

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
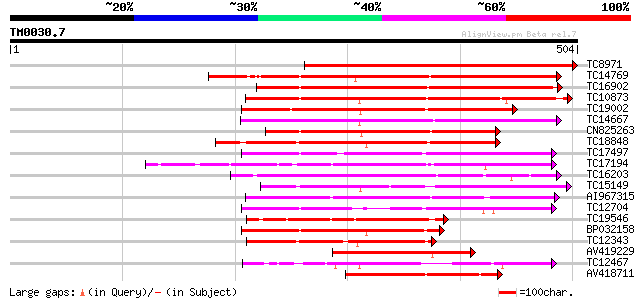
Query= TM0030.7
(504 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 480 e-136
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 236 8e-63
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 219 8e-58
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 208 2e-54
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 197 4e-51
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 194 2e-50
CN825263 183 7e-47
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 183 7e-47
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 177 3e-45
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 172 9e-44
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 172 1e-43
TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like pro... 150 5e-37
AI967315 149 1e-36
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 147 3e-36
TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine k... 145 2e-35
BP032158 141 2e-34
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 140 6e-34
AV419229 139 8e-34
TC12467 UP|CAE02597 (CAE02597) Nod-factor receptor 5, complete 139 1e-33
AV418711 134 3e-32
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 480 bits (1235), Expect = e-136
Identities = 242/242 (100%), Positives = 242/242 (100%)
Frame = +3
Query: 263 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAI 322
EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAI
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAI 185
Query: 323 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVK 382
DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVK
Sbjct: 186 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVK 365
Query: 383 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE 442
GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE
Sbjct: 366 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE 545
Query: 443 GSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADYLKSAR 502
GSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADYLKSAR
Sbjct: 546 GSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADYLKSAR 725
Query: 503 RE 504
RE
Sbjct: 726 RE 731
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 236 bits (601), Expect = 8e-63
Identities = 132/316 (41%), Positives = 198/316 (61%), Gaps = 2/316 (0%)
Frame = +3
Query: 177 KIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKAN 236
K P + S PS+ K + L +V AT + +TL IG+GGFG+VY+
Sbjct: 2052 KYPMETNIIFSLPSKDDFFIKSVSIQAFTLEYIEV--ATERY-KTL-IGEGGFGSVYRGT 2219
Query: 237 LEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVP 296
L DG VAVK + R EF +E+ LL+ I H NLV LLG+ ++ +++IL+ ++
Sbjct: 2220 LNDGQEVAVKVRSSTSTQGTR-EFDNELNLLSAIQHENLVPLLGYCNESDQQILVYPFMS 2396
Query: 297 NGTLREHLDG--LRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTES 354
NG+L++ L G + KILD+ RL IA+ A GL YLH + + +IHRD+KSSNILL S
Sbjct: 2397 NGSLQDRLYGEPAKRKILDWPTRLSIALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHS 2576
Query: 355 MRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEI 414
M AKVADFGF++ P GD +++S +V+GT GYLDPEY KT QL+ KSDV+SFG++LLEI
Sbjct: 2577 MCAKVADFGFSKYAPQEGD-SYVSLEVRGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEI 2753
Query: 415 LTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAA 474
