

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0029b.1
(1377 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
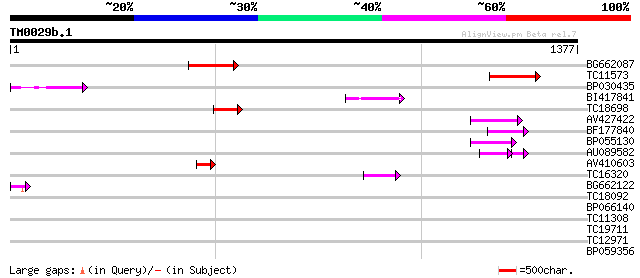
Score E
Sequences producing significant alignments: (bits) Value
BG662087 214 1e-55
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 172 3e-43
BP030435 157 8e-39
BI417841 77 2e-14
TC18698 75 7e-14
AV427422 70 2e-12
BF177840 66 3e-11
BP055130 62 8e-10
AU089582 52 2e-08
AV410603 55 1e-07
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 51 1e-06
BG662122 47 2e-05
TC18092 35 0.081
BP066140 31 1.5
TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoa... 31 1.5
TC19711 31 1.5
TC12971 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14... 30 3.4
BP059356 29 5.8
>BG662087
Length = 373
Score = 214 bits (544), Expect = 1e-55
Identities = 101/120 (84%), Positives = 109/120 (90%)
Frame = +1
Query: 435 NGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADED 494
+GKWRM VDYTDLNKACPKDSYPLPSID LVDGAS NELLSLMDAYSGYHQI+MHP+DED
Sbjct: 13 SGKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDED 192
Query: 495 KTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDH 554
KTAFMTARVNYCY+T+PFGLKNAGATYQ LMDRVF VGRNMEVY+D+MIVKS +H
Sbjct: 193 KTAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 172 bits (436), Expect = 3e-43
Identities = 80/124 (64%), Positives = 100/124 (80%)
Frame = +2
Query: 1165 DNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLD 1224
+NG QF+S QT++FC MGIQMRF+SV+HPQTNGQ E+AN+VIL+G++RRL EA+G W+D
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 1225 ELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELD 1284
ELP VLWSYNT QS+ +ETPF +TYG D MLPVEI NF+W T+ EEEN+ NMAV +
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEISNFSW*TKARHEEENRMNMAVGWE 367
Query: 1285 LLSE 1288
L E
Sbjct: 368 KLPE 379
>BP030435
Length = 533
Score = 157 bits (398), Expect = 8e-39
Identities = 90/186 (48%), Positives = 108/186 (57%)
Frame = +1
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C +H++ GH T++CRTLQRE+ KLI G K +
Sbjct: 103 CVYHQAMGHITEECRTLQREVGKLIATG----------------------------KPTR 198
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHVRAVNSVHEVAFG 121
Q A KG ++TIAGGFGGGG T AARKR+ RAVN+V EV FG
Sbjct: 199 IQGGNAPVPKG---------------VHTIAGGFGGGGITSAARKRYARAVNTVTEVLFG 333
Query: 122 FVHPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLT 181
F HP IT S DF IKPH D+PIV+ LR+N N++RVLLDQGSSADIIYG FD+LGL
Sbjct: 334 FSHPVITFSSDDFIRIKPHLDNPIVILLRVNQLNVQRVLLDQGSSADIIYGGVFDRLGLN 513
Query: 182 DNDLTP 187
+ DLTP
Sbjct: 514 EADLTP 531
>BI417841
Length = 617
Score = 77.0 bits (188), Expect = 2e-14
Identities = 50/146 (34%), Positives = 77/146 (52%), Gaps = 4/146 (2%)
Frame = +1
Query: 816 TLFVDGSSN----SSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLARE 871
TL DGSS S+G+GA + E ++ L + + + TNNQAEY LI GLK A E
Sbjct: 139 TLEFDGSSKGNPGSAGAGAVLRAEDGSKVYLREGVGNQ---TNNQAEYRGLILGLKHAHE 309
Query: 872 VKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRA 931
+ + ++ DSQLV QV+G+++ ++PN+ + L + FQ + +VPR N A
Sbjct: 310 QGYQHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEA 489
Query: 932 DALAKLASTRKPGNNKSVIQETLAYP 957
D A L G+ ++E +P
Sbjct: 490 DVQANLGVNLPAGH----VEEYCEFP 555
