

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
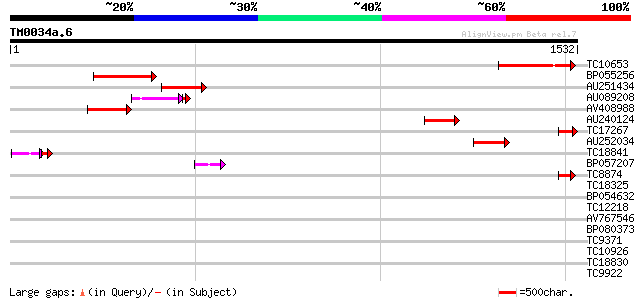
Query= TM0034a.6
(1532 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10653 similar to UP|O24517 (O24517) Unconventional myosin, par... 228 7e-60
BP055256 215 5e-56
AU251434 205 4e-53
AU089208 116 1e-26
AV408988 115 7e-26
AU240124 106 2e-23
TC17267 similar to PIR|T05200|T05200 myosin heavy chain F4I10.13... 105 5e-23
AU252034 102 4e-22
TC18841 50 1e-09
BP057207 51 2e-06
TC8874 similar to PIR|T00727|T00727 myosin heavy chain PCR43 - A... 45 1e-04
TC18325 36 0.040
BP054632 31 1.3
TC12218 31 1.7
AV767546 31 1.7
BP080373 30 2.2
TC9371 30 2.9
TC10926 similar to UP|CALX_SOYBN (Q39817) Calnexin homolog precu... 30 3.8
TC18830 weakly similar to UP|O40637 (O40637) Protease, partial (5%) 30 3.8
TC9922 29 4.9
>TC10653 similar to UP|O24517 (O24517) Unconventional myosin, partial (15%)
Length = 1173
Score = 228 bits (580), Expect = 7e-60
Identities = 114/207 (55%), Positives = 148/207 (71%)
Frame = +1
Query: 1321 GQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGEY 1380
G W +I+ L++L+ L N V I+K+ TQ FS+IN+ LFNSLLLRR+CCTFSNGEY
Sbjct: 58 GHWQSIIESLNNLLCTLKENFVAPVLIQKIFTQTFSYINVQLFNSLLLRRDCCTFSNGEY 237
Query: 1381 MKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTV 1440
+K+GLAELE W AKEEYAG+SW EL +IRQAVGFLVIHQK + S DEI DLCP ++V
Sbjct: 238 VKAGLAELELWCAQAKEEYAGSSWDELKHIRQAVGFLVIHQKYRISYDEIINDLCPIMSV 417
Query: 1441 RQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDI 1500
+Q+YRI T+YWD Y T+SVS +V+S MR ++ ++D+ N S+SFLLDD SIPFS ED+
Sbjct: 418 QQLYRICTLYWDANYNTRSVSPDVLSSMR-VLMAEDSNNAQSDSFLLDDSSSIPFSVEDL 594
Query: 1501 DMAIPAIDPDEIDLPAFVPEYSCAQFL 1527
A+ D ++ + E QFL
Sbjct: 595 STAMQEKDFSDLKPAEELLENPAFQFL 675
>BP055256
Length = 523
Score = 215 bits (547), Expect = 5e-56
Identities = 109/171 (63%), Positives = 128/171 (74%), Gaps = 1/171 (0%)
Frame = +3
Query: 226 GRISGAAIRTYLLERSRVVQITDPERNYHCFYQLCAFETDA-EKYELGHPSHFHYLNQSK 284
GRISGAAIRTYLLER Q++DPERNYHCFY LCA + +KY+LG+P FHYLNQS
Sbjct: 9 GRISGAAIRTYLLERFTCCQVSDPERNYHCFYMLCAAPQEVVQKYKLGNPRTFHYLNQSN 188
Query: 285 IYELDGVSNVEEYVRTRRAMNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIK 344
YEL+GV +EY T+RAM++VGIS E+QEAIFR +AA+LHLGNIEF+ GKE DSSV K
Sbjct: 189 XYELEGVDEFKEYCDTKRAMDVVGISSEEQEAIFRVVAAVLHLGNIEFTKGKEIDSSVPK 368
Query: 345 DEKSRFHMQMAANLFMCDVDLLLSTLCTRSIQTREGSIVKALDCNAAVAGR 395
DEKS FH+Q AA LFMCD L +LC R I TR+ +I K LD AA R
Sbjct: 369 DEKSWFHLQTAAELFMCDAKALEDSLCKRVIVTRDENITKWLDPEAAALSR 521
>AU251434
Length = 371
Score = 205 bits (522), Expect = 4e-53
Identities = 95/122 (77%), Positives = 107/122 (86%)
Frame = +1
Query: 409 WLVDKINRSVGQDINSQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKME 468
WLVDKIN S+GQD S+ IGVLDIYGFE F NSFEQFCIN NEKLQQHFN+HVFKME
Sbjct: 1 WLVDKINNSIGQDPTSKSLIGVLDIYGFESFNTNSFEQFCINLTNEKLQQHFNQHVFKME 180
Query: 469 QEEYGREEINWSYIEFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFP 528
QEEY +EEI+WSYIEFVDNQD+L+LIEKKP GI+ALLDEACMFP+STHETF+ KL+Q F
