

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0033b.7
(715 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
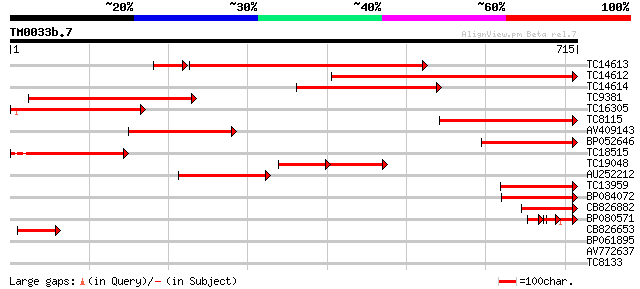
Sequences producing significant alignments: (bits) Value
TC14613 homologue to UP|PAL2_PEA (Q04593) Phenylalanine ammonia-... 577 0.0
TC14612 similar to UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-... 596 e-171
TC14614 homologue to UP|PAL1_SOYBN (P27991) Phenylalanine ammoni... 355 2e-98
TC9381 similar to UP|PALY_STYHU (P45732) Phenylalanine ammonia-l... 351 3e-97
TC16305 similar to UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-... 302 1e-82
TC8115 similar to PIR|A24727|A24727 phenylalanine ammonia-lyase ... 293 8e-80
AV409143 253 6e-68
BP052646 250 5e-67
TC18515 homologue to UP|PAL1_SOYBN (P27991) Phenylalanine ammoni... 244 3e-65
TC19048 homologue to PIR|A24727|A24727 phenylalanine ammonia-lya... 132 2e-63
AU252212 202 2e-52
TC13959 similar to UP|PALY_MEDSA (P27990) Phenylalanine ammonia-... 194 4e-50
BP084072 191 3e-49
CB826882 152 1e-37
BP080571 56 5e-16
CB826653 71 7e-13
BP061895 33 0.001
AV772637 28 5.1
TC8133 similar to GB|BAA97157.1|8809606|AB018117 ethylene respon... 27 8.7
>TC14613 homologue to UP|PAL2_PEA (Q04593) Phenylalanine ammonia-lyase 2 ,
partial (47%)
Length = 1048
Score = 577 bits (1487), Expect(2) = 0.0
Identities = 293/301 (97%), Positives = 298/301 (98%)
Frame = +3
Query: 227 SGEVVNAKDAFQLAGIDSGFFELQPKEGLALVNGTAVGSGLASIVLFDANILAILAEVLS 286
SG+V+NAK+AFQ AGIDSGFFELQPKEGLALVNGTAVGSGLASIVLFDANILAIL+EVLS
Sbjct: 144 SGKVLNAKEAFQSAGIDSGFFELQPKEGLALVNGTAVGSGLASIVLFDANILAILSEVLS 323
Query: 287 AIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHEVDPLQKPK 346
AIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHEVDPLQKPK
Sbjct: 324 AIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHEVDPLQKPK 503
Query: 347 QDRYALRTSPQWLGPLIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGV 406
QDRYALRTSPQWLGPLIEVIRFSTKSIER+INSVNDNPLIDVSRNKALHGGNFQGTPIGV
Sbjct: 504 QDRYALRTSPQWLGPLIEVIRFSTKSIERDINSVNDNPLIDVSRNKALHGGNFQGTPIGV 683
Query: 407 SMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCS 466
SMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSLDYG KGAEIAMASYCS
Sbjct: 684 SMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSLDYGLKGAEIAMASYCS 863
Query: 467 ELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRH 526
ELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKT EAIEILKLMSSTFLIALCQAIDLRH
Sbjct: 864 ELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTYEAIEILKLMSSTFLIALCQAIDLRH 1043
Query: 527 L 527
L
Sbjct: 1044L 1046
Score = 81.3 bits (199), Expect(2) = 0.0
Identities = 39/43 (90%), Positives = 41/43 (94%)
Frame = +1
Query: 182 TKLINNNITPCLPLRGTVTASGDLVPLSYIAGLLTGRPNSKAV 224
TKLINNNITPCLPLRGTVTASGDLVPLSYIAGLLTGR NS ++
Sbjct: 1 TKLINNNITPCLPLRGTVTASGDLVPLSYIAGLLTGRQNSXSL 129
>TC14612 similar to UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1 ,
partial (43%)
Length = 1049
Score = 596 bits (1537), Expect = e-171
Identities = 299/309 (96%), Positives = 305/309 (97%)
Frame = +3
Query: 407 SMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCS 466
SMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSL YGFKGAEIAMASYCS
Sbjct: 3 SMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSLYYGFKGAEIAMASYCS 182
Query: 467 ELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRH 526
ELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDL+H
Sbjct: 183 ELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLKH 362
Query: 527 LEENLKYSVKSTVSQVAKRTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATY 586
LEENLKYSV+STVSQVA+RTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATY
Sbjct: 363 LEENLKYSVQSTVSQVAERTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATY 542
Query: 587 PLMQKLRQVLVDHALVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPA 646
PLMQKLRQVLV HALVNGE EKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPA
Sbjct: 543 PLMQKLRQVLVXHALVNGENEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPA 722
Query: 647 MPNKINECRSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGE 706
+PNKINECRSYPLYKFVR+ELGTELLTGEKTRSPGEE DKLFTAICQGKIIDPL+ECLGE
Sbjct: 723 IPNKINECRSYPLYKFVREELGTELLTGEKTRSPGEEFDKLFTAICQGKIIDPLMECLGE 902
Query: 707 WNGAPLPIC 715
WNGAPLPIC
Sbjct: 903 WNGAPLPIC 929
>TC14614 homologue to UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1
, partial (26%)
Length = 550
Score = 355 bits (910), Expect = 2e-98
Identities = 181/183 (98%), Positives = 182/183 (98%)
Frame = +2
Query: 362 LIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKL 421
LIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKL
Sbjct: 2 LIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKL 181
Query: 422 MFAQFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQS 481
MFAQFTELVDDHYNNGLPSNLTASRNPSLDYG KGAEIAMASYCSELQYLANPVTTHVQS
Sbjct: 182 MFAQFTELVDDHYNNGLPSNLTASRNPSLDYGLKGAEIAMASYCSELQYLANPVTTHVQS 361
Query: 482 AEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQ 541
AEQHNQDVNSLGLISSRKTNEA+EILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQ
Sbjct: 362 AEQHNQDVNSLGLISSRKTNEAVEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQ 541
Query: 542 VAK 544
VAK
Sbjct: 542 VAK 550
>TC9381 similar to UP|PALY_STYHU (P45732) Phenylalanine ammonia-lyase ,
partial (30%)
Length = 997
Score = 351 bits (900), Expect = 3e-97
Identities = 178/213 (83%), Positives = 195/213 (90%), Gaps = 1/213 (0%)
Frame = +2
Query: 24 DPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAANHQG-VSVELC 82
DPL+WG AAE+MKGSHLDEVKRMV EYRK VV+LGGETLTIAQVAA+AA+ QG V+VEL
Sbjct: 230 DPLNWGAAAEAMKGSHLDEVKRMVEEYRKPVVRLGGETLTIAQVAAVAAHDQGAVTVELS 409
Query: 83 ESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFLNAGIFGN 142
ES+RAGV+AS +WV +SMN GTDSYGVTTGFGATSHRRTK G ALQ ELIRFLNAGIFGN
Sbjct: 410 ESSRAGVEASRNWVRDSMNKGTDSYGVTTGFGATSHRRTKQGGALQKELIRFLNAGIFGN 589
Query: 143 GTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLPLRGTVTAS 202
GTES TLP ATRAAMLVRINTLLQGYSGIRFEILEAITKLIN+N+ PCLPLRGT+TAS
Sbjct: 590 GTESNCTLPHTATRAAMLVRINTLLQGYSGIRFEILEAITKLINSNVAPCLPLRGTITAS 769
Query: 203 GDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKD 235
GDLVPLSYIAGLLTGRPNSKAVGPSGE++ A++
Sbjct: 770 GDLVPLSYIAGLLTGRPNSKAVGPSGEILTAQE 868
>TC16305 similar to UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1 ,
