

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0029a.5
(1393 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
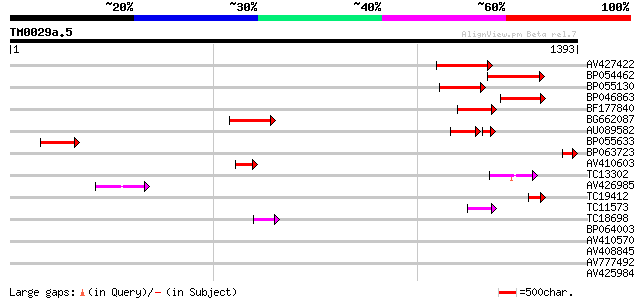
Sequences producing significant alignments: (bits) Value
AV427422 204 1e-52
BP054462 171 6e-43
BP055130 131 6e-31
BP046863 130 1e-30
BF177840 100 2e-21
BG662087 95 7e-20
AU089582 74 2e-19
BP055633 82 6e-16
BP063723 76 3e-14
AV410603 74 2e-13
TC13302 70 2e-12
AV426985 65 1e-10
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 51 1e-06
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 49 7e-06
TC18698 45 1e-04
BP064003 38 0.013
AV410570 37 0.021
AV408845 35 0.11
AV777492 34 0.18
AV425984 31 1.5
>AV427422
Length = 417
Score = 204 bits (518), Expect = 1e-52
Identities = 86/137 (62%), Positives = 110/137 (79%)
Frame = +1
Query: 1050 GLPKAMGKDTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDR 1109
GLPK+ G + +LVVVDR +K++HF+ L HPY AK IA++F++EVVRLHG P SIVSDRD
Sbjct: 7 GLPKSKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDP 186
Query: 1110 VFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSW 1169
+F+S FW E+FK+ GTKLK S+AYHP++DGQTEVVNRC+ETYLRC +PK W W+ W
Sbjct: 187 LFMSNFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPW 366
Query: 1170 AEFWYNTNYHSAIKTTP 1186
AE+WYNT+YH + TP
Sbjct: 367 AEYWYNTSYHVSTGQTP 417
>BP054462
Length = 422
Score = 171 bits (434), Expect = 6e-43
Identities = 79/140 (56%), Positives = 108/140 (76%)
Frame = +1
Query: 1175 NTNYHSAIKTTPFKALYGREPPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQN 1234
N+NY+ + K +PF+ALYGREPPV+ +G + + V L R+ +L +L++NL K+Q+
Sbjct: 1 NSNYNRSAKMSPFQALYGREPPVLLQGTTIPSKIAAVNDLQVGRDELLSDLRANLLKSQD 180
Query: 1235 RMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAY 1294
MR ANK RRDV Y++GD V+LK+QPY+ +SLAK+ N+KLSPRYYGPYPI+AKI AY
Sbjct: 181 MMRTYANKKRRDVDYQIGDEVFLKLQPYRRRSLAKKMNEKLSPRYYGPYPIVAKIGAVAY 360
Query: 1295 KLQLPEGSQVHPVFHISLLK 1314
+L+LP S+VHPVFH+SLLK
Sbjct: 361 RLELPAHSRVHPVFHVSLLK 420
>BP055130
Length = 567
Score = 131 bits (330), Expect = 6e-31
Identities = 58/112 (51%), Positives = 74/112 (65%)
Frame = +2
Query: 1056 GKDTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTF 1115
G I+VVVDR +KY HF Y + ++AE F+ +V+LHG P +IVSDRD+ F S F
Sbjct: 179 GFTVIIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAF 358
Query: 1116 WSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWL 1167
W FKL GT L SS+YHPQTDGQTE +N+C+E YLRC P+ W +L
Sbjct: 359 WKHFFKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLWVSYL 514
>BP046863
Length = 580
Score = 130 bits (328), Expect = 1e-30
Identities = 63/109 (57%), Positives = 85/109 (77%)
Frame = +1
Query: 1207 SVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKS 1266
SV+ V L ER+ +L EL+ NL KAQ++MR QANKHRR V Y+VG+ V+LK+QPYKL++
Sbjct: 253 SVEAVNMLIEERDALLLELRGNLLKAQDQMRAQANKHRRYVDYQVGNWVFLKLQPYKLQN 432
Query: 1267 LAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQVHPVFHISLLKK 1315
LA+R NQKLSPR+YGP+ ++ ++ AY L L S+VHPVFH+SLL+K
Sbjct: 433 LAQRKNQKLSPRFYGPFKVLERVVQVAY*LDLXSESRVHPVFHLSLLEK 579
>BF177840
Length = 410
Score = 100 bits (248), Expect = 2e-21
Identities = 49/95 (51%), Positives = 59/95 (61%)
Frame = +2
Query: 1101 TSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKP 1160
TSIVSDRD F+S FW ++ GTKL +S+ HPQTDGQTEVVN+ + T LR V
Sbjct: 11 TSIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNL 190
