

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0032.12
(921 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
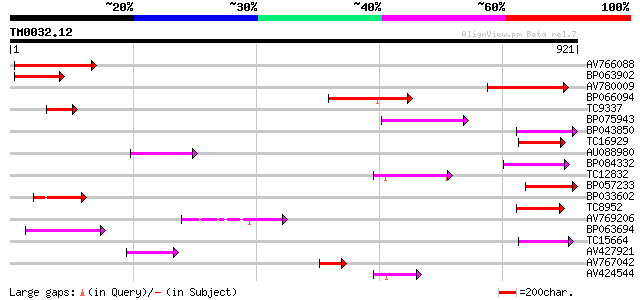
Sequences producing significant alignments: (bits) Value
AV766088 196 2e-50
BP063902 152 2e-37
AV780009 125 2e-29
BP066094 107 1e-23
TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B pre... 92 4e-19
BP075943 86 3e-17
BP043850 83 2e-16
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 82 3e-16
AU088980 82 4e-16
BP084332 79 3e-15
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 78 7e-15
BP057233 73 2e-13
BP033602 72 4e-13
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 71 9e-13
AV769206 69 3e-12
BP063694 69 3e-12
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 65 4e-11
AV427921 65 5e-11
AV767042 60 2e-09
AV424544 58 8e-09
>AV766088
Length = 501
Score = 196 bits (497), Expect = 2e-50
Identities = 90/132 (68%), Positives = 107/132 (80%)
Frame = -3
Query: 9 QTGMPLLRCNSLGDLYPVTCSSHFAGLASGVWHNRLGHPTSSALNHLRNNKLIYCDPSRS 68
QTG+PL+RCNS GDLYPVT S FAG+A +WH+RLGHP+SSAL +LR+NK I +
Sbjct: 397 QTGIPLMRCNSPGDLYPVTLSFPFAGIAQSLWHSRLGHPSSSALRYLRSNKFISYELLNY 218
Query: 69 STVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILSTAGHRYYVLVLDDHTDFL 128
S VC+SCV GKHVRLPF SS +T+ FDILHSDLWTSP+LS+AGHR+YVL LDD TDFL
Sbjct: 217 SPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTDFL 38
Query: 129 WMFPISKKSQVY 140
W FP+S KSQV+
Sbjct: 37 WTFPLSNKSQVF 2
>BP063902
Length = 494
Score = 152 bits (384), Expect = 2e-37
Identities = 71/80 (88%), Positives = 76/80 (94%)
Frame = -3
Query: 9 QTGMPLLRCNSLGDLYPVTCSSHFAGLASGVWHNRLGHPTSSALNHLRNNKLIYCDPSRS 68
+TGMPLLRCNSLGDLYPVT SS FAGLAS VWH+RLGHP SSALNHLRNNKLI+C+PSRS
Sbjct: 273 KTGMPLLRCNSLGDLYPVTRSSPFAGLASSVWHHRLGHPASSALNHLRNNKLIFCEPSRS 94
Query: 69 STVCDSCVLGKHVRLPFSSS 88
S+VCDSCVLGKHVRLPFSSS
Sbjct: 93 SSVCDSCVLGKHVRLPFSSS 34
>AV780009
Length = 529
Score = 125 bits (315), Expect = 2e-29
Identities = 61/133 (45%), Positives = 85/133 (63%)
Frame = -1
Query: 776 GMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLEL 835
G TRRS +GY +FLG +LISW +K+Q T+S SS+EAEYR +A V E W+ L L
Sbjct: 418 GCPDTRRSVTGYSIFLGTSLISWRTKKQTTVSRSSSEAEYRALAATVCEVQWLSYLFQFL 239
Query: 836 HFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQI 895
+ + CDN S+++++ NP H+RTKHIE+D H VR K+ G +L + + HQ+
Sbjct: 238 KLNVPLPVPLFCDNQSALHIAHNPTFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQL 59
Query: 896 ADIFTKGLPRVLF 908
ADIFTK P + F
Sbjct: 58 ADIFTKPCPHLCF 20
>BP066094
Length = 532
Score = 107 bits (266), Expect = 1e-23
Identities = 55/147 (37%), Positives = 90/147 (60%), Gaps = 11/147 (7%)
Frame = +2
Query: 518 KPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKS 577
K IR ++S +++ + +H ++V +AFL+G + E VY+HQP G D ++ D++ + KKS
Sbjct: 89 KTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKKS 268
Query: 578 LYGLKQAPRAWYQRFADYV-----------SSIGFQHSSSDHSYILLYVDDIILVASSHD 626