partial (21%)
Length = 637
Score = 302 bits (774), Expect = 1e-82
Identities = 157/174 (90%), Positives = 160/174 (91%), Gaps = 3/174 (1%)
Frame = +1
Query: 1 MAPNTNA---TQNGSIFCNTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKL 57
MAP TN +QN SIFC K A +DPLSWGVAAESMKGSHLDEVKRMVAEYRK VVKL
Sbjct: 118 MAPTTNGHHQSQNDSIFCTAAK-AGSDPLSWGVAAESMKGSHLDEVKRMVAEYRKPVVKL 294
Query: 58 GGETLTIAQVAAIAANHQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATS 117
GGETLTIAQVAAIAAN QGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATS
Sbjct: 295 GGETLTIAQVAAIAANDQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATS 474
Query: 118 HRRTKNGNALQLELIRFLNAGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYS 171
HRRTKNGNALQLELIRFLNAGIFGNGTE+ HTLPQPATRAAMLVRINTLLQGYS
Sbjct: 475 HRRTKNGNALQLELIRFLNAGIFGNGTETNHTLPQPATRAAMLVRINTLLQGYS 636
>TC8115 similar to PIR|A24727|A24727 phenylalanine ammonia-lyase - kidney
bean (fragment) {Phaseolus vulgaris;} ,
partial (34%)
Length = 760
Score = 293 bits (749), Expect = 8e-80
Identities = 140/173 (80%), Positives = 154/173 (88%)
Frame = +2
Query: 543 AKRTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALV 602
A+ TLTTGVNGELHPSRFCEKDLL VVDRE +FAYIDDPC TYPLMQKLRQ LVDHAL
Sbjct: 8 ARGTLTTGVNGELHPSRFCEKDLLTVVDREDVFAYIDDPCSGTYPLMQKLRQSLVDHALE 187
Query: 603 NGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKF 662
N + EK+ TSIFQKIA FEDELK++LPKEVESAR AY++G A+PNKI ECRSYPLYKF
Sbjct: 188 NADAEKNVNTSIFQKIAIFEDELKAILPKEVESARVAYDNGESAIPNKIKECRSYPLYKF 367
Query: 663 VRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
VR+ELGT LLTGEK +SPGE+ DKLFTA+CQGKIIDP+LECLGEWNGAPLPIC
Sbjct: 368 VREELGTGLLTGEKVKSPGEDFDKLFTAMCQGKIIDPILECLGEWNGAPLPIC 526
>AV409143
Length = 415
Score = 253 bits (647), Expect = 6e-68
Identities = 129/137 (94%), Positives = 136/137 (99%)
Frame = +1
Query: 150 LPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLPLRGTVTASGDLVPLS 209
LPQPATRAAMLVRINTLLQGYSGIRFEILEAITKL+NNNITPCLPLRGTVTASGDLVPLS
Sbjct: 4 LPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLLNNNITPCLPLRGTVTASGDLVPLS 183
Query: 210 YIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFELQPKEGLALVNGTAVGSGLAS 269
YIAGLLTGRPNSKAVGPSGEV+NAK+AF+LA IDSGFFELQPKEGLALVNGTAVGSGLAS
Sbjct: 184 YIAGLLTGRPNSKAVGPSGEVLNAKEAFELASIDSGFFELQPKEGLALVNGTAVGSGLAS 363
Query: 270 IVLFDANILAILAEVLS 286
IVLF+ANILA+L+EVLS
Sbjct: 364 IVLFEANILAVLSEVLS 414
>BP052646
Length = 514
Score = 250 bits (639), Expect = 5e-67
Identities = 121/121 (100%), Positives = 121/121 (100%)
Frame = -1
Query: 595 VLVDHALVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINEC 654
VLVDHALVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINEC
Sbjct: 514 VLVDHALVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINEC 335
Query: 655 RSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPI 714
RSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPI
Sbjct: 334 RSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPI 155
Query: 715 C 715
C
Sbjct: 154 C 152
>TC18515 homologue to UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1
, partial (19%)
Length = 575
Score = 244 bits (623), Expect = 3e-65
