

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
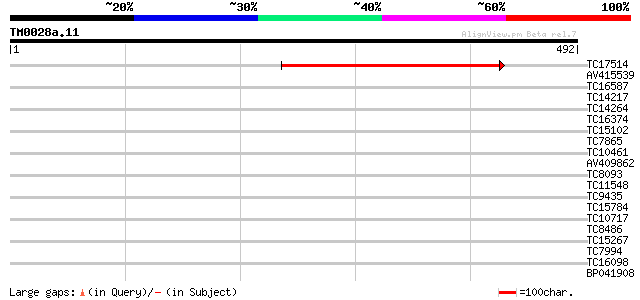
Query= TM0028a.11
(492 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17514 weakly similar to UP|O48683 (O48683) F3I6.9 protein, par... 375 e-105
AV415539 39 0.001
TC16587 weakly similar to UP|Q9JGS5 (Q9JGS5) PORF1, partial (3%) 38 0.003
TC14217 36 0.012
TC14264 similar to UP|Q84NF9 (Q84NF9) Histone H1 (Fragment), par... 35 0.021
TC16374 similar to UP|Q7LZA6 (Q7LZA6) Protamine II (Fragment), p... 35 0.028
TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, ... 35 0.036
TC7865 homologue to UP|SAHH_ARATH (O23255) Adenosylhomocysteinas... 34 0.047
TC10461 homologue to UP|BAD11346 (BAD11346) BRI1-KD interacting ... 34 0.062
AV409862 34 0.062
TC8093 homologue to UP|EFTU_TOBAC (P41342) Elongation factor Tu,... 34 0.062
TC11548 similar to UP|Q9FG37 (Q9FG37) Gamma-tubulin interacting ... 34 0.062
TC9435 similar to UP|Q9AXJ7 (Q9AXJ7) Phosphatase-like protein Mt... 34 0.062
TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precurso... 33 0.081
TC10717 similar to UP|Q8S564 (Q8S564) 4-coumarate:coenzyme A lig... 33 0.081
TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Ara... 33 0.081
TC15267 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Ce... 33 0.081
TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomeras... 33 0.081
TC16098 homologue to UP|Q8LAQ5 (Q8LAQ5) Flower pigmentation prot... 33 0.11
BP041908 33 0.14
>TC17514 weakly similar to UP|O48683 (O48683) F3I6.9 protein, partial (9%)
Length = 582
Score = 375 bits (964), Expect = e-105
Identities = 193/193 (100%), Positives = 193/193 (100%)
Frame = +2
Query: 237 ESNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSE 296
ESNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSE
Sbjct: 2 ESNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSE 181
Query: 297 ESKKVANKPLHKSLNLEPSNPDPAPLPTLRKSFIMESMGDKDIVKRAFKTFKTFQNNYSQ 356
ESKKVANKPLHKSLNLEPSNPDPAPLPTLRKSFIMESMGDKDIVKRAFKTFKTFQNNYSQ
Sbjct: 182 ESKKVANKPLHKSLNLEPSNPDPAPLPTLRKSFIMESMGDKDIVKRAFKTFKTFQNNYSQ 361
Query: 357 PKSSGEDRSSVKKQVPSRGTVSKVPTSSSLRKENGRPNKVESKDTRGGNTVQTTLSRKSD 416
PKSSGEDRSSVKKQVPSRGTVSKVPTSSSLRKENGRPNKVESKDTRGGNTVQTTLSRKSD
Sbjct: 362 PKSSGEDRSSVKKQVPSRGTVSKVPTSSSLRKENGRPNKVESKDTRGGNTVQTTLSRKSD 541
Query: 417 IISGKGKESSRKL 429
IISGKGKESSRKL
Sbjct: 542 IISGKGKESSRKL 580
>AV415539
Length = 419
Score = 39.3 bits (90), Expect = 0.001
Identities = 29/80 (36%), Positives = 35/80 (43%), Gaps = 12/80 (15%)