++GR P+ +K+ E + WA V E++DP ++ +A+ + ++++++ QC
Sbjct: 2754 VSGREPLNIKRPRTEWSLVEWATPYIRGSKVDEIVDPGIKGGYHAEAMWRVVEVALQCLE 2933
Query: 475 PIRTDRPNMKSVGEQL 490
P T RP+M ++ +L
Sbjct: 2934 PFSTYRPSMVAIVREL 2981
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 219 bits (558), Expect = 8e-58
Identities = 117/272 (43%), Positives = 169/272 (62%)
Frame = +3
Query: 220 ETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLL 279
E L +G GGFG VY ++ G VA+KR + + EF +E+E+L+K+ HR+LV L+
Sbjct: 3 EALLLGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVH-EFQTEIEMLSKLRHRHLVSLI 179
Query: 280 GFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQI 339
G+ ++ E IL+ +++ GTLREHL + L + QRLEI I A GL YLH A+ I
Sbjct: 180 GYCEENTEMILVYDHMAYGTLREHLYKTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTI 359
Query: 340 IHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLT 399
IHRDVK++NILL E AKV+DFG ++ GP D TH+ST VKG+ GYLDPEY + QLT
Sbjct: 360 IHRDVKTTNILLDEKWVAKVSDFGLSKTGPTL-DNTHVSTVVKGSFGYLDPEYFRRQQLT 536
Query: 400 PKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNA 459
KSDVYSFG++L EIL R + A ++ WA YN+G + +++DP ++ +
Sbjct: 537 DKSDVYSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAP 716
Query: 460 DVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
+ K + + +C + +RP+M G+ LW
Sbjct: 717 ECFKKFAETAMKCVSDQGIERPSM---GDVLW 803
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 208 bits (529), Expect = 2e-54
Identities = 117/295 (39%), Positives = 180/295 (60%), Gaps = 4/295 (1%)
Frame = +1
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAK 269
++ ATRNF E IG+GGFG VYK L G VAVK+ + + + EF EV +L+
Sbjct: 427 ELADATRNFKEANLIGEGGFGKVYKGRLTTGEAVAVKQLSHDGRQGFQ-EFVMEVLMLSL 603
Query: 270 IDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRG--KILDFNQRLEIAIDVAHG 327
+ H NLV+L+G+ G++R+L+ EY+P G+L +HL L + L+++ R+++A+ A G
Sbjct: 604 LHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVGAARG 783
Query: 328 LTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGY 387
L YLH A+ +I+RD+KS+NILL K++DFG A+LGPV GD TH+ST+V GT GY
Sbjct: 784 LEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPV-GDNTHVSTRVMGTYGY 960
Query: 388 LDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY--NEGSV 445
PEY + +LT KSD+YSFG++LLE+LTGRR ++ + E+ + WA R Y +
Sbjct: 961 CAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLVSWA-RPYFSDRRRF 1137
Query: 446 VELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADYLKS 500
++DPL++ + L + + ++ C RP + + + +YL S
Sbjct: 1138GHMVDPLLQGRFPSRCLHQAIAITAMCLQEQPKFRPLITDI-----VVALEYLAS 1287
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 197 bits (500), Expect = 4e-51
Identities = 107/247 (43%), Positives = 154/247 (62%), Gaps = 2/247 (0%)
Frame = +1
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVEL 266
+L ++ AT NF+ ++G+GGFG+VY L DG +AVKR K EF+ EVE+
Sbjct: 31 SLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLK-VWSNKADMEFAVEVEI 207
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGK--ILDFNQRLEIAIDV 324
LA++ H+NL+ L G+ +G ER+++ +Y+PN +L HL G +LD+N+R+ IAI
Sbjct: 208 LARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAIGS 387
Query: 325 AHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGT 384
A G+ YLH A IIHRD+K+SN+LL +A+VADFGFA+L P TH++T+VKGT
