

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0035.12
(125 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
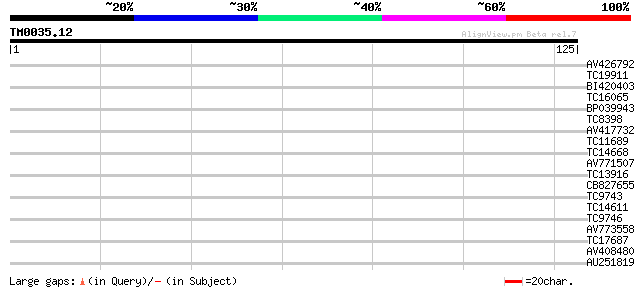
Score E
Sequences producing significant alignments: (bits) Value
AV426792 30 0.12
TC19911 weakly similar to PIR|A44805|A44805 eggshell protein pre... 29 0.27
BI420403 28 0.45
TC16065 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich... 28 0.59
BP039943 27 0.77
TC8398 weakly similar to UP|Q84KK1 (Q84KK1) S haplotype-specific... 27 0.77
AV417732 27 0.77
TC11689 similar to GB|AAD34673.1|4966342|F3O9 ESTs gb|T04357 and... 27 1.0
TC14668 UP|BAC98256 (BAC98256) MKIAA1813 protein (Fragment), par... 27 1.0
AV771507 27 1.0
TC13916 homologue to UP|Q9FK48 (Q9FK48) Similarity to transcript... 27 1.0
CB827655 27 1.3
TC9743 similar to UP|AAQ89667 (AAQ89667) At5g25170, partial (36%) 26 1.7
TC14611 similar to UP|Q8LPB2 (Q8LPB2) Glycine-rich RNA binding p... 26 1.7
TC9746 26 1.7
AV773558 26 1.7
TC17687 weakly similar to GB|AAH05628.1|13542865|BC005628 acidic... 26 1.7
AV408480 26 2.3
AU251819 25 3.8
>AV426792
Length = 432
Score = 30.0 bits (66), Expect = 0.12
Identities = 22/64 (34%), Positives = 34/64 (52%), Gaps = 7/64 (10%)
Frame = +1
Query: 69 NKTQIKTPLNKASTF---EATKLRSGTCIF----RSGAAVRFVILGFAERREMEEEEKRK 121
N TQI PL+K S+ +AT+ R G + G R ++ A R++ EEEKRK
Sbjct: 145 NLTQIPKPLSKLSSTKVEDATQARQGEKSTSEGPQRGEKFRQIMEQEARRKKQREEEKRK 324
Query: 122 QKQS 125
+++
Sbjct: 325 AEEA 336
>TC19911 weakly similar to PIR|A44805|A44805 eggshell protein precursor -
fluke (Schistosoma haematobium) {Schistosoma
haematobium;} , partial (24%)
Length = 350
Score = 28.9 bits (63), Expect = 0.27
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = -2
Query: 6 GQHHHSHRRSFFTHINWAPDQHPH 29
G HHH+HR+ H W+ + +PH
Sbjct: 154 GYHHHNHRQR--VHDRWSQNAYPH 89
>BI420403
Length = 425
Score = 28.1 bits (61), Expect = 0.45
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = -3
Query: 4 NPGQHHHSHRRSFFTHINWAPDQHPHEVLAFMPSL 38
+P +HHH H F D PH L+++PS+
Sbjct: 387 HPPRHHHHHHHLF-------DDDQPHAPLSWVPSM 304
>TC16065 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich
glycoprotein DZ-HRGP precursor, partial (20%)
Length = 608
Score = 27.7 bits (60), Expect = 0.59
Identities = 11/25 (44%), Positives = 13/25 (52%)
Frame = +2
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQH 27
HNP HHH RR H++ D H
Sbjct: 272 HNPAHHHHHSRR----HVHLVVDHH 334
>BP039943
Length = 481
Score = 27.3 bits (59), Expect = 0.77
Identities = 11/31 (35%), Positives = 21/31 (67%)
Frame = +2
Query: 67 IWNKTQIKTPLNKASTFEATKLRSGTCIFRS 97
++ K++IKT L+ ++ TKL + TC ++S
