AAQ89652 (AAQ89652) At3g54880, partial (37%) 27 1.3
TC14314 similar to UP|Q9FPI4 (Q9FPI4) AT5g16110 (AT5g16110/T21H1... 27 1.3
AV780325 26 1.7
BP051017 26 2.2
TC17493 similar to GB|AAO64904.1|29029050|BT005969 At3g57140 {Ar... 25 3.7
TC18333 similar to UP|Q9FXM3 (Q9FXM3) Replication factor C 37kDa... 25 3.7
TC9827 weakly similar to UP|Q8S9A2 (Q8S9A2) Glucosyltransferase-... 25 3.7
TC15861 similar to UP|S61G_SCHPO (Q09827) Probable protein trans... 25 3.7
TC14073 homologue to UP|AAQ08403 (AAQ08403) Methionine synthase,... 25 4.8
TC10196 homologue to UP|Q39804 (Q39804) BiP isoform B, partial (... 25 4.8
AV423238 25 4.8
AV771048 25 4.8
TC8409 similar to UP|Q9FPS8 (Q9FPS8) Ubiquitin-specific protease... 25 4.8
>TC19475
Length = 524
Score = 30.0 bits (66), Expect = 0.12
Identities = 19/44 (43%), Positives = 27/44 (61%), Gaps = 1/44 (2%)
Frame = +3
Query: 26 HSSTCNFSELLEILSRHVVAGIFYKGL-NIVLCILILQKDKLIR 68
H+STCNF L +LSR + + L ++LCI IL DK++R
Sbjct: 240 HTSTCNFFSL*-LLSRVLFCCSITQSLCYVILCI*ILLTDKIVR 368
>TC15762 similar to GB|BAA25822.1|3088342|AB007158 ribosomal protein S23
{Homo sapiens;} , partial (24%)
Length = 572
Score = 28.9 bits (63), Expect = 0.26
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = -1
Query: 92 YPPPLFRSLKIGELSKMPLFKKKKF 116
YPPPLF SL +S + +K KKF
Sbjct: 344 YPPPLFPSLSQPNVSLLASYKGKKF 270
>BP083536
Length = 377
Score = 28.5 bits (62), Expect = 0.34
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = +1
Query: 30 CNFSELLEILSRHVVAGIFYK 50
CN EL+EI RH +AG+ Y+
Sbjct: 313 CNLWELVEIPCRHALAGVHYR 375
>TC8046 similar to UP|Q7XJC2 (Q7XJC2) Adenine phosphoribosyltransferase
(Fragment), partial (85%)
Length = 1298
Score = 28.5 bits (62), Expect = 0.34
Identities = 20/58 (34%), Positives = 30/58 (51%), Gaps = 1/58 (1%)
Frame = +3
Query: 18 RFIVSLKDHSSTCNFSEL-LEILSRHVVAGIFYKGLNIVLCILILQKDKLIRTHTCTL 74
+FIV + HS TC FS++ LS+ LNI++ I+I+ D L + T L
Sbjct: 96 QFIVGVSRHSHTCLFSQIPSTTLSKD-------SKLNIIIIIIIILLDSLPKFSTTPL 248
>TC15568
Length = 637
Score = 26.9 bits (58), Expect = 0.98
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = -2
Query: 14 LRSDRFIVSLKDHSSTC 30
+RSDRF+ +LK SSTC
Sbjct: 63 MRSDRFLQNLKQPSSTC 13
>TC15237 similar to UP|AAQ24632 (AAQ24632) GPRP, partial (64%)
Length = 814
Score = 26.9 bits (58), Expect = 0.98
Identities = 10/28 (35%), Positives = 19/28 (67%)
Frame = -2
Query: 81 KSKVFDANCYSYPPPLFRSLKIGELSKM 108
K+ + D++C S PPP+F++ K + +M
Sbjct: 156 KTPLSDSSCLSLPPPIFQNQKPN*ICRM 73
>TC13958 similar to AAQ89652 (AAQ89652) At3g54880, partial (37%)
Length = 516
Score = 26.6 bits (57), Expect = 1.3
Identities = 13/37 (35%), Positives = 22/37 (59%)
Frame = +3
Query: 31 NFSELLEILSRHVVAGIFYKGLNIVLCILILQKDKLI 67
NFS +IL ++G+ + + +V+ LI+ KDK I
Sbjct: 312 NFS*CTDILINFTISGLLFSTIYLVVANLII*KDKNI 422
>TC14314 similar to UP|Q9FPI4 (Q9FPI4) AT5g16110 (AT5g16110/T21H19_30),
partial (12%)
Length = 1145
Score = 26.6 bits (57), Expect = 1.3
Identities = 15/51 (29%), Positives = 25/51 (48%), Gaps = 1/51 (1%)
Frame = +2
Query: 21 VSLKDHSSTCNFSELLEILSRHVVAGIFYKGLNIVLCILILQKDK-LIRTH 70
+ L+ S CN E + R +V IF+ G I C L++ + ++ TH
Sbjct: 983 LKLRSSSCNCNLEEDED*SLRSLVKNIFFVGAPIFRCYLLIDDIRDIVYTH 1135
>AV780325
Length = 524
Score = 26.2 bits (56), Expect = 1.7
Identities = 10/19 (52%), Positives = 15/19 (78%)
Frame = +3
Query: 34 ELLEILSRHVVAGIFYKGL 52
E+ +I+SRH + G+ YKGL
Sbjct: 222 EVTKIISRHWLVGVSYKGL 278
>BP051017
Length = 522
Score = 25.8 bits (55), Expect = 2.2
Identities = 13/42 (30%), Positives = 23/42 (53%)
Frame = -1
Query: 20 IVSLKDHSSTCNFSELLEILSRHVVAGIFYKGLNIVLCILIL 61
I KDH++ + L+ H++ Y+ LN++LC L+L
Sbjct: 414 IEKFKDHNAVPTPKKKLDPAQLHLM---IYRPLNVLLCFLLL 298