Sbjct: 388 AEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIP--DGATHVTTRVKGT 561
Query: 385 VGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGS 444
+GYL PEY + DV+SFGILLLE+ +G++P+E + +R WA
Sbjct: 562 LGYLAPEYAMLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSINDWALPLACAKK 741
Query: 445 VVELMDP 451
E DP
Sbjct: 742 FTEFADP 762
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 194 bits (494), Expect = 2e-50
Identities = 111/292 (38%), Positives = 165/292 (56%), Gaps = 7/292 (2%)
Frame = +2
Query: 206 LNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVE 265
L+L ++ T NF IG+G +G VY A L DG VAVK+ EF ++V
Sbjct: 323 LSLDELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEFLTQVS 502
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRG-------KILDFNQRL 318
+++++ + N V+L G+ +GN R+L E+ G+L + L G +G LD+ QR+
Sbjct: 503 MVSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRV 682
Query: 319 EIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHIS 378
IA+D A GL YLH + IIHRD++SSN+L+ E +AK+ADF + P + H S
Sbjct: 683 RIAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLH-S 859
Query: 379 TKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFR 438
T+V GT GY PEY T QLT KSDVYSFG++LLE+LTGR+PV+ ++ + WA
Sbjct: 860 TRVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATP 1039
Query: 439 KYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
+ +E V + +DP ++ + K+ ++ C RPNM V + L
Sbjct: 1040RLSEDKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKAL 1195
>CN825263
Length = 663
Score = 183 bits (464), Expect = 7e-47
Identities = 105/211 (49%), Positives = 134/211 (62%), Gaps = 2/211 (0%)
Frame = +1
Query: 228 GFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNE 287
GFG VYK L DG VAVK KR+ R EF +EVE+L+++ HRNLVKL+G +
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGR-EFLAEVEMLSRLHHRNLVKLIGICIEKQT 177
Query: 288 RILITEYVPNGTLREHLDGLRGKI--LDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVK 345
R LI E VPNG++ HL G + LD+N R++IA+ A GL YLH + +IHRD K
Sbjct: 178 RCLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFK 357
Query: 346 SSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVY 405
SSNILL KV+DFG AR G++ HIST V GT GYL PEY T L KSDVY
Sbjct: 358 SSNILLECDFTPKVSDFGLARTALDEGNK-HISTHVMGTFGYLAPEYAMTGHLLVKSDVY 534
Query: 406 SFGILLLEILTGRRPVELKKAADERVTLRWA 436
S+G++LLE+LTG +PV+L + + + WA
Sbjct: 535 SYGVVLLELLTGTKPVDLSQPPGQENLVTWA 627
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 183 bits (464), Expect = 7e-47
Identities = 102/255 (40%), Positives = 154/255 (60%), Gaps = 2/255 (0%)
Frame = +2
Query: 184 RVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVV 243
+V P+ F R+ T ++ AT FS+ ++G+GGFG+VY DGL +
Sbjct: 212 KVEEGPTSFGSVNNSWRIFTYK----ELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQI 379
Query: 244 AVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREH 303
AVK+ K + ++ EF+ EVE+L ++ H+NL+ L G+ ++R+++ +Y+PN +L H
Sbjct: 380 AVKKLKAMNSKA-EMEFAVEVEVLGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSH 556
Query: 304 LDGLRGKILDFN--QRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVAD 361
L G + N +R++IAI A G+ YLH IIHRD+K+SN+LL VAD
Sbjct: 557 LHGQFAVEVQLNWQKRMKIAIGSAEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVAD 736
Query: 362 FGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPV 421