Frame = +2
Query: 252 SKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKK----------- 300
S +S S R+S PA+ PS TS ST SP S P+ S E S
Sbjct: 71 SPSSSTSPSRSSSPAAAPSHSATSRQSTA---SPCPSSPPLFSPEHSPPPPPRSATRAGS 241
Query: 301 -VANKPLHKSLNLEPSNPDP 319
A KPL ++ PS PDP
Sbjct: 242 GAAPKPLLNGSSVSPSEPDP 301
>TC16587 weakly similar to UP|Q9JGS5 (Q9JGS5) PORF1, partial (3%)
Length = 706
Score = 38.1 bits (87), Expect = 0.003
Identities = 26/91 (28%), Positives = 47/91 (51%)
Frame = +1
Query: 238 SNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEE 297
SN+A T ++S + + S+ S PAS+P+ P S S++ ++ +L +P+T+
Sbjct: 208 SNSA-TNRRSHCSEPRFHSASSACRSNPASSPATPRNSHSTSPHSSNLALPSEPLTALTT 384
Query: 298 SKKVANKPLHKSLNLEPSNPDPAPLPTLRKS 328
K + PS+ PAP P++R+S
Sbjct: 385 PKTPS-----------PSSSKPAPDPSVRRS 444
>TC14217
Length = 985
Score = 36.2 bits (82), Expect = 0.012
Identities = 37/156 (23%), Positives = 66/156 (41%), Gaps = 14/156 (8%)
Frame = +1
Query: 145 EEVAVSRDCYQSSSIEVEESEEQESI-----SHGAEPIDEPEEVVCIKQEENLHVEAEDV 199
E V VS + + +VEE E E+ S E + + V K+EEN + +++
Sbjct: 91 EAVQVSSREVEVETKKVEEQNEAEAATTETESVKVENVKNDTQAVDDKKEENADTKVDEI 270
Query: 200 KEISHVEYKET--EKASEVEAKDVKLDHPKESKVASV-------NRESNAAKTKKKSMLP 250
+ET K + E+ + +D+ ++ +V +ES+ KT K LP
Sbjct: 271 STAISEPVRETLASKFKDEESIETGVDNTEKEQVEEPVKTEVQDTKESDVTKTSKD--LP 444
Query: 251 KSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPS 286
K ++ + +++ S + L A G SPS
Sbjct: 445 KETPAKPAQKQSNNIISKVKQSLAKAKKAITGKSPS 552
>TC14264 similar to UP|Q84NF9 (Q84NF9) Histone H1 (Fragment), partial (76%)
Length = 1277
Score = 35.4 bits (80), Expect = 0.021
Identities = 26/86 (30%), Positives = 36/86 (41%), Gaps = 4/86 (4%)
Frame = +2
Query: 240 AAKTKKKSM-LPKSKASQISTPRNSKPAST---PSKPLTSASSTKKGNSPSLSRKPITSS 295
AA TK K+ PKSKA+ T +KPA+ +KP A K + + K S
Sbjct: 557 AAATKAKAKPAPKSKAAATKTTAKAKPAAAAKPKAKPAAKAKPAAKAKAVAAPAKAKASP 736
Query: 296 EESKKVANKPLHKSLNLEPSNPDPAP 321
+ K A +N+ P P P
Sbjct: 737 AKPKAKAKSKTAPRMNVNPPVPKAKP 814
Score = 29.6 bits (65), Expect = 1.2
Identities = 27/99 (27%), Positives = 39/99 (39%), Gaps = 15/99 (15%)
Frame = +2
Query: 208 KETEKASEVEAKDVKLDHPKESKVASVNRESNA-AKTKKKSMLPKSKASQISTPR----- 261
K T KA A K ++K A+ + A AK K PK+KA + PR
Sbjct: 614 KTTAKAKPAAAAKPKAKPAAKAKPAAKAKAVAAPAKAKASPAKPKAKAKSKTAPRMNVNP 793
Query: 262 ---------NSKPASTPSKPLTSASSTKKGNSPSLSRKP 291
+KPA+ P+K +++ T G KP
Sbjct: 794 PVPKAKPAAKAKPAARPAKASRTSTRTTPGKKVPPPPKP 910
>TC16374 similar to UP|Q7LZA6 (Q7LZA6) Protamine II (Fragment), partial