>TC17493 similar to GB|AAO64904.1|29029050|BT005969 At3g57140 {Arabidopsis
thaliana;}, partial (34%)
Length = 891
Score = 25.0 bits (53), Expect = 3.7
Identities = 18/45 (40%), Positives = 23/45 (51%), Gaps = 6/45 (13%)
Frame = +1
Query: 19 FIVSLKDHSSTCNFSELLEIL------SRHVVAGIFYKGLNIVLC 57
F VSLK H + CNFS E S+HVV +F + + LC
Sbjct: 328 FRVSLKTHGTQCNFSIKWEGFSQL*RGSQHVV--LFMRSDSCKLC 456
>TC18333 similar to UP|Q9FXM3 (Q9FXM3) Replication factor C 37kDa subunit,
partial (12%)
Length = 400
Score = 25.0 bits (53), Expect = 3.7
Identities = 17/48 (35%), Positives = 26/48 (53%), Gaps = 3/48 (6%)
Frame = +1
Query: 23 LKDHSSTCNFSELLEILSRHVVA---GIFYKGLNIVLCILILQKDKLI 67
L + +TC+F I + V G+ Y+GLNI L L+ Q++ LI
Sbjct: 94 LMELMNTCSFLMCSAIQRKPFVTCXEGLSYRGLNIRLSGLLNQEELLI 237
>TC9827 weakly similar to UP|Q8S9A2 (Q8S9A2) Glucosyltransferase-7
(Fragment), partial (42%)
Length = 611
Score = 25.0 bits (53), Expect = 3.7
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = +3
Query: 58 ILILQKDKLIRTHTCTLLHQLMGKSKVFDANCYSYPPPLFRSL 100
IL+L+++ LI+ LLH + + + CY+Y LFR++
Sbjct: 348 ILLLREEDLIKILQSLLLHWITSELLYINIFCYAY--ILFRTV 470
>TC15861 similar to UP|S61G_SCHPO (Q09827) Probable protein transport
protein SEC61 gamma subunit, partial (21%)
Length = 801
Score = 25.0 bits (53), Expect = 3.7
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = -3
Query: 95 PLFRSLKIGELSKMPLFKKKKFSLSPLT 122
PL +G +P FKK + SL+PLT
Sbjct: 220 PLSGFKALGSSGMLPTFKKPEDSLNPLT 137
>TC14073 homologue to UP|AAQ08403 (AAQ08403) Methionine synthase, complete
Length = 2757
Score = 24.6 bits (52), Expect = 4.8
Identities = 16/50 (32%), Positives = 29/50 (58%), Gaps = 4/50 (8%)
Frame = +3
Query: 41 RHVVAGIFYKGLNIVLCIL-ILQKDKLIRTHTCTLLH---QLMGKSKVFD 86
R++ A LN + + I+ KDKL+ + +C+LLH L+ ++K+ D
Sbjct: 1062 RNIWANDLASSLNTLAALEGIVGKDKLVVSTSCSLLHTAVDLVNETKLDD 1211
>TC10196 homologue to UP|Q39804 (Q39804) BiP isoform B, partial (17%)
Length = 562
Score = 24.6 bits (52), Expect = 4.8
Identities = 8/31 (25%), Positives = 16/31 (50%)
Frame = +2
Query: 1 MSDELDWYEPNDSLRSDRFIVSLKDHSSTCN 31
+ + L+W + N S + + LK+ + CN
Sbjct: 158 VKEALEWLDDNQSAEKEEYEEKLKEVEAVCN 250
>AV423238
Length = 373
Score = 24.6 bits (52), Expect = 4.8
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = -2
Query: 61 LQKDKLIRTHTCTLLHQLMGKSKVFDANC 89
LQK K+ + H C++ L+ SK D NC
Sbjct: 90 LQKTKMRKLHPCSVFFSLLFLSKE-DVNC 7
>AV771048
Length = 430
Score = 24.6 bits (52), Expect = 4.8
Identities = 19/55 (34%), Positives = 25/55 (44%), Gaps = 5/55 (9%)
Frame = +2
Query: 46 GIFYKGLNI----VLCIL-ILQKDKLIRTHTCTLLHQLMGKSKVFDANCYSYPPP 95
GIF G +I + CIL QK K+ Q+M KS + A C +P P
Sbjct: 53 GIFRNGPDIFTTTLTCILPFTQKKKVTEREKIK*KGQIMAKSIISLAKCGYFPTP 217
>TC8409 similar to UP|Q9FPS8 (Q9FPS8) Ubiquitin-specific protease 16,
partial (4%)
Length = 709
Score = 24.6 bits (52), Expect = 4.8
Identities = 14/43 (32%), Positives = 18/43 (41%), Gaps = 6/43 (13%)
Frame = -1
Query: 68 RTHTCTLLHQLMGKSKVFDANC------YSYPPPLFRSLKIGE 104
R+H C L H K+ NC Y L +SL +GE
Sbjct: 157 RSHQCWLNHAYFQYCKILHQNCC*KGMNYHLMSLLVKSLSVGE 29
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.140 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,560,048
Number of Sequences: 28460
Number of extensions: 36115
Number of successful extensions: 239
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 238
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 238
length of query: 122
length of database: 4,897,600
effective HSP length: 79
effective length of query: 43
effective length of database: 2,649,260
effective search space: 113918180
effective search space used: 113918180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0036b.1