LYGLKQAPRAWY+R + ++ +++ + D + +YVDDII +++
Sbjct: 269 LYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANPS 448
Query: 627 LRKSFMALLAFEFAMKDLGPLSYFLGI 653
L K F ++ EF M+ +G L YFLGI
Sbjct: 449 LCKEFSEMMQAEFEMRMMGELKYFLGI 529
>TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B precursor
(Pectin methylesterase) (PE) , partial (54%)
Length = 1145
Score = 92.0 bits (227), Expect = 4e-19
Identities = 43/51 (84%), Positives = 46/51 (89%)
Frame = +3
Query: 60 LIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILS 110
L PSRS++VCDSCVLGKHVRLPFSSSETITLR FDILHSDLWTSP+LS
Sbjct: 993 LFKLSPSRSNSVCDSCVLGKHVRLPFSSSETITLRPFDILHSDLWTSPVLS 1145
>BP075943
Length = 547
Score = 85.5 bits (210), Expect = 3e-17
Identities = 48/142 (33%), Positives = 77/142 (53%), Gaps = 1/142 (0%)
Frame = -3
Query: 604 SSSDHSYILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLF 663
SS + +LLYVDD++L + + + L +F +KDLG YFLG+ + R G+
Sbjct: 542 SSGSFTALLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIV 363
Query: 664 LSQSTYASEIIARAGMASCNPSATPVDTK*K-LSSSSGTPCEDVTLYQSLAGALQYLTFT 722
L+Q YA ++I+ +G P T + L +++GTP D+ Y+ + G L YL T
Sbjct: 362 LNQRKYALQLISDSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTT 183
Query: 723 RPNISYVVQQVCLHMHAPHTEH 744
RP+I++ V Q+ + AP H
Sbjct: 182 RPDITFAVNQLSQFLSAPTDIH 117
>BP043850
Length = 515
Score = 83.2 bits (204), Expect = 2e-16
Identities = 46/99 (46%), Positives = 58/99 (58%)
Frame = -1
Query: 823 SESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARG 882
SE W+R LL EL F SQ T +H DN S+I ++ NPV+H+ T+HIE+D H VRE R
Sbjct: 515 SEIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRR 336
Query: 883 QARILHVPSRHQIADIFTKGLPRVLFDDFRSSLSFGEPP 921
+ HV + QIADI TK L R + S L E P
Sbjct: 335 VITLPHVSTSVQIADILTKSLTRQRHNFLVSKLMLVESP 219
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 82.4 bits (202), Expect = 3e-16
Identities = 36/77 (46%), Positives = 52/77 (66%)
Frame = +1
Query: 827 WIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARI 886
W+ LL +L P Q LV+CDN S+ +++ NPV H+RTKHIE+D H VRE++ +G +
Sbjct: 4 WLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIHL 183
Query: 887 LHVPSRHQIADIFTKGL 903
L + S +ADI+TK L
Sbjct: 184 LPISSSEPLADIYTKAL 234
>AU088980
Length = 360
Score = 82.0 bits (201), Expect = 4e-16
Identities = 44/110 (40%), Positives = 59/110 (53%)
Frame = +2
Query: 196 NGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHR 255
N ERK R I N+ R L+ HS +P + W A++ A YL+N LP L+ P Q L +
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 256 DPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDL 305
P+ L+VFG LC+ + KL PR+ CVFLG+ GY DL
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMDL 331
>BP084332
Length = 368
Score = 79.0 bits (193), Expect = 3e-15
Identities = 42/107 (39%), Positives = 63/107 (58%)
Frame = +2
Query: 803 QPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHH 862
Q T++ S+AEAEY A ++ W+++ L + L ++CDN ++I LS NP+ H