(63%)
Length = 1130
Score = 35.0 bits (79), Expect = 0.028
Identities = 47/232 (20%), Positives = 97/232 (41%), Gaps = 29/232 (12%)
Frame = +3
Query: 78 LLAQEKETEKDSFGSDDQNGVDLSGNDCGTDAEFDVSNAQ--GSTTEGVEQEISSVGEV- 134
L + E++ K F D+ D S + +++ +N GST+ G E E SSV EV
Sbjct: 99 LASLERKMIKRRFYRLDRGDKDASDASSSEEDDYEQNNDDDAGSTSSGYESEDSSVKEVD 278
Query: 135 --SRNLLDA------LKEEEVAVSRDC----YQSSSIEVEESEEQESISHGAEPIDEPEE 182
S LL + ++E++V ++R+ Y S +EE+ + I + +
Sbjct: 279 VNSAGLLSSDDDAGTIREKQVLINRELPNKRYSEKSNASAMAEEKPLPTEMTSYILQCKS 458
Query: 183 V---------VCIKQEENLHVEAEDVKEISHVEYKETEKASEVEAK-----DVKLDHPKE 228
V +C+ ++ + ++ H ++ K ++A +++ D E
Sbjct: 459 VFKCRICPRIICLSED----TLRDHLQSKRHARSEKLYKQGRLKAMLNINGEIENDEDSE 626
Query: 229 SKVASVNRESNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTK 280
+++ + + E N K KKK+ + + + + S ++T T + K
Sbjct: 627 TEIQTEDTEENVQKKKKKNTKGEKQNKKRLGKKKSDKSNTRKMRSTKRQAKK 782
>TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, partial
(68%)
Length = 1097
Score = 34.7 bits (78), Expect = 0.036
Identities = 33/86 (38%), Positives = 41/86 (47%), Gaps = 7/86 (8%)
Frame = +1
Query: 239 NAAKTKKKSMLPKSKASQISTPRN----SKPAST-PSKP--LTSASSTKKGNSPSLSRKP 291
N + S P S ISTP + KPAST PS P TS+SS+ N+P RKP
Sbjct: 499 NTTPSSLSSAPPSPTTSPISTPSSPENQEKPASTPPSAPPRSTSSSSSSPINTP---RKP 669
Query: 292 ITSSEESKKVANKPLHKSLNLEPSNP 317
S +K A+ P L+L S P
Sbjct: 670 ---SSATKAPASSPPGSELSLHHSPP 738
>TC7865 homologue to UP|SAHH_ARATH (O23255) Adenosylhomocysteinase
(S-adenosyl-L-homocysteine hydrolase) (AdoHcyase) ,
complete
Length = 1852
Score = 34.3 bits (77), Expect = 0.047
Identities = 31/99 (31%), Positives = 42/99 (42%), Gaps = 9/99 (9%)
Frame = +2
Query: 240 AAKTKKKSML-PKSKASQISTPRNSKPASTPSKPLTSASSTKKG-NSPSLSRKPI----- 292
AA T+ ++ L P S AS S+PR+ PAS P+ P ++ S K SP S P
Sbjct: 95 AASTRSRTSLRPTSAASSSSSPRSRCPASWPAVPSSALLSPSKALESPDPST*PFKPPSS 274
Query: 293 --TSSEESKKVANKPLHKSLNLEPSNPDPAPLPTLRKSF 329
S + + A P S P P P P + F
Sbjct: 275 LKPSPPSAPRFAGAPATSS---PPRTTPPLPSPAIALRF 382
>TC10461 homologue to UP|BAD11346 (BAD11346) BRI1-KD interacting protein 118
(Fragment), partial (7%)
Length = 636
Score = 33.9 bits (76), Expect = 0.062
Identities = 19/42 (45%), Positives = 27/42 (64%), Gaps = 2/42 (4%)
Frame = +1
Query: 448 KEEKEADMKKLKHN--FKATPLPAFYGGQKVSKVRPEKVDAK 487
KE +EA++KKL+ + FKA P+P+FY K P KV+ K
Sbjct: 19 KENQEAEIKKLRKSLAFKAAPMPSFY------KEPPPKVELK 126
>AV409862
Length = 420
Score = 33.9 bits (76), Expect = 0.062