Sbjct: 191 LFGKSRIKTKLHNGPIYKKTKLVTHTCAWKS 283
>TC8398 weakly similar to UP|Q84KK1 (Q84KK1) S haplotype-specific F-box
protein a, partial (9%)
Length = 1602
Score = 27.3 bits (59), Expect = 0.77
Identities = 25/96 (26%), Positives = 37/96 (38%), Gaps = 8/96 (8%)
Frame = +3
Query: 3 HNP--GQHHHSHRRSFFTHINWA------PDQHPHEVLAFMPSLFKSNDLARRKRASDGD 54
H+P + H H +F W P PH + + PSLF S + + K
Sbjct: 282 HSPLLSKFHIVHDETFTADEPWPVHALREPFNQPHVSICYAPSLFSSQNNDKLKGHCRLI 461
Query: 55 KSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRS 90
+ N +N + T L+ + ATKLRS
Sbjct: 462 GACNGLVSVINEGYNSAESATKLSCCNLNPATKLRS 569
>AV417732
Length = 418
Score = 27.3 bits (59), Expect = 0.77
Identities = 17/51 (33%), Positives = 25/51 (48%), Gaps = 7/51 (13%)
Frame = -1
Query: 27 HPHEVLAFMPSLFK-------SNDLARRKRASDGDKSLNKTQEEKASIWNK 70
H H+V AF P LFK S + AR + G +SL + + A W++
Sbjct: 178 HCHQVQAFSPLLFKEARSREMSPEKARSEHFDWGRESLPEPRRSHAGRWSR 26
>TC11689 similar to GB|AAD34673.1|4966342|F3O9 ESTs gb|T04357 and
gb|AA595092 come from this gene. {Arabidopsis thaliana;}
, partial (26%)
Length = 573
Score = 26.9 bits (58), Expect = 1.0
Identities = 17/61 (27%), Positives = 29/61 (46%)
Frame = +3
Query: 21 NWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLNKA 80
N APD+HP L F+ +L + K G + N+ ++ +W K+ PLN+
Sbjct: 57 NLAPDRHPERRLKAAFKAFEEAELPKLKEEKPG-LTYNQYRDMIWKLWKKSP-DNPLNQV 230
Query: 81 S 81
+
Sbjct: 231 A 233
>TC14668 UP|BAC98256 (BAC98256) MKIAA1813 protein (Fragment), partial (3%)
Length = 597
Score = 26.9 bits (58), Expect = 1.0
Identities = 19/75 (25%), Positives = 32/75 (42%), Gaps = 6/75 (8%)
Frame = +2
Query: 8 HHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDK-----SLNKTQE 62
HHH H I ++ + VL + FK +R+R G K SL++T++
Sbjct: 122 HHHHHHHQKIMKI*FSFPPNFSSVLKYSSFFFKYVKRKKRRRKKRGRKFPSSASLSRTEQ 301
Query: 63 EKASI-WNKTQIKTP 76
++ I W +P
Sbjct: 302 KRGEIQWEGPTCPSP 346
>AV771507
Length = 480
Score = 26.9 bits (58), Expect = 1.0
Identities = 11/16 (68%), Positives = 15/16 (93%)
Frame = +2
Query: 110 ERREMEEEEKRKQKQS 125
E+R+M++EEKR QKQS
Sbjct: 62 EQRKMKKEEKRNQKQS 109
>TC13916 homologue to UP|Q9FK48 (Q9FK48) Similarity to transcription
regulator, partial (17%)
Length = 603
Score = 26.9 bits (58), Expect = 1.0
Identities = 19/51 (37%), Positives = 23/51 (44%), Gaps = 3/51 (5%)
Frame = +2
Query: 38 LFKSNDLARRKRASDGDKSLNKTQEEKA---SIWNKTQIKTPLNKASTFEA 85
LF + A RK + D+ L K QE SIWNK N+ FEA
Sbjct: 140 LFDESMGASRKLQGEIDRVLKKVQEGVEVFDSIWNKVYDTDNANQKEKFEA 292
>CB827655
Length = 472
Score = 26.6 bits (57), Expect = 1.3
Identities = 10/19 (52%), Positives = 16/19 (83%)
Frame = -2
Query: 106 LGFAERREMEEEEKRKQKQ 124