Identities = 127/152 (83%), Positives = 135/152 (88%), Gaps = 2/152 (1%)
Frame = +3
Query: 1 MAPNTNATQNGSIFCNTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGE 60
MA TN+ QN S+FC +DPL+WGVAAESMKGSHLDEVKRMVAEYRK VV+LGGE
Sbjct: 129 MAATTNS-QNSSLFCTA--KGGSDPLNWGVAAESMKGSHLDEVKRMVAEYRKPVVRLGGE 299
Query: 61 TLTIAQVAAIAAN--HQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSH 118
TLTIAQVAA+AA+ H GVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSH
Sbjct: 300 TLTIAQVAAVAAHDQHNGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSH 479
Query: 119 RRTKNGNALQLELIRFLNAGIFGNGTESTHTL 150
RRTK GNALQ ELIRFLNAGIFGNGTES+HTL
Sbjct: 480 RRTKQGNALQQELIRFLNAGIFGNGTESSHTL 575
>TC19048 homologue to PIR|A24727|A24727 phenylalanine ammonia-lyase -
kidney bean (fragment) {Phaseolus
vulgaris;} , partial (27%)
Length = 414
Score = 132 bits (332), Expect(2) = 2e-63
Identities = 64/78 (82%), Positives = 71/78 (90%)
Frame = +1
Query: 399 FQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAE 458
F+ P+ V+MDNTRLALA+IGKL+FAQF+ELV+D YNNG PSNL SRNPSLDYGFKGAE
Sbjct: 181 FKEPPLRVAMDNTRLALASIGKLLFAQFSELVNDFYNNGXPSNLYGSRNPSLDYGFKGAE 360
Query: 459 IAMASYCSELQYLANPVT 476
IAMASYCSELQYLANPVT
Sbjct: 361 IAMASYCSELQYLANPVT 414
Score = 128 bits (321), Expect(2) = 2e-63
Identities = 61/65 (93%), Positives = 63/65 (96%)
Frame = +3
Query: 340 DPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNF 399
DPLQKPKQDRYALRTS QWLGPLIEV RFSTKSIERE+NSVNDNPLIDV+RNKALHGGNF
Sbjct: 3 DPLQKPKQDRYALRTSRQWLGPLIEVSRFSTKSIERELNSVNDNPLIDVARNKALHGGNF 182
Query: 400 QGTPI 404
QGTPI
Sbjct: 183 QGTPI 197
>AU252212
Length = 350
Score = 202 bits (513), Expect = 2e-52
Identities = 99/116 (85%), Positives = 110/116 (94%)
Frame = +3
Query: 214 LLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFELQPKEGLALVNGTAVGSGLASIVLF 273
LLTGRPNSKAVGPSGE++ AK+AF+LA I FFEL+PKEGLALVNGTAVGSGLA+IVLF
Sbjct: 3 LLTGRPNSKAVGPSGEILTAKEAFELANIGHDFFELEPKEGLALVNGTAVGSGLAAIVLF 182
Query: 274 DANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSY 329
+ANILA+L+E++SAIFAEVM GKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSY
Sbjct: 183 EANILAVLSELISAIFAEVMLGKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSY 350
>TC13959 similar to UP|PALY_MEDSA (P27990) Phenylalanine ammonia-lyase ,
partial (13%)
Length = 494
Score = 194 bits (493), Expect = 4e-50
Identities = 91/96 (94%), Positives = 94/96 (97%)
Frame = +1
Query: 620 TFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTGEKTRS 679
TFEDELKSLLP EVESARAAYESGNP +PNKINECRSYPLYKFVR+ELGTELLTGEKTRS
Sbjct: 1 TFEDELKSLLPXEVESARAAYESGNPTIPNKINECRSYPLYKFVREELGTELLTGEKTRS 180
Query: 680 PGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
PGEECD+LFTAICQGKIIDPLLECLGEWNGAPLPIC
Sbjct: 181 PGEECDQLFTAICQGKIIDPLLECLGEWNGAPLPIC 288
>BP084072
Length = 457
Score = 191 bits (486), Expect = 3e-49
Identities = 90/95 (94%), Positives = 93/95 (97%)
Frame = -1
Query: 621 FEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTGEKTRSP 680
FEDELKSLLPKEVESARAAYESGNP + NKINECRSYPLYKFVR+ELGTELLTGEK+RSP
Sbjct: 457 FEDELKSLLPKEVESARAAYESGNPTISNKINECRSYPLYKFVREELGTELLTGEKSRSP 278
Query: 681 GEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
GEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC
Sbjct: 277 GEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 173
>CB826882
Length = 364
Score = 152 bits (385), Expect = 1e-37
Identities = 70/70 (100%), Positives = 70/70 (100%)
Frame = +2
Query: 646 AMPNKINECRSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLG 705
AMPNKINECRSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLG
Sbjct: 2 AMPNKINECRSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLG 181
Query: 706 EWNGAPLPIC 715
EWNGAPLPIC
Sbjct: 182 EWNGAPLPIC 211
>BP080571
Length = 338
Score = 55.8 bits (133), Expect(2) = 5e-16
Identities = 27/41 (65%), Positives = 29/41 (69%), Gaps = 3/41 (7%)
Frame = -2
Query: 678 RSPGEECDKLFTAICQ---GKIIDPLLECLGEWNGAPLPIC 715
+SPG + L Q GKIIDPLLECLGEWNGAPLPIC
Sbjct: 268 KSPGHQVRSLTNCSQQYATGKIIDPLLECLGEWNGAPLPIC 146
Score = 45.8 bits (107), Expect(2) = 5e-16
Identities = 20/21 (95%), Positives = 21/21 (99%)
Frame = -3
Query: 653 ECRSYPLYKFVRKELGTELLT 673
ECRSYPLYKFVR+ELGTELLT
Sbjct: 336 ECRSYPLYKFVREELGTELLT 274
Score = 41.6 bits (96), Expect = 4e-04
Identities = 18/21 (85%), Positives = 19/21 (89%)
Frame = -1
Query: 674 GEKTRSPGEECDKLFTAICQG 694
GEK+RSPGEE DKLFTAIC G
Sbjct: 272 GEKSRSPGEEFDKLFTAICHG 210
>CB826653
Length = 450
Score = 70.9 bits (172), Expect = 7e-13
Identities = 35/55 (63%), Positives = 41/55 (73%)
Frame = +2
Query: 10 NGSIFCNTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTI 64
NG NT DPL+WG AAE+++GSHLDEVKRMV EYRK +V LGG+TLTI
Sbjct: 284 NGGGKSNTHHHHHDDPLNWGAAAEALRGSHLDEVKRMVEEYRKPLVMLGGKTLTI 448
>BP061895
Length = 246
Score = 32.7 bits (73), Expect(2) = 0.001
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = -3
Query: 704 LGEWNGAPLPIC 715
LGEWNGAPLPIC
Sbjct: 190 LGEWNGAPLPIC 155
Score = 26.2 bits (56), Expect(2) = 0.001
Identities = 11/15 (73%), Positives = 11/15 (73%)
Frame = -2
Query: 689 TAICQGKIIDPLLEC 703
T GKIIDPLLEC
Sbjct: 236 TQYATGKIIDPLLEC 192
>AV772637
Length = 551
Score = 28.1 bits (61), Expect = 5.1
Identities = 13/30 (43%), Positives = 20/30 (66%)
Frame = -1
Query: 323 ILDGSSYMKAAKKLHEVDPLQKPKQDRYAL 352
+ DGS Y++A K+ H +PL P +DR+ L
Sbjct: 521 VADGSHYIQALKQTH-YEPLVPPVRDRHLL 435
>TC8133 similar to GB|BAA97157.1|8809606|AB018117 ethylene responsive
element binding factor 5 (ATERF5) {Arabidopsis
thaliana;} , partial (22%)
Length = 785
Score = 27.3 bits (59), Expect = 8.7
Identities = 20/59 (33%), Positives = 27/59 (44%)
Frame = -3
Query: 223 AVGPSGEVVNAKDAFQLAGIDSGFFELQPKEGLALVNGTAVGSGLASIVLFDANILAIL 281
A G G VVN G D G +PK ALV +G LA L D++++ +L
Sbjct: 753 AKGKRGSVVN------FGGFDGGVECSEPKSRSALVRVPDLGGELAPRSLADSSVVLLL 595
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,320,851
Number of Sequences: 28460
Number of extensions: 120151
Number of successful extensions: 563
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 560
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 561
length of query: 715
length of database: 4,897,600
effective HSP length: 97
effective length of query: 618
effective length of database: 2,136,980
effective search space: 1320653640
effective search space used: 1320653640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0033b.7