Identities = 25/77 (32%), Positives = 33/77 (42%), Gaps = 1/77 (1%)
Frame = +3
Query: 250 PKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEE-SKKVANKPLHK 308
P A+ +PRN P + P P T+ +S K P+ S TSS SK A+ P
Sbjct: 72 PSPFAASPPSPRNPSPTTLP--PPTTTNSPSKPEPPATSTPSATSSTSASKTAASTPSTP 245
Query: 309 SLNLEPSNPDPAPLPTL 325
S + P P TL
Sbjct: 246 SASSPPPTSPPPSSTTL 296
>TC8093 homologue to UP|EFTU_TOBAC (P41342) Elongation factor Tu,
chloroplast precursor (EF-Tu), partial (58%)
Length = 1146
Score = 33.9 bits (76), Expect = 0.062
Identities = 26/90 (28%), Positives = 40/90 (43%), Gaps = 11/90 (12%)
Frame = +2
Query: 247 SMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNS---PSLSRKP--------ITSS 295
++ P AS S P P S P L +SST++ N+ PS + P +SS
Sbjct: 164 TLTPLPPASS-SDPNYPPPTSPPPSILPQSSSTRRRNNPPPPSTAAPPPSPSAPLAASSS 340
Query: 296 EESKKVANKPLHKSLNLEPSNPDPAPLPTL 325
+ + P S +P +P P+P P+L
Sbjct: 341 ARNPMSTSAPSATSTTAKPPSPPPSPWPSL 430
>TC11548 similar to UP|Q9FG37 (Q9FG37) Gamma-tubulin interacting
protein-like, partial (6%)
Length = 480
Score = 33.9 bits (76), Expect = 0.062
Identities = 28/88 (31%), Positives = 41/88 (45%), Gaps = 2/88 (2%)
Frame = +3
Query: 239 NAAKTKKKSMLPKSKASQISTPRNSK-PASTPSKPLTSASSTKKGNSPSLSRKPITSSEE 297
N++ ++ P + TPRN K P++TPS + A+ST P+ P S+
Sbjct: 42 NSSTVSSATISPPTHHH*TLTPRNFKNPSATPSAS-SPAASTPLSPPPTPPPSPNKSNAV 218
Query: 298 SKKVANKPLH-KSLNLEPSNPDPAPLPT 324
S A P H +S PS+P PL T
Sbjct: 219 SPLTAAPPTHSRSPTYTPSSPPEPPLLT 302
>TC9435 similar to UP|Q9AXJ7 (Q9AXJ7) Phosphatase-like protein Mtc923,
partial (17%)
Length = 486
Score = 33.9 bits (76), Expect = 0.062
Identities = 30/119 (25%), Positives = 46/119 (38%), Gaps = 17/119 (14%)
Frame = +3
Query: 225 HPKESKVASVNRESNAAKTKKKSMLPKSKASQIST--PRNSKPASTPSKPLTSASSTKKG 282
H S N+ S+++K + P S S +T P ++ P P P + + TK
Sbjct: 51 HSARSPPPPNNKPSSSSKATISTSTPPSPPSSTTTTTP*STPPQPPPQPPPPTTTKTKTT 230
Query: 283 NSPSLSRKPITSSEESKKVANKPL---------------HKSLNLEPSNPDPAPLPTLR 326
S P+TS+ A PL H + NL P P+PA + + R
Sbjct: 231 PFHPPSLNPLTSTLPRLPHARLPLPTPPLAPPTSSDPAGHSARNLPPRLPEPAAVESAR 407
>TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor (Atperox
P31) (ATP41) , partial (38%)
Length = 475
Score = 33.5 bits (75), Expect = 0.081
Identities = 38/133 (28%), Positives = 55/133 (40%), Gaps = 15/133 (11%)
Frame = +1
Query: 204 HVEYKETEKASEVEAKDVKLDHPKESKVASVN-RESNAAKTK------KKSMLPK----- 251
H + +S + H ++S S N R N+ K+ KS P
Sbjct: 34 HHHHHHHSSSSSQHSHSSPPSHTRDSHSTSTNTRAHNSPKSSATPSPTNKSPPPPPAPPR 213