Sbjct: 181 QEEYKKEEIDWSYIEFVDNQDILDLIEKKPGGIIALLDEACMFPRSTHETFAQKLYQTFK 360
Query: 529 SH 530
+H
Sbjct: 361 NH 366
>AU089208
Length = 488
Score = 116 bits (291), Expect(2) = 1e-26
Identities = 62/142 (43%), Positives = 85/142 (59%), Gaps = 1/142 (0%)
Frame = +1
Query: 328 GNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDVDLLLSTLCTRSIQTREGSIVKALD 387
GN+ F+ + + + FH+ A L CD++ L L TR ++ R +IV+ L
Sbjct: 10 GNVSFTVIDNENHVQAVENEGLFHV---AKLIGCDIEXLKLALSTRKMKVRNENIVQKLT 180
Query: 388 CNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQD-INSQMQIGVLDIYGFECFKDNSFEQ 446
+ AV RD LAK++YA LFDWLV++IN S+ + I +LDIYGFE F N FE
Sbjct: 181 LSQAVDARDALAKSIYACLFDWLVEQINXSLAVGKXRTGRSIXILDIYGFESFNXNXFEX 360
Query: 447 FCINFANEKLQQHFNEHVFKME 468
FCI +ANE LQQHFN H+F +E
Sbjct: 361 FCIXYANEXLQQHFNRHLFXLE 426
Score = 21.6 bits (44), Expect(2) = 1e-26
Identities = 7/20 (35%), Positives = 13/20 (65%)
Frame = +2
Query: 468 EQEEYGREEINWSYIEFVDN 487
+ EEY + I+W+ + F D+
Sbjct: 425 KHEEYXQHGIDWAKLXFXDH 484
>AV408988
Length = 365
Score = 115 bits (287), Expect = 7e-26
Identities = 60/121 (49%), Positives = 78/121 (63%), Gaps = 2/121 (1%)
Frame = +1
Query: 210 NSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVVQITDPERNYHCFYQLCAFETDA--E 267
NSSRFGK +EI F G+ISGA I+T+LLE+SRVVQ + ER+YH FYQLCA + E
Sbjct: 1 NSSRFGKLIEIHFSETGKISGANIQTFLLEKSRVVQCNEGERSYHIFYQLCAGAPPSLRE 180
Query: 268 KYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRAMNIVGISHEDQEAIFRTLAAILHL 327
K L + YL QS Y + GV + EE+ A+++V IS DQ+ +F LAA+L L
Sbjct: 181 KLNLQSAQDYKYLRQSSCYSITGVDDAEEFHAVMEALDVVHISKGDQDNVFAMLAAVLWL 360
Query: 328 G 328
G
Sbjct: 361 G 363
>AU240124
Length = 300
Score = 106 bits (265), Expect = 2e-23
Identities = 50/95 (52%), Positives = 67/95 (69%)
Frame = +2
Query: 1121 LSRCIKENLGFKNGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVL 1180
L RCI ++LGF +P+AA IIYKCLLHW +FE ERT++FD II+ I ++ +D++DVL
Sbjct: 14 LIRCIAQHLGFSGNRPIAACIIYKCLLHWRSFEVERTSVFDRIIQTIGHAIETQDNNDVL 193
Query: 1181 PYWLSNTSALLCLLQRNLRSNGFLTTTGQRYTGSA 1215
YWLSN S LL LLQR L+++G QR S+
Sbjct: 194 AYWLSNASTLLLLLQRTLKASGAAGMAPQRRRSSS 298
>TC17267 similar to PIR|T05200|T05200 myosin heavy chain F4I10.130 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(3%)
Length = 513
Score = 105 bits (262), Expect = 5e-23
Identities = 50/50 (100%), Positives = 50/50 (100%)
Frame = +1
Query: 1483 NSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPAFVPEYSCAQFLNPIQT 1532
NSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPAFVPEYSCAQFLNPIQT
Sbjct: 1 NSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPAFVPEYSCAQFLNPIQT 150
>AU252034
Length = 305
Score = 102 bits (255), Expect = 4e-22
Identities = 50/97 (51%), Positives = 69/97 (70%)
Frame = +2
Query: 1254 VEARYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSG 1313
VEA+YPA+LFKQQLTA +EKI+G++RD+LKKE+SPLLGLCIQAP+T RQ K +
Sbjct: 14 VEAKYPALLFKQQLTAFLEKIYGMIRDNLKKEISPLLGLCIQAPRTSRQGLVKGRSHANA 193
Query: 1314 LPQQPSGGQWANIVNFLDSLMSKLHGNHVPSFFIRKL 1350
+ QQ W +IV L++ + + N+ P F +RK+