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILH 208
Query: 863 QRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFD 909
R KHIE+ HF+R+ V +G + V + HQ ADIFTK L F+
Sbjct: 209 SRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFN 349
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 77.8 bits (190), Expect = 7e-15
Identities = 51/147 (34%), Positives = 73/147 (48%), Gaps = 20/147 (13%)
Frame = +2
Query: 592 FADYVSSIGFQHSSSDHS------------YILLYVDDIILVASSHDLRKSFMALLAFEF 639
F ++ S+G+ SSDH +LLYVDD+++V + D + A LA EF
Sbjct: 2 FDSFIMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREF 181
Query: 640 AMKDLGPLSYFLGIAV--TRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSS 697
MKDLGP + LG+ + R ++LSQ Y +++ R M CNP +TP+ KLSS
Sbjct: 182 DMKDLGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSS 361
Query: 698 SSGTPCEDVTL------YQSLAGALQY 718
S E + Y S G+L Y
Sbjct: 362 SMIPSSEAERMEMSRVPYASAVGSLMY 442
>BP057233
Length = 473
Score = 73.2 bits (178), Expect = 2e-13
Identities = 32/84 (38%), Positives = 52/84 (61%)
Frame = -2
Query: 838 PLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIAD 897
P+ ++ ++ CDN+S+ L+ NPV H R+KHIE+D+H++R++V + + + +VP+ QIAD
Sbjct: 466 PMPRKPILWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQIAD 287
Query: 898 IFTKGLPRVLFDDFRSSLSFGEPP 921
TK L F R L P
Sbjct: 286 CLTKPLSHTRFSQLRDKLGVIHSP 215
>BP033602
Length = 533
Score = 72.0 bits (175), Expect = 4e-13
Identities = 37/88 (42%), Positives = 53/88 (60%), Gaps = 1/88 (1%)
Frame = +3
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
+WHNRLGHP L HL + ++ + + SS C+SCVL K+ R P+ S R F +
Sbjct: 276 LWHNRLGHPNFQYLRHLFPD--LFKNVNCSSLECESCVLAKNQRAPYYSQPYHASRPFYL 449
Query: 99 LHSDLW-TSPILSTAGHRYYVLVLDDHT 125
+HSD+W S I + G R++V +DDHT
Sbjct: 450 IHSDVWGPSKITTQFGKRWFVTFIDDHT 533
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 70.9 bits (172), Expect = 9e-13
Identities = 33/78 (42%), Positives = 50/78 (63%)
Frame = +3
Query: 824 ESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQ 883
E+ W+ L +L +++CDN S+++L+ N V H+RT++IE+D H V KV G
Sbjct: 21 EALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFGI 200
Query: 884 ARILHVPSRHQIADIFTK 901
+LHVPS Q+AD+FTK
Sbjct: 201 LHLLHVPSSDQVADVFTK 254
>AV769206
Length = 536
Score = 69.3 bits (168), Expect = 3e-12
Identities = 63/179 (35%), Positives = 89/179 (49%), Gaps = 8/179 (4%)
Frame = -3
Query: 280 KLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADL-PLESTSSYDC 338
KL S CVF+GY +NH+GYKC D S R + IS+ V+F E RFP+ L P E+TSS
Sbjct: 510 KLSLHSKECVFIGYSINHKGYKCLDQSGR-IYISKDVLFHEHRFPYPSLFPDETTSSTS- 337
Query: 339 FTEDLPPSLIHHWQTTSTRPPDL-SIPPSSPTDSTMPSPAPTSSPTSSAPL-----PLPP 392
E P S I + T PP L S SS +T+PS + +PT+ L PL
Sbjct: 336 -AEFFPLSTIP--IVSHTTPPALASSGFSSLGSATLPS---SFNPTAKLTLTEW*QPLTE 175
Query: 393 VPPTPPTR-TMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQP 450
+ P T T + ++ + + ++SI + ISP+ N P +K+ M P
Sbjct: 174 LTAVPLTELTAVQNRINNCVQLAFQLAHTLSITNIFISPIYSNTNLLYETP-YKYTMAP 1