FGFA+L P +H++T+VKGT+GYL PEY +++ DVYSFGILLLE++TGR+P+
Sbjct: 737 FGFAKLIPEG--VSHMTTRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLELVTGRKPI 910
Query: 422 ELKKAADERVTLRWA 436
E +R WA
Sbjct: 911 EKLPGGVKRTITEWA 955
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 177 bits (450), Expect = 3e-45
Identities = 105/281 (37%), Positives = 165/281 (58%), Gaps = 1/281 (0%)
Frame = +1
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVEL 266
+ S + AT +FS + ++G+GGFG VYK L +G +AVKR + + EF +E++L
Sbjct: 1639 DFSTISSATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGME-EFKNEIKL 1815
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAH 326
+A++ HRNLVKL G +E N ++ LD R K++D+N+RL+I +A
Sbjct: 1816 IARLQHRNLVKLFGCSVHQDENSHA-----NKKMKILLDSTRSKLVDWNKRLQIIDGIAR 1980
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTK-VKGTV 385
GL YLH + +IIHRD+K+SNILL + M K++DFG AR+ GDQ TK V GT
Sbjct: 1981 GLLYLHQDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARI--FIGDQVEARTKRVMGTY 2154
Query: 386 GYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSV 445
GY+ PEY + KSDV+SFG+++LEI++G++ L A+R + E
Sbjct: 2155 GYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIGRFYDPHHHLNLLSHAWRLWIEERP 2334
Query: 446 VELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
+EL+D L+++ V +++ + ++ C +RP+M S+
Sbjct: 2335 LELVDELLDDPVIPTEILRYIHVALLCVQRRPENRPDMLSI 2457
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 172 bits (437), Expect = 9e-44
Identities = 123/372 (33%), Positives = 200/372 (53%), Gaps = 6/372 (1%)
Frame = +1
Query: 121 VGIFAGGALLACCAVLCPCFYGKKRKATAHAVLAKDPISMDSATSFDASVSASVSGKIPA 180
VGI G + +L C Y + +K ISM +T +AS S +
Sbjct: 814 VGISIAGTFVLL--LLAFCMYVRYQKKEEEKAKLPTDISMALSTQ---DGNASSSAEYET 978
Query: 181 SPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDG 240
S P + S ++ + ++ + ++ +AT NFS +IGQGGFG VY A L G
Sbjct: 979 SGSSGPGTASATGLT-SIMVAKSMEFSYQELAKATNNFSLDNKIGQGGFGAVYYAELR-G 1152
Query: 241 LVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTL 300
A+K+ + TEF E+++L + H NLV+L+G+ +G+ L+ E++ NG L
Sbjct: 1153 KKTAIKKMDVQ----ASTEFLCELKVLTHVHHLNLVRLIGYCVEGS-LFLVYEHIDNGNL 1317
Query: 301 REHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVA 360
++L G + L ++ R++IA+D A GL Y+H + IHRDVKS+NIL+ +++R KVA
Sbjct: 1318 GQYLHGSGKEPLPWSSRVQIALDAARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVA 1497
Query: 361 DFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRP 420
DFG +L V G+ T + T++ GT GY+ PEY + ++PK DVY+FG++L E+++ +
Sbjct: 1498 DFGLTKLIEV-GNST-LQTRLVGTFGYMPPEYAQYGDISPKIDVYAFGVVLFELISAKNA 1671
Query: 421 V----ELKKAADERVTL-RWAFRKYNE-GSVVELMDPLMEEAVNADVLMKMLDLSFQCAA 474
V EL + V L A K + ++ +L+DP + E D ++K+ L C
Sbjct: 1672 VLKTGELVAESKGLVALFEEALNKSDPCDALRKLVDPRLGENYPIDSVLKIAQLGRACTR 1851
Query: 475 PIRTDRPNMKSV 486
RP+M+S+
Sbjct: 1852 DNPLLRPSMRSL 1887
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 172 bits (436), Expect = 1e-43
Identities = 108/301 (35%), Positives = 164/301 (53%), Gaps = 7/301 (2%)
Frame = +2
Query: 197 KLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESL 256
KL+ L + VV + E IG+GG G VY+ ++ +G VA+KR +
Sbjct: 2138 KLTAFQRLEIKAEDVVECLK---EENIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRN 2308
Query: 257 RTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQ 316