>TC18698
Length = 808
Score = 75.1 bits (183), Expect = 7e-14
Identities = 37/71 (52%), Positives = 47/71 (66%)
Frame = -2
Query: 495 KTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDH 554
KT RVNY Y+ MP GLKN TYQRLMD++F +Q+ +N+EVYV+DMIVKS + H
Sbjct: 801 KTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*FH 622
Query: 555 HQDLEEAFGEI 565
DL +I
Sbjct: 621 RGDLSRDLSDI 589
>AV427422
Length = 417
Score = 70.1 bits (170), Expect = 2e-12
Identities = 39/127 (30%), Positives = 70/127 (54%), Gaps = 1/127 (0%)
Frame = +1
Query: 1120 ILVAVDYFTKWIEAEPLAKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREF 1178
+LV VD +K+ PL +AK++ + + + +V G+P +IVSD F S+ +E
Sbjct: 37 VLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMSNFWKEL 216
Query: 1179 CKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQ 1238
K G +++ ++ HP+++GQ E NR + LR +A+ +W +P + YNT+
Sbjct: 217 FKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYWYNTSYH 396
Query: 1239 STTRETP 1245
+T +TP
Sbjct: 397 VSTGQTP 417
>BF177840
Length = 410
Score = 66.2 bits (160), Expect = 3e-11
Identities = 33/100 (33%), Positives = 56/100 (56%)
Frame = +2
Query: 1161 AIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKG 1220
+IVSD T+F S R ++G ++ +++ HPQT+GQ E N+ + LR L
Sbjct: 14 SIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLK 193
Query: 1221 AWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEI 1260
W LP + ++YN STT+ +PF + YG + + P+++
Sbjct: 194 MWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>BP055130
Length = 567
Score = 61.6 bits (148), Expect = 8e-10
Identities = 39/111 (35%), Positives = 59/111 (53%), Gaps = 1/111 (0%)
Frame = +2
Query: 1120 ILVAVDYFTKWIEAEP-LAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREF 1178
I+V VD +K+ P A TS+++ + IV G+PRAIVSD F+S+ + F
Sbjct: 191 IIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAFWKHF 370
Query: 1179 CKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAV 1229
K G + +S HPQT+GQ E+ N+ + LR + E W+ L A+
Sbjct: 371 FKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLWVSYLLAL 523
>AU089582
Length = 383
Score = 52.0 bits (123), Expect(2) = 2e-08
Identities = 25/80 (31%), Positives = 45/80 (56%)
Frame = +2
Query: 1141 SAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQV 1200
+++ Y IV G+P +I+SD G QF+S R F +G +++ ++ HPQT+GQ
Sbjct: 11 ASQYAKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQS 190
Query: 1201 ESANRVILRGLRRRLAEAKG 1220
E +++ LR + + +G
Sbjct: 191 ERTIQILEDMLRACVXDLRG 250
Score = 24.6 bits (52), Expect(2) = 2e-08
Identities = 11/40 (27%), Positives = 20/40 (49%)
Frame = +3
Query: 1219 KGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPV 1258
+G+W L + ++YN + +S+ PF YG P+
Sbjct: 246 EGSWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSPI 365
>AV410603
Length = 162
Score = 54.7 bits (130), Expect = 1e-07
Identities = 23/48 (47%), Positives = 32/48 (65%)
Frame = +1
Query: 453 KDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMT 500
KDS+P+P++D L+D G++ S +D SGYHQI + P D KT F T
Sbjct: 10 KDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRT 153
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 50.8 bits (120), Expect = 1e-06
Identities = 27/91 (29%), Positives = 45/91 (48%)