LGF ++R+ EE E++KQK+
Sbjct: 84 LGFQKKRKEEENERKKQKR 28
>TC9743 similar to UP|AAQ89667 (AAQ89667) At5g25170, partial (36%)
Length = 552
Score = 26.2 bits (56), Expect = 1.7
Identities = 9/20 (45%), Positives = 11/20 (55%)
Frame = +1
Query: 4 NPGQHHHSHRRSFFTHINWA 23
N HHH HRR +NW+
Sbjct: 232 NSHHHHHHHRRHHQCCVNWS 291
>TC14611 similar to UP|Q8LPB2 (Q8LPB2) Glycine-rich RNA binding protein,
partial (20%)
Length = 760
Score = 26.2 bits (56), Expect = 1.7
Identities = 10/27 (37%), Positives = 13/27 (48%)
Frame = -2
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHPH 29
H+P HHH H+ H + P PH
Sbjct: 291 HSPHYHHHHHQALEGAHDHTQPPPVPH 211
>TC9746
Length = 637
Score = 26.2 bits (56), Expect = 1.7
Identities = 9/25 (36%), Positives = 13/25 (52%)
Frame = +3
Query: 5 PGQHHHSHRRSFFTHINWAPDQHPH 29
P HHH+ H++W P + PH
Sbjct: 342 P*PHHHTLLHRTLQHLHWPPLRPPH 416
>AV773558
Length = 469
Score = 26.2 bits (56), Expect = 1.7
Identities = 9/29 (31%), Positives = 13/29 (44%)
Frame = +3
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHPHEV 31
H HHH H H ++ P Q+ H +
Sbjct: 312 HLCSHHHHHHHHHHHHHPHYTPTQNHHPI 398
>TC17687 weakly similar to GB|AAH05628.1|13542865|BC005628 acidic nuclear
phosphoprotein 32 family, member B {Mus musculus;},
partial (9%)
Length = 702
Score = 26.2 bits (56), Expect = 1.7
Identities = 10/28 (35%), Positives = 13/28 (45%)
Frame = -3
Query: 1 MIHNPGQHHHSHRRSFFTHINWAPDQHP 28
++H H HSH R+ H P HP
Sbjct: 388 LLHLHCTHSHSHHRTLHPHYPRPPQPHP 305
>AV408480
Length = 262
Score = 25.8 bits (55), Expect = 2.3
Identities = 12/32 (37%), Positives = 15/32 (46%)
Frame = +3
Query: 7 QHHHSHRRSFFTHINWAPDQHPHEVLAFMPSL 38
+HHH HR +N PD HP +P L
Sbjct: 156 RHHHHHRPPL--RLNLRPDPHPKMAPTRIPPL 245
>AU251819
Length = 368
Score = 25.0 bits (53), Expect = 3.8
Identities = 11/31 (35%), Positives = 14/31 (44%)
Frame = +2
Query: 2 IHNPGQHHHSHRRSFFTHINWAPDQHPHEVL 32
IH P ++HH HR H+ P H L
Sbjct: 134 IHIPLKNHHHHRHHHHHHLYLHPHHQYHPPL 226
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.129 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,337,815
Number of Sequences: 28460
Number of extensions: 31617
Number of successful extensions: 435
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 410
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 426
length of query: 125
length of database: 4,897,600
effective HSP length: 80
effective length of query: 45
effective length of database: 2,620,800
effective search space: 117936000
effective search space used: 117936000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 50 (23.9 bits)
Lotus: description of TM0035.12