F +E+E L KI HRN+++LLG++ + +L+ EY+PNG+L E L G +G L +
Sbjct: 2309 DYGFRAEIETLGKIRHRNIMRLLGYVSNKDTNLLLYEYMPNGSLGEWLHGAKGGHLRWEM 2488
Query: 317 RLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTH 376
R +IA++ A GL Y+H IIHRDVKS+NILL A VADFG A+ G
Sbjct: 2489 RYKIAVEAARGLCYMHHDCSPLIIHRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQS 2668
Query: 377 ISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWA 436
+S+ + G+ GY+ PEY T ++ KSDVYSFG++LLE++ GR+PV + D + W
Sbjct: 2669 MSS-IAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIIGRKPV--GEFGDGVDIVGWV 2839
Query: 437 FRKYNEGS-------VVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQ 489
+ +E S V+ ++DP + V+ M +++ C + RP M+ V
Sbjct: 2840 NKTMSELSQPSDTALVLAVVDPRLSGYPLTSVI-HMFNIAMMCVKEMGPARPTMREVVHM 3016
Query: 490 L 490
L
Sbjct: 3017 L 3019
>TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like protein kinase
protein (AT3g17840/MEB5_6), partial (45%)
Length = 1489
Score = 150 bits (379), Expect = 5e-37
Identities = 101/281 (35%), Positives = 149/281 (52%), Gaps = 5/281 (1%)
Frame = +3
Query: 224 IGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFID 283
+G+G FGT YKA LE GLVVAVKR K EF ++E + +DH++LV L +
Sbjct: 294 LGKGTFGTAYKAVLETGLVVAVKRLKDVTISE--KEFKDKIETVGAMDHQSLVPLRAYYF 467
Query: 284 KGNERILITEYVPNGTLREHLDGLRGK---ILDFNQRLEIAIDVAHGLTYLHLYAEKQII 340
+E++L+ +Y+P G+L L G +G L++ R IA+ A G+ YLH +
Sbjct: 468 SRDEKLLVYDYMPMGSLSALLHGNKGAGRTPLNWEIRSGIALGAARGIEYLHSQGPN-VS 644
Query: 341 HRDVKSSNILLTESMRAKVADFGFARL-GPVNGDQTHISTKVKGTVGYLDPEYMKTHQLT 399
H ++K+SNILLT+S AKV+DFG A L GP S+ GY PE +++
Sbjct: 645 HGNIKASNILLTKSYEAKVSDFGLAHLVGP--------SSTPNRVAGYRAPEVTDPRKVS 800
Query: 400 PKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMD-PLMEEAVN 458
K+DVYSFG+LLLE+LTG+ P + RW E E+ D L+
Sbjct: 801 QKADVYSFGVLLLELLTGKAPTHALLNEEGVDLPRWVQSLVREEWTSEVFDLELLRYQNV 980
Query: 459 ADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRADYLK 499
+ ++++L L+ CAAP RP++ V + + LK
Sbjct: 981 EEEMVQLLQLAVDCAAPYPDRRPSIAEVTRSIEELCRSNLK 1103
>AI967315
Length = 1308
Score = 149 bits (376), Expect = 1e-36
Identities = 94/281 (33%), Positives = 147/281 (51%), Gaps = 2/281 (0%)
Frame = +1
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKRE-HFESLRTEFSSEVELLA 268
++ AT FS +G+GG+ VYK LE G +AVKR R E EF +E+ +
Sbjct: 124 ELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRLTRTCRDERKEKEFLTEIGTIG 303
Query: 269 KIDHRNLVKLLGF-IDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHG 327
+ H N++ LLG ID G L+ E G++ + + LD+ R +I + A G
Sbjct: 304 HVCHSNVMPLLGCCIDNG--LYLVFELSTVGSVASLIHDEKMAPLDWKTRYKIVLGTARG 477
Query: 328 LTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGY 387
L YLH +++IIHRD+K+SNILLTE +++DFG A+ P H ++GT G+
Sbjct: 478 LHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTH-HSIAPIEGTFGH 654
Query: 388 LDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVE 447
L PEY + K+DV++FG+ LLE+++GR+PV+ + WA ++ + +
Sbjct: 655 LAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKPVD----GSHQSLHTWAKPILSKWEIEK 822
Query: 448 LMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGE 488
L+DP +E + ++ + C T RP M V E
Sbjct: 823 LVDPRLEGCYDVTQFNRVAFAASLCIRASSTWRPTMSEVLE 945
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 147 bits (372), Expect = 3e-36