Frame = +2
Query: 859 YEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEV 918
Y LI GLK A + + + ++ DS LV NQV+G +++K+ N+ + L F
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 919 VVEYVPRAENQRADALAKLASTRKPGNNKSV 949
+ ++PR N AD A + + G + V
Sbjct: 182 KINHIPREYNSEADVQANFGISLRAGQVEEV 274
>BG662122
Length = 386
Score = 47.4 bits (111), Expect = 2e-05
Identities = 25/58 (43%), Positives = 34/58 (58%), Gaps = 10/58 (17%)
Frame = +2
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAG----------YQGNRQGQWRNGGDQNKAHK 49
CE+ RS HD DDC TL+REI+KLI+ G QG+RQ R GD+ + ++
Sbjct: 212 CEYRRSVVHDIDDCFTLKREIEKLIKMGRLKQYDRGSRQQGDRQQGGRLQGDRQQGNR 385
>TC18092
Length = 519
Score = 35.0 bits (79), Expect = 0.081
Identities = 19/35 (54%), Positives = 24/35 (68%)
Frame = -2
Query: 440 MCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELL 474
MCV TD+NK CP+ PL SI++LVD +S E L
Sbjct: 101 MCVC*TDINKDCPR--XPLTSINALVDYSSSYEYL 3
>BP066140
Length = 320
Score = 30.8 bits (68), Expect = 1.5
Identities = 26/73 (35%), Positives = 34/73 (45%)
Frame = -2
Query: 865 GLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVP 924
GL LA E IR LL +D + V + G + YL R L+ E ++
Sbjct: 316 GLSLAWERGIRKLLCISDCEDVVKALTGLGSFHLDSWAAYLAEARSLIHPEWECLLLPHH 137
Query: 925 RAENQRADALAKL 937
R N+ ADALAKL
Sbjct: 136 RERNKPADALAKL 98
>TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (4%)
Length = 756
Score = 30.8 bits (68), Expect = 1.5
Identities = 14/42 (33%), Positives = 27/42 (63%)
Frame = -2
Query: 1131 IEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSS 1172
++ L ++++AK V F R+V RF +PR ++ D+G + +S
Sbjct: 509 VQGTSLLQVSTAKTVEFSDDRVV-RFPLPRDVIGDDGAETTS 387
>TC19711
Length = 579
Score = 30.8 bits (68), Expect = 1.5
Identities = 17/41 (41%), Positives = 23/41 (55%), Gaps = 3/41 (7%)
Frame = -2
Query: 261 LSSGKVGRLKVDQKMARECYNNCLNLYGKKSALV---GHRC 298
L SGK RL++DQK +E NN +N+ K + HRC
Sbjct: 485 LHSGKTKRLEIDQKKRQEKQNNIINILR*KFKIYIKKKHRC 363
>TC12971 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;}, partial (7%)
Length = 721
Score = 29.6 bits (65), Expect = 3.4
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = +3
Query: 41 GGDQNKAHKREEERADTKGKKKQE 64
GGD+ ++ REEE D K K K+E
Sbjct: 261 GGDEIPSNNREEEEGDNKSKNKKE 332
>BP059356
Length = 436
Score = 28.9 bits (63), Expect = 5.8
Identities = 28/92 (30%), Positives = 41/92 (44%), Gaps = 9/92 (9%)
Frame = +1
Query: 43 DQNKAHKREEERADTKGKKKQESAAIAT----KGADDTFAQHS----GPPVGTINTIAGG 94
D+N +K+E A+ KK E+ + + + D F + PP G AGG
Sbjct: 184 DKNPNNKKE---AEANFKKISEAYDVLSDPQKRAVYDQFGEEGLNGQVPPPG-----AGG 339
Query: 95 FGGGGDTHAARKR-HVRAVNSVHEVAFGFVHP 125
F GGGD A R + R+ + + FGF P
Sbjct: 340 FSGGGDGGPASFRFNPRSADDILSEFFGFSSP 435
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,957,489
Number of Sequences: 28460
Number of extensions: 283044
Number of successful extensions: 1259
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1254
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1259
length of query: 1377
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1276
effective length of database: 2,023,140
effective search space: 2581526640
effective search space used: 2581526640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0029b.1