>BP063694
Length = 511
Score = 68.9 bits (167), Expect = 3e-12
Identities = 37/132 (28%), Positives = 61/132 (46%), Gaps = 2/132 (1%)
Frame = -2
Query: 26 VTCSSHFAGLASGVWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVC-DSCVLGKHVRLP 84
+T SS + +WH+RLGH + L + + +C SC + K RL
Sbjct: 399 LTTSSPLPRASFELWHSRLGHVNFDIIKQLHKHGCLDVSSILPKPICCTSCQMAKSKRLV 220
Query: 85 FSSSETITLRSFDILHSDL-WTSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIF 143
F + D++H DL SP+ S G Y+V+ +DD + F W +P+ +KS D+
Sbjct: 219 FHDNNKRASAVLDLIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVL 40
Query: 144 TTLATLIKTQFS 155
++ +FS
Sbjct: 39 LRFKVFMENRFS 4
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 65.5 bits (158), Expect = 4e-11
Identities = 33/89 (37%), Positives = 51/89 (57%)
Frame = +1
Query: 827 WIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARI 886
W+ +L +L P +L++CD+ S+ +++ N V H+RTKH+++D H VREK+ +
Sbjct: 13 WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAKLFHL 192
Query: 887 LHVPSRHQIADIFTKGLPRVLFDDFRSSL 915
L + S Q ADI TK L F S L
Sbjct: 193 LPISSVDQTADILTKPLESGPFSHLVSKL 279
>AV427921
Length = 284
Score = 65.1 bits (157), Expect = 5e-11
Identities = 33/84 (39%), Positives = 47/84 (55%)
Frame = +1
Query: 191 HTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQ 250
+T QNG SERK RTI NM+R+LL S +P SF A+ + ++LN P ++N P +
Sbjct: 19 YTPQQNGVSERKNRTILNMVRSLLTMSGLPKSFLPEAVMWSLHILNRSPTLVVQNMMPEE 198
Query: 251 RLYHRDPSYSHLRVFGCLCFPFFP 274
R P+ H R+ CL + P
Sbjct: 199 AWSGRQPAVDHFRISRCLAYAHVP 270
>AV767042
Length = 444
Score = 60.1 bits (144), Expect = 2e-09
Identities = 29/44 (65%), Positives = 36/44 (80%)
Frame = -3
Query: 503 QIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFL 546
Q AGVDC++TF+ VVKPA IRTV +IALSRSWPI L+V N ++
Sbjct: 349 QTAGVDCNKTFSPVVKPAPIRTVFTIALSRSWPIPQLNVQNLWI 218
>AV424544
Length = 276
Score = 57.8 bits (138), Expect = 8e-09
Identities = 34/90 (37%), Positives = 48/90 (52%), Gaps = 11/90 (12%)
Frame = +3
Query: 591 RFADYVSSIGFQHSSSDHSY-----------ILLYVDDIILVASSHDLRKSFMALLAFEF 639
+ + Y+ +G+ S+ DHS IL+YVDD+IL + + + L +F
Sbjct: 6 KLSSYLHILGYIQSAHDHSLFTKFRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQF 185
Query: 640 AMKDLGPLSYFLGIAVTRYVGGLFLSQSTY 669
+KDLG L YFLG+ V R GLFLSQ Y
Sbjct: 186 RIKDLGTLKYFLGLEVARSSCGLFLSQRKY 275
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,219,667
Number of Sequences: 28460
Number of extensions: 384776
Number of successful extensions: 10587
Number of sequences better than 10.0: 930
Number of HSP's better than 10.0 without gapping: 5394
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7786
length of query: 921
length of database: 4,897,600
effective HSP length: 99
effective length of query: 822
effective length of database: 2,080,060
effective search space: 1709809320
effective search space used: 1709809320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0032.12