Query: 1161 KQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGREP 1195
K W WL EF YN HS K +PF+ +YG P
Sbjct: 191 KMWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNP 295
>BG662087
Length = 373
Score = 95.1 bits (235), Expect = 7e-20
Identities = 48/113 (42%), Positives = 69/113 (60%)
Frame = +1
Query: 540 GGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPK 599
G WR VDY LNKA D +P+P ID+L+D + S +D SGYHQI+M D K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 600 TAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYS 652
TAF T +Y Y +PFGL NA +T+Q LM++V + + + V+ D++++ S
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKS 354
>AU089582
Length = 383
Score = 74.3 bits (181), Expect(2) = 2e-19
Identities = 36/75 (48%), Positives = 49/75 (65%), Gaps = 1/75 (1%)
Frame = +2
Query: 1082 AKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQT 1141
A + A++++ E+V LHG P SI+SDR F S FW GT+LK S+A+HPQTDGQ+
Sbjct: 11 ASQYAKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQS 190
Query: 1142 EVVNRCVETYLR-CV 1155
E + +E LR CV
Sbjct: 191 ERTIQILEDMLRACV 235
Score = 39.7 bits (91), Expect(2) = 2e-19
Identities = 17/31 (54%), Positives = 22/31 (70%)
Frame = +3
Query: 1163 WPKWLSWAEFWYNTNYHSAIKTTPFKALYGR 1193
W ++LS EF YN +Y S+I PF+ALYGR
Sbjct: 255 WDQYLSLMEFAYNNSYRSSI*MAPFEALYGR 347
>BP055633
Length = 528
Score = 82.0 bits (201), Expect = 6e-16
Identities = 40/96 (41%), Positives = 61/96 (62%)
Frame = -2
Query: 75 WEPFKQALFRRFQPALLQNPFGPLLSVKQKGSVMEYRENFELLAAPMRNADREVLKGVFL 134
W FK+A+ +FQ +PF LL++KQ+GSV E+ FE A ++ D E +F+
Sbjct: 518 WTKFKEAMLEQFQLTSNSSPFAALLALKQEGSVEEFVGQFERFAGMLKGIDEEHYMDIFV 339
Query: 135 NGLQEEIKAEMKLYPADDLAELMDRALLLEEKNTAM 170
NGL+EEI AE+KLY L ++ +AL++E+KN A+
Sbjct: 338 NGLKEEIAAEIKLYEPKSLTIMVKKALMVEQKNLAV 231
>BP063723
Length = 479
Score = 76.3 bits (186), Expect = 3e-14
Identities = 33/36 (91%), Positives = 34/36 (93%)
Frame = -3
Query: 1358 IRWKDLPTFEDSWEDFSKLLDQFPNHQLEDKLNLQG 1393
IRWKDLPTFEDSWEDF KLLD FPNHQLED+LNLQG
Sbjct: 477 IRWKDLPTFEDSWEDFCKLLDPFPNHQLEDQLNLQG 370
>AV410603
Length = 162
Score = 73.6 bits (179), Expect = 2e-13
Identities = 33/54 (61%), Positives = 41/54 (75%)
Frame = +1
Query: 554 ATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEG 607
+T+ D FP+P +DELLDE+ + FSKLDL+SGYHQI +K ED KT FRTH G
Sbjct: 1 STVKDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>TC13302
Length = 533
Score = 70.5 bits (171), Expect = 2e-12
Identities = 54/148 (36%), Positives = 67/148 (44%), Gaps = 31/148 (20%)
Frame = -2
Query: 1179 HSAIKTTPFKALYGREPPVIFKGNDSLTSVDEVEKLT--AERNLILEELKSNLE------ 1230
H AIKT PFKALY R ++ KG SL SVD+V L E+ I N
Sbjct: 520 HLAIKTNPFKALYERNTLMVIKGGVSLASVDDVITLNRVVEKPCIDYRYGENSPYKG*SK 341
Query: 1231 -----------------------KAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSL 1267
KA N+MRQQ +HRRD+Q VGD+V P
Sbjct: 340 PHPLS*LLR*S*A*PKF**EQ*AKATNQMRQQV-RHRRDIQLHVGDMV*SN-NPTL*TRA 167
Query: 1268 AKRSNQKLSPRYYGPYPIIAKINPAAYK 1295
+R N KL PR+YG + ++ KINP K
Sbjct: 166 YQRVNHKLCPRFYGTHEMVEKINPVV*K 83
>AV426985
Length = 422
Score = 64.7 bits (156), Expect = 1e-10
Identities = 37/135 (27%), Positives = 67/135 (49%), Gaps = 3/135 (2%)
Frame = +1
Query: 212 LTQTELQERSRKGLCFKCGDKWGKEHICSMKNYQLILMEVEEDEEE---EEIFEEAEDGE 268
++ E+Q R K LC+ C +K+ H C K ++ + E D ++ +++ D
Sbjct: 16 ISPAEMQLRREKNLCYWCDEKFSFNHKCPNKYLMMLQLTDENDSDDTSNQQVTTTISDDS 195
