

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0033b.5
(88 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
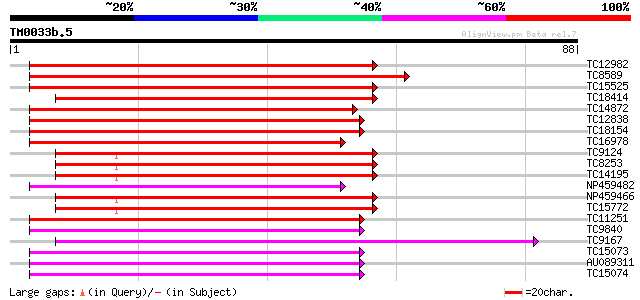
TC12982 weakly similar to UP|Q9AWA1 (Q9AWA1) GTP binding protein... 89 1e-19
TC8589 UP|Q40210 (Q40210) RAB5B, complete 72 2e-14
TC15525 UP|Q40209 (Q40209) RAB5A, complete 70 5e-14
TC18414 weakly similar to UP|RAB5_TOBAC (P29687) Ras-related pro... 65 2e-12
TC14872 UP|Q40208 (Q40208) RAB2A, complete 54 4e-09
TC12838 UP|Q40196 (Q40196) RAB11F, complete 54 6e-09
TC18154 UP|Q40192 (Q40192) RAB11B, complete 48 3e-07
TC16978 UP|R11A_LOTJA (Q40191) Ras-related protein Rab11A, complete 45 2e-06
TC9124 homologue to UP|Q41024 (Q41024) Small GTP-binding protein... 45 2e-06
TC8253 UP|Q40219 (Q40219) RAB8E, complete 45 3e-06
TC14195 UP|Q40215 (Q40215) RAB8A, complete 45 3e-06
NP459482 RAB11J [Lotus japonicus] 45 3e-06
NP459466 RAB8C [Lotus japonicus] 45 3e-06
TC15772 UP|Q40218 (Q40218) RAB8D, complete 45 3e-06
TC11251 UP|R11E_LOTJA (Q40195) Ras-related protein Rab11E, complete 45 3e-06
TC9840 UP|R11D_LOTJA (Q40194) Ras-related protein Rab11D, complete 44 4e-06
TC9167 UP|Q40202 (Q40202) RAB1B (Fragment), complete 44 5e-06
TC15073 UP|Q40199 (Q40199) RAB11I (Fragment), complete 44 7e-06
AU089311 43 9e-06
TC15074 similar to UP|R11B_TOBAC (Q40521) Ras-related protein Ra... 41 3e-05
>TC12982 weakly similar to UP|Q9AWA1 (Q9AWA1) GTP binding protein
(Fragment), partial (54%)
Length = 526
Score = 89.4 bits (220), Expect = 1e-19
Identities = 43/54 (79%), Positives = 48/54 (88%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LVMAL AN+SDLEPKREVE EEG +FA ENGMFYMETSAKT ENIN+LF+EI +
Sbjct: 136 LVMALAANQSDLEPKREVETEEGEKFAPENGMFYMETSAKTAENINELFYEIAK 297
>TC8589 UP|Q40210 (Q40210) RAB5B, complete
Length = 1241
Score = 71.6 bits (174), Expect = 2e-14
Identities = 33/59 (55%), Positives = 46/59 (77%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESP 62
+VMALV NK+DL KREV +++G +A++NGMF++ETSAKT +NIN+LF EI + P
Sbjct: 640 IVMALVGNKADLLEKREVAVQDGIDYAEKNGMFFIETSAKTADNINELFEEIAKRLPRP 816
>TC15525 UP|Q40209 (Q40209) RAB5A, complete
Length = 1176
Score = 70.5 bits (171), Expect = 5e-14
Identities = 31/54 (57%), Positives = 42/54 (77%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMA NKSDLE KR+V +E +A+ENG+F+METSAKT N+ND+F+EI +
Sbjct: 599 MVMAFAGNKSDLEDKRKVTADEARVYAEENGLFFMETSAKTAANVNDVFYEIAK 760
>TC18414 weakly similar to UP|RAB5_TOBAC (P29687) Ras-related protein
Rab5, partial (27%)
Length = 609
Score = 65.1 bits (157), Expect = 2e-12
Identities = 29/50 (58%), Positives = 41/50 (82%)
Frame = +2
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LVANK+DLE +R+V E+G ++AQENGM + ETSAKT N+++LF+EI +
Sbjct: 5 LVANKADLEDERKVRNEDGEEYAQENGMTFFETSAKTAPNVHELFYEIAK 154
>TC14872 UP|Q40208 (Q40208) RAB2A, complete
Length = 1125
Score = 54.3 bits (129), Expect = 4e-09
Identities = 23/51 (45%), Positives = 35/51 (68%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFE 54
+ + L+ NK DL +R V EEG QFA+E+G+ +ME SAKT +N+ + F +
Sbjct: 540 MTIMLIGNKCDLAHRRAVSTEEGEQFAKEHGLIFMEASAKTAQNVEEAFIK 692
>TC12838 UP|Q40196 (Q40196) RAB11F, complete
Length = 1099
Score = 53.5 bits (127), Expect = 6e-09
Identities = 27/52 (51%), Positives = 36/52 (68%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LVANKSDL REVE EEG +FA+ G+ +METSA N+ + F E+
Sbjct: 464 MVVVLVANKSDLVQSREVEKEEGEEFAEIEGLCFMETSALQNLNVEEAFLEM 619
>TC18154 UP|Q40192 (Q40192) RAB11B, complete
Length = 1183
Score = 48.1 bits (113), Expect = 3e-07
Identities = 22/52 (42%), Positives = 34/52 (65%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ L+ NK DL +R V E+ +FA+E G+F+ ETSA T EN+ F ++
Sbjct: 686 IVIMLIGNKGDLVDQRVVLSEDAVEFAEEQGLFFSETSALTGENVESAFLKL 841
>TC16978 UP|R11A_LOTJA (Q40191) Ras-related protein Rab11A, complete
Length = 984
Score = 45.4 bits (106), Expect = 2e-06
Identities = 19/49 (38%), Positives = 32/49 (64%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
+V+ L+ NK DL +R+V E+ +FA++ G+F++ETSA N+ F
Sbjct: 428 IVIILIGNKCDLVNQRDVPTEDAKEFAEKEGLFFLETSALEATNVESAF 574
>TC9124 homologue to UP|Q41024 (Q41024) Small GTP-binding protein, complete
Length = 1109
Score = 45.1 bits (105), Expect = 2e-06
Identities = 24/51 (47%), Positives = 33/51 (64%), Gaps = 1/51 (1%)
Frame = +3