Identities = 105/305 (34%), Positives = 157/305 (51%), Gaps = 25/305 (8%)
Frame = +1
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVEL 266
+ S + T +FSE+ ++G+GGFG VYK L +G +AVKR + + EF +EV+L
Sbjct: 1420 DFSTISSTTNHFSESNKLGEGGFGPVYKGVLANGQEIAVKRLSNTSGQGME-EFKNEVKL 1596
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAH 326
+A++ HRNLVKLLG +E +LI E++ N +L + F+ RL
Sbjct: 1597 IARLQHRNLVKLLGCSIHHDEMLLIYEFMHNRSLDYFI---------FDSRL-------- 1725
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTK-VKGTV 385
+IIHRD+K+SNILL M K++DFG AR+ GDQ TK V GT
Sbjct: 1726 -----------RIIHRDLKTSNILLDSEMNPKISDFGLARI--FTGDQVEAKTKRVMGTY 1866
Query: 386 GYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRR--------------------PVELKK 425
GY+ PEY + KSDV+SFG+++LEI++G++ V L K
Sbjct: 1867 GYMSPEYAVHGSFSVKSDVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHSSNFAVFLIK 2046
Query: 426 AAD----ERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRP 481
A E V R A+R + E +EL+D L++ +++ + ++ C RP
Sbjct: 2047 ALRICMFENVKNRKAWRLWIEERPLELVDELLDGLAIPTEILRYIHIALLCVQQRPEYRP 2226
Query: 482 NMKSV 486
+M SV
Sbjct: 2227 DMLSV 2241
>TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine kinase ,
partial (46%)
Length = 731
Score = 145 bits (365), Expect = 2e-35
Identities = 86/181 (47%), Positives = 123/181 (67%), Gaps = 1/181 (0%)
Frame = +1
Query: 211 VVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKI 270
+ +AT+NF+ L GQG FGTVYKA + G VVAVK + + EF +EV LL ++
Sbjct: 214 IQKATQNFTTIL--GQGSFGTVYKATMPTGEVVAVK-VLAPNSKQGEHEFQTEVHLLGRL 384
Query: 271 DHRNLVKLLGF-IDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLT 329
HRNLV L+GF +DKG +RIL+ +++ NG+L L G K L +++RL+IA+D++HG+
Sbjct: 385 HHRNLVNLVGFCVDKG-QRILVYQFMSNGSLANLLYG-EEKELSWDERLQIAMDISHGIE 558
Query: 330 YLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLD 389
YLH A +IHRD+KS+NILL +SMRA VADFG ++ +G ++ +KGT GY+D
Sbjct: 559 YLHEGAVPPVIHRDLKSANILLDDSMRAMVADFGLSKEEIFDGR----NSGLKGTYGYMD 726
Query: 390 P 390
P
Sbjct: 727 P 729
>BP032158
Length = 555
Score = 141 bits (356), Expect = 2e-34
Identities = 74/182 (40%), Positives = 118/182 (64%), Gaps = 2/182 (1%)
Frame = +2
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVEL 266
+L Q+ AT NF +IG+GGFG VYK L +G V+AVK+ + + R EF +E+ +
Sbjct: 17 SLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNR-EFINEIGM 193
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFN--QRLEIAIDV 324
++ + H NLVKL G +GN+ +L+ EY+ N +L L G + L+ N R++I + +
Sbjct: 194 ISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVGI 373
Query: 325 AHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGT 384
A GL YLH + +I+HRD+K++N+LL + ++AK++DFG A+L + THIST++ GT
Sbjct: 374 AKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLD--EEENTHISTRIAGT 547
Query: 385 VG 386
+G
Sbjct: 548 IG 553
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 140 bits (352), Expect = 6e-34
Identities = 78/174 (44%), Positives = 111/174 (62%), Gaps = 5/174 (2%)
Frame = +3
Query: 211 VVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKI 270
+V AT+NF ++G+GGFG V+K L DG +AVK+ R + RT+F +E +LL ++
Sbjct: 222 LVAATKNFHAVNKLGEGGFGPVFKGKLNDGREIAVKKLSRRSNQG-RTQFINEAKLLTRV 398
Query: 271 DHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGL-----RGKILDFNQRLEIAIDVA 325
HRN+V L G+ G+E++L+ EYVP RE LD L + + LD+ +R +I VA