Query: 269 FVLEGKVLQLSLNSKEGLTSNRSFKVKGKIGNREVLILIDCGATSNFISQDLVVELEIPV 328
V + SLN+ G T + + G+IGN V +L+D G++ F+ + L++P+
Sbjct: 196 PVEDH---HFSLNAMRGFTGVGTIRFTGQIGNISVQVLVDGGSSDCFLQPHIAKFLQLPI 366
Query: 329 IATSEYVVEVGNGAK 343
+ + V VGNG K
Sbjct: 367 ESKPNFKVLVGNGQK 411
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 50.8 bits (120), Expect = 1e-06
Identities = 21/43 (48%), Positives = 33/43 (75%), Gaps = 1/43 (2%)
Frame = +3
Query: 1274 KLSPRYYGPYPIIAKINPAAYKLQL-PEGSQVHPVFHISLLKK 1315
KLSPR+ GP+ ++ ++ +Y+L L P+ S VHPVFH+S+L+K
Sbjct: 48 KLSPRFIGPFEVLERVGSVSYRLALPPDLSAVHPVFHVSMLRK 176
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 48.5 bits (114), Expect = 7e-06
Identities = 26/71 (36%), Positives = 38/71 (52%)
Frame = +2
Query: 1124 GTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIK 1183
G +++FSS HPQT+GQTE N+ + ++ +W L + YNT S+IK
Sbjct: 62 GIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWIDELPIVLWSYNTMPQSSIK 241
Query: 1184 TTPFKALYGRE 1194
TPF YG +
Sbjct: 242 ETPF*LTYGAD 274
>TC18698
Length = 808
Score = 44.7 bits (104), Expect = 1e-04
Identities = 21/64 (32%), Positives = 37/64 (57%)
Frame = -2
Query: 599 KTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKNEELH 658
KT + + +Y Y V+P GL N +T+Q LM+++ + K V V+ +D+++ S E H
Sbjct: 801 KTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*FH 622
Query: 659 KDHL 662
+ L
Sbjct: 621 RGDL 610
>BP064003
Length = 509
Score = 37.7 bits (86), Expect = 0.013
Identities = 31/98 (31%), Positives = 51/98 (51%), Gaps = 7/98 (7%)
Frame = +1
Query: 845 LGSKFVIHTDQ-RSLRFLADQRIMGEEQQKWMSKLMGYDFEIKYKPGIE------NKAAD 897
LGS+FVI TD + FL +R + +W L G +++ P ++ NK AD
Sbjct: 112 LGSRFVIKTDNVATSSFLTQKR--APTRARWQDFLSG---GVRHGPRVQAREDQSNKVAD 276
Query: 898 ALSRKLQFSAISSVQCAEWADLEAEILEDERYRKVLQE 935
ALSRK + ++ + A+ + L+ I ER R+ L++
Sbjct: 277 ALSRKAELASNKLEEIADMSQLKGAI--PERIREGLEK 384
>AV410570
Length = 412
Score = 37.0 bits (84), Expect = 0.021
Identities = 18/46 (39%), Positives = 24/46 (52%)
Frame = +2
Query: 201 QPEKKWNGGQRLTQTELQERSRKGLCFKCGDKWGKEHICSMKNYQL 246
QP+ K R++ E+Q R KGLCF C +K+ H C K L
Sbjct: 71 QPQAK-----RISPAEMQLRREKGLCFTCDEKYSWNHKCPNKQLML 193
>AV408845
Length = 429
Score = 34.7 bits (78), Expect = 0.11
Identities = 17/32 (53%), Positives = 22/32 (68%)
Frame = -3
Query: 239 CSMKNYQLILMEVEEDEEEEEIFEEAEDGEFV 270
C +KN + + EE+EEEEE EE E+GEFV
Sbjct: 178 CGVKNEKRVRG*KEEEEEEEEEEEEEEEGEFV 83
>AV777492
Length = 472
Score = 33.9 bits (76), Expect = 0.18
Identities = 11/19 (57%), Positives = 17/19 (88%)
Frame = +1
Query: 1144 VNRCVETYLRCVTGSKPKQ 1162
+NRC+E Y RC+TG++PK+
Sbjct: 403 LNRCLEVYFRCLTGARPKK 459
>AV425984
Length = 351
Score = 30.8 bits (68), Expect = 1.5
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = +1
Query: 1356 VLIRWKDLPTFEDSWEDFS 1374
VLI+WK LP E SWE F+
Sbjct: 16 VLIQWKHLPISESSWESFT 72
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,403,484
Number of Sequences: 28460
Number of extensions: 292666
Number of successful extensions: 1652
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 1592
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1642
length of query: 1393
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1291
effective length of database: 1,994,680
effective search space: 2575131880
effective search space used: 2575131880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0029a.5