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N++++FF I R
Sbjct: 624 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNLNVDEVFFSIAR 776
>TC8253 UP|Q40219 (Q40219) RAB8E, complete
Length = 1106
Score = 44.7 bits (104), Expect = 3e-06
Identities = 24/51 (47%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +3
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF I R
Sbjct: 555 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNLNVEEVFFSIAR 707
>TC14195 UP|Q40215 (Q40215) RAB8A, complete
Length = 1171
Score = 44.7 bits (104), Expect = 3e-06
Identities = 24/51 (47%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +2
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF I R
Sbjct: 575 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNLNVEEVFFSIAR 727
>NP459482 RAB11J [Lotus japonicus]
Length = 672
Score = 44.7 bits (104), Expect = 3e-06
Identities = 23/49 (46%), Positives = 29/49 (58%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
+V LV NKSDL+ REV EG A+ G+F+METSA N+ F
Sbjct: 358 VVTILVGNKSDLKDAREVTTAEGKALAEAQGLFFMETSALDSSNVAAAF 504
>NP459466 RAB8C [Lotus japonicus]
Length = 639
Score = 44.7 bits (104), Expect = 3e-06
Identities = 24/51 (47%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +1
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF I R
Sbjct: 373 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNLNVEEVFFSIAR 525
>TC15772 UP|Q40218 (Q40218) RAB8D, complete
Length = 1139
Score = 44.7 bits (104), Expect = 3e-06
Identities = 24/51 (47%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +2
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF I R
Sbjct: 611 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNLNVEEVFFSIAR 763
>TC11251 UP|R11E_LOTJA (Q40195) Ras-related protein Rab11E, complete
Length = 950
Score = 44.7 bits (104), Expect = 3e-06
Identities = 20/52 (38%), Positives = 32/52 (61%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ L+ NKSDL V E+G FA++ +++METSA N+ + F E+
Sbjct: 416 IVVMLIGNKSDLRHLVTVSTEDGKAFAEKESLYFMETSALEATNVENAFSEV 571
>TC9840 UP|R11D_LOTJA (Q40194) Ras-related protein Rab11D, complete
Length = 1302
Score = 44.3 bits (103), Expect = 4e-06
Identities = 20/52 (38%), Positives = 31/52 (59%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ L+ NKSDL V E+G FA+ +++METSA N+ + F E+
Sbjct: 602 IVVMLIGNKSDLRHLVAVPTEDGKSFAERESLYFMETSALEATNVENAFTEV 757
>TC9167 UP|Q40202 (Q40202) RAB1B (Fragment), complete
Length = 951
Score = 43.9 bits (102), Expect = 5e-06
Identities = 26/75 (34%), Positives = 34/75 (44%)
Frame = +2
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPGRSSY 67
LV NKSDL R V E FA E G+ +METSAK N+ F + ++ S
Sbjct: 443 LVGNKSDLTANRAVSYETAKAFADEIGIPFMETSAKDATNVEQAFMAMAASIKNRMASQP 622
Query: 68 APTKEWASAGLSSMP 82
A + +S P
Sbjct: 623 ANNARPPTVNISGKP 667
>TC15073 UP|Q40199 (Q40199) RAB11I (Fragment), complete
Length = 733
Score = 43.5 bits (101), Expect = 7e-06
Identities = 21/52 (40%), Positives = 31/52 (59%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LV NK+DL R V EE FA+ ++METSA N+++ F E+
Sbjct: 202 IVIMLVGNKADLRHLRAVPTEEATAFAERENTYFMETSALESLNVDNAFIEV 357
>AU089311
Length = 344
Score = 43.1 bits (100), Expect = 9e-06
Identities = 21/52 (40%), Positives = 29/52 (55%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ L+ NK DL R V E+ +FAQ +F+METSA N+ F I
Sbjct: 26 IVIMLIGNKCDLGSLRAVPTEDAEEFAQRENLFFMETSALESTNVETCFLTI 181
>TC15074 similar to UP|R11B_TOBAC (Q40521) Ras-related protein Rab11B,
partial (89%)
Length = 973
Score = 41.2 bits (95), Expect = 3e-05
Identities = 20/52 (38%), Positives = 30/52 (57%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LV NK+DL R V EE FA+ ++METSA N+++ F +
Sbjct: 516 IVIMLVGNKADLRHLRAVPTEEATAFAERENTYFMETSALESLNVDNAFIXV 671
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,312,793
Number of Sequences: 28460
Number of extensions: 13322
Number of successful extensions: 92
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 92
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 92
length of query: 88
length of database: 4,897,600
effective HSP length: 64
effective length of query: 24
effective length of database: 3,076,160
effective search space: 73827840
effective search space used: 73827840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0033b.5