Sbjct: 399 QHRNVVSLFGYCAHGSEKLLVYEYVP----RESLDKLLFRSQKKEQLDWKRRFDIISGVA 566
Query: 326 HGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHIST 379
GL YLH + IIHRD+K++NILL E K+ADFG AR+ P DQTH++T
Sbjct: 567 RGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLARIFP--EDQTHVNT 722
>AV419229
Length = 395
Score = 139 bits (351), Expect = 8e-34
Identities = 71/131 (54%), Positives = 92/131 (70%), Gaps = 4/131 (3%)
Frame = +1
Query: 288 RILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSS 347
++L+ E++PNGTLR+HL + L F+ RL++A+ A GL YLH A+ I HRDVK++
Sbjct: 1 QMLVYEFMPNGTLRDHLSASSKEPLSFSTRLKVALGSAKGLAYLHTEADPPIFHRDVKAT 180
Query: 348 NILLTESMRAKVADFGFARLGPVNGDQ----THISTKVKGTVGYLDPEYMKTHQLTPKSD 403
NILL AKVADFG +RL PV + H+ST VKGT GYLDPEY TH+LT KSD
Sbjct: 181 NILLDSRFSAKVADFGLSRLAPVPDLEGIVPGHVSTVVKGTPGYLDPEYFLTHKLTDKSD 360
Query: 404 VYSFGILLLEI 414
VYS G++LLE+
Sbjct: 361 VYSLGVVLLEL 393
>TC12467 UP|CAE02597 (CAE02597) Nod-factor receptor 5, complete
Length = 2292
Score = 139 bits (350), Expect = 1e-33
Identities = 94/291 (32%), Positives = 153/291 (52%), Gaps = 12/291 (4%)
Frame = +3
Query: 208 LSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELL 267
+ +++ AT++FS+ ++G+ +VYKAN+E G VVAVK+ K + E+++L
Sbjct: 1056 IDEIMEATKDFSDECKVGE----SVYKANIE-GRVVAVKKIKEGGA-------NEELKIL 1199
Query: 268 AKIDHRNLVKLLGFIDKGNER--ILITEYVPNGTLREHLDGLRG---KILDFNQRLEIAI 322
K++H NLVKL+G + G + L+ EY NG+L E L L ++QR+ IA+
Sbjct: 1200 QKVNHGNLVKLMG-VSSGYDGNCFLVYEYAENGSLAEWLFSKSSGTPNSLTWSQRISIAV 1376
Query: 323 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVK 382
DVA GL Y+H + +IIHRD+ +SNILL + +AK+A+F AR
Sbjct: 1377 DVAVGLQYMHEHTYPRIIHRDITTSNILLDSNFKAKIANFAMAR---------------- 1508
Query: 383 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWA------ 436
T+ + PK DV++FG+LL+E+LTGR+ + K+ + V + W
Sbjct: 1509 ----------TSTNPMMPKIDVFAFGVLLIELLTGRKAMTTKENGE--VVMLWKDMWEIF 1652
Query: 437 -FRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
+ E + + MDP +E + D + + L+ C A RP+M +
Sbjct: 1653 DIEENREERIRKWMDPNLESFYHIDNALSLASLAVNCTADKSLSRPSMAEI 1805
>AV418711
Length = 419
Score = 134 bits (338), Expect = 3e-32
Identities = 75/141 (53%), Positives = 93/141 (65%), Gaps = 1/141 (0%)
Frame = +3
Query: 299 TLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAK 358
TL+ HL G L + +RL+I I A GL YLH K +IHRDVKS+NILL +++ AK
Sbjct: 3 TLKSHLYGSGFPSLSWKERLDICIGSARGLHYLHTGYAKAVIHRDVKSANILLDDNLMAK 182
Query: 359 VADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGR 418
VADFG ++ GP DQTH+ST VKG+ GYLDPEY + QLT KSDVYSFG++L E+L
Sbjct: 183 VADFGLSKTGP-ELDQTHVSTAVKGSFGYLDPEYFRRQQLTEKSDVYSFGVVLFEVLCA- 356
Query: 419 RPVELKKAADERVTL-RWAFR 438
RPV E V L WA +
Sbjct: 357 RPVIDPSLPREMVNLAEWAMK 419
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,463,512
Number of Sequences: 28460
Number of extensions: 91258
Number of successful extensions: 985
Number of sequences better than 10.0: 288
Number of HSP's better than 10.0 without gapping: 805
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 809
length of query: 504
length of database: 4,897,600
effective HSP length: 94
effective length of query: 410
effective length of database: 2,222,360
effective search space: 911167600
effective search space used: 911167600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0030.7