Sbjct: 194 VAQQALIAHWQSIVKSLNNYLKIMKANYAPPFLVRKV 304
>TC18841
Length = 568
Score = 50.4 bits (119), Expect(2) = 1e-09
Identities = 21/28 (75%), Positives = 24/28 (85%)
Frame = +2
Query: 87 LNDIYTYTGSILIAVNPFTKLPHLYDNH 114
LN IYTYTG+ILIA+ PF +LPHLYD H
Sbjct: 485 LNAIYTYTGNILIAITPFHRLPHLYDTH 568
Score = 30.8 bits (68), Expect(2) = 1e-09
Identities = 27/92 (29%), Positives = 39/92 (42%), Gaps = 3/92 (3%)
Frame = +1
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VWV+D W+ E S + GK++ ++ PR E G
Sbjct: 241 VGSHVWVEDPALTWIDGE-FHSKSAVKKSMFALGMGKQLSKISQRFFPR--IHEASPGRC 411
Query: 65 DMT--RLAYLNEPGVLYNLKRRY-TLNDIYTY 93
MT +PG L+NL RY T ++Y Y
Sbjct: 412 GMT*QNCHICMQPGSLHNLAARYATQCNLYIY 507
>BP057207
Length = 249
Score = 50.8 bits (120), Expect = 2e-06
Identities = 31/85 (36%), Positives = 44/85 (51%), Gaps = 1/85 (1%)
Frame = -1
Query: 499 IGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFAISHYAGKVTYHTDT 558
+G+++LLDE FP T TF+ KL QH S+ E+ + F +SHYAG+VTY T
Sbjct: 249 LGLLSLLDEESTFPNGTDLTFADKLKQHLNSNSCFKGER--EKAFTVSHYAGEVTYDTPD 76
Query: 559 FLDKNRDYVVVEH-CNLLSSSKCPF 582
+ Y + H +L PF
Sbjct: 75 SWRRTGTYCLRFHPTAILKGGPYPF 1
>TC8874 similar to PIR|T00727|T00727 myosin heavy chain PCR43 - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (3%)
Length = 715
Score = 44.7 bits (104), Expect = 1e-04
Identities = 22/46 (47%), Positives = 29/46 (62%)
Frame = +2
Query: 1482 SNSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPAFVPEYSCAQFL 1527
S SFLLDDD SIPFS +DI +I ++ ++D P + E S FL
Sbjct: 5 STSFLLDDDSSIPFSVDDISKSIQQVEVADVDPPPLIRENSGFGFL 142
>TC18325
Length = 513
Score = 36.2 bits (82), Expect = 0.040
Identities = 26/68 (38%), Positives = 40/68 (58%), Gaps = 3/68 (4%)
Frame = +2
Query: 1463 EVVSEMREIVSSKDNQNITSN-SFLLDDDMSIPFSAEDI--DMAIPAIDPDEIDLPAFVP 1519
EV+S MR ++ ++D+ NI N SFLL+ D SIPF E++ M+ + ++D P +
Sbjct: 2 EVISRMR-VLMTEDSDNIPPNHSFLLEVDSSIPFLMEEMFRSMSDIRLSDMDVDPPPILR 178
Query: 1520 EYSCAQFL 1527
S QFL
Sbjct: 179 LRSDFQFL 202
>BP054632
Length = 514
Score = 31.2 bits (69), Expect = 1.3
Identities = 19/52 (36%), Positives = 31/52 (59%), Gaps = 1/52 (1%)
Frame = +3
Query: 1026 LEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSN-RQGFAKSLSERYSNAVA 1076
LE VK+ EEK+ +LEDE + L +K +A + N ++G K ++ AV+
Sbjct: 330 LEDQVKTYEEKVQTLEDEINELNEKLSAANSEINTKEGLVKQHAKVAEEAVS 485
>TC12218
Length = 737
Score = 30.8 bits (68), Expect = 1.7
Identities = 31/108 (28%), Positives = 48/108 (43%), Gaps = 7/108 (6%)
Frame = +2
Query: 908 KKIRVSNEDAKQIEISKL-----QKMIEALNLELDAAKLATI--NECNKNAVLQNQFELS 960
+K+R + + +E+S + Q + + DA ++ T NE K + N +
Sbjct: 290 EKLRRDKLNERFLELSSILEPGRQPKTDKAAIISDAVRVVTQLRNEAEKLKEMNNDLQEK 469
Query: 961 IKEKSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKG 1008
IKE A K +EIR E LK+ + EKK V+L N Q G
Sbjct: 470 IKELKAEK------NEIRDEKNKLKLDKEKLEKK-----VKLRNVQPG 580
>AV767546
Length = 589
Score = 30.8 bits (68), Expect = 1.7
Identities = 17/61 (27%), Positives = 33/61 (53%), Gaps = 2/61 (3%)
Frame = +3
Query: 893 LEKQLDELTWRLHLEKKIRVSNEDAKQI--EISKLQKMIEALNLELDAAKLATINECNKN 950
+E L + TW +KKI E+ K + I+++QK++ + L+ + LA + ++N
Sbjct: 402 VENALAKFTWAKEAQKKIEKLKEEGKPMPTSIAEVQKLVGSSPLDHARSNLAQSGQISRN 581
Query: 951 A 951
A
Sbjct: 582 A 584
>BP080373
Length = 401
Score = 30.4 bits (67), Expect = 2.2
Identities = 17/42 (40%), Positives = 19/42 (44%)
Frame = +2
Query: 71 YLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYD 112
Y N P N Y LNDIYTY + I F L H Y+
Sbjct: 122 YKNLPADTTNHLSHYKLNDIYTYQPTCCIYFVLFVILHHFYE 247
>TC9371
Length = 661
Score = 30.0 bits (66), Expect = 2.9
Identities = 26/90 (28%), Positives = 40/90 (43%), Gaps = 1/90 (1%)
Frame = +2
Query: 959 LSIKEKSALKRELV-AVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLR 1017
L + EKS R A D ++KEN MLK ++ K + A K + E
Sbjct: 17 LEVLEKSISARARAEAADVLQKENLMLKERIEVLIKDKNSFTY----AFKYQRERYSDYE 184
Query: 1018 EFEQKCSQLEQNVKSLEEKMLSLEDENHVL 1047
E Q+ L+ V +E++ +LE N+ L
Sbjct: 185 EKVQELQHLKPLVSQYQEQIRTLEVNNYAL 274
>TC10926 similar to UP|CALX_SOYBN (Q39817) Calnexin homolog precursor, partial
(31%)
Length = 629
Score = 29.6 bits (65), Expect = 3.8
Identities = 17/44 (38%), Positives = 23/44 (51%), Gaps = 4/44 (9%)
Frame = +1
Query: 1390 KWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRK----KSLDE 1429
+WIV+ K+EY G H+ + G LV + RK K LDE
Sbjct: 166 RWIVSQKDEYNGAWKHDKSEGHDDYGLLVSEKARKYAIVKELDE 297
>TC18830 weakly similar to UP|O40637 (O40637) Protease, partial (5%)
Length = 850
Score = 29.6 bits (65), Expect = 3.8
Identities = 47/196 (23%), Positives = 82/196 (40%), Gaps = 20/196 (10%)
Frame = +2
Query: 905 HLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELSIKEK 964
HL + + S + E L NL A + +++ N+++ +++ SI+
Sbjct: 68 HLTEGVEASYGNENIGEDDDLLPENSVTNLSWGATEEKSVD--NQSSNMEDPLIESIRSL 241
Query: 965 SALKRELVAVDEIRKENAMLKVSLDAFEKKCTA-----------LEVELINAQKGRDETI 1013
A++ L + +E + VS D KC++ L ++A G +E
Sbjct: 242 LAVQEALEEEVQKFRETGIEVVSPDDDLAKCSSASAGTIGVDIGLHNSCVSAHSGAEEIK 421
Query: 1014 EKLRE-FEQKCSQLEQNVKSLEEK------MLSLEDENHVLRQKALSAPLKSNRQGFAKS 1066
+ E + L QN+ SLE K M++L+D V + ALS+ KS ++ A +
Sbjct: 422 QTASSSLELQVLNLTQNISSLENKLKELQGMVALKDSRIVELETALSSG-KSPKEESAST 598
Query: 1067 --LSERYSNAVASRTE 1080
LSE V S E
Sbjct: 599 IGLSEEICKEVESEVE 646
>TC9922
Length = 528
Score = 29.3 bits (64), Expect = 4.9
Identities = 13/34 (38%), Positives = 20/34 (58%), Gaps = 2/34 (5%)
Frame = -2
Query: 253 YHCFYQL--CAFETDAEKYELGHPSHFHYLNQSK 284
+HCF+ L C++ HPSH+H L+Q+K
Sbjct: 128 FHCFHDLIVCSY----------HPSHYHLLSQAK 57
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.135 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,453,509
Number of Sequences: 28460
Number of extensions: 328481
Number of successful extensions: 1723
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 1704
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1717
length of query: 1532
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1430
effective length of database: 1,994,680
effective search space: 2852392400
effective search space used: 2852392400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0034a.6