Query: 252 --SKASQISTPRNSKP-ASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVANKPLHK 308
S ++ STP + P +S+P P T ++T SPS P TSS E K +N
Sbjct: 214 SASSSTTASTPTAATPPSSSPQLPSTKPNATPTSTSPS-PATPSTSSSELKPPSN----- 375
Query: 309 SLNLEPSNPDPAP 321
+L P+ P PAP
Sbjct: 376 --SLAPT-PSPAP 405
>TC10717 similar to UP|Q8S564 (Q8S564) 4-coumarate:coenzyme A ligase ,
partial (32%)
Length = 691
Score = 33.5 bits (75), Expect = 0.081
Identities = 22/73 (30%), Positives = 30/73 (40%)
Frame = +1
Query: 254 ASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVANKPLHKSLNLE 313
AS +TP S P T + P + ++PS S + S S + PL S
Sbjct: 223 ASSTATPEKSTPMPTSTSP---PGRSPPASTPSASATVM*SCSSSVTAPSLPLPSSAPRL 393
Query: 314 PSNPDPAPLPTLR 326
P P P P P+ R
Sbjct: 394 PEPPSPPPTPSTR 432
>TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;}, partial (25%)
Length = 856
Score = 33.5 bits (75), Expect = 0.081
Identities = 28/84 (33%), Positives = 41/84 (48%), Gaps = 3/84 (3%)
Frame = +2
Query: 225 HPKESKVASVNRESNAAKTK-KKSMLPKSKASQIS--TPRNSKPASTPSKPLTSASSTKK 281
H + S+V + + S++A T + S P + +S S TPR+S TPS +S SS
Sbjct: 119 HGRTSRVHTTTKPSSSAATAGRSSHAPTTTSSPTSAATPRSSPSTGTPSSQRSSPSSPP- 295
Query: 282 GNSPSLSRKPITSSEESKKVANKP 305
S +L RK SS S + P
Sbjct: 296 --SATLHRKSSPSSTSSPARISTP 361
>TC15267 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein), partial (6%)
Length = 1003
Score = 33.5 bits (75), Expect = 0.081
Identities = 21/61 (34%), Positives = 27/61 (43%)
Frame = +1
Query: 238 SNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEE 297
S+ S P A ++ + PAS PS P T+ T +SPS S P TSS
Sbjct: 235 SHRTNNNNLSTQPTPSAPSATSLASRTPASPPSPPPTTTPPTHNPSSPSPSAPPWTSSPP 414
Query: 298 S 298
S
Sbjct: 415 S 417
>TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomerase
precursor , partial (81%)
Length = 1355
Score = 33.5 bits (75), Expect = 0.081
Identities = 22/70 (31%), Positives = 33/70 (46%)
Frame = +2
Query: 255 SQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVANKPLHKSLNLEP 314
S S P P+S+PS SAS + NSP+ S P S+ S++ + S P
Sbjct: 461 SSASAPAPPPPSSSPS----SASFSPPANSPTSSASPPPSARRSRRDPSASRSPSSTTIP 628
Query: 315 SNPDPAPLPT 324
++ P+ PT
Sbjct: 629 ASISPSTAPT 658
>TC16098 homologue to UP|Q8LAQ5 (Q8LAQ5) Flower pigmentation protein ATAN11,
partial (79%)
Length = 919
Score = 33.1 bits (74), Expect = 0.11
Identities = 25/89 (28%), Positives = 35/89 (39%), Gaps = 3/89 (3%)
Frame = +1
Query: 243 TKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEESKKVA 302
T+ K+ SK + STP STP ++A+ + SP S T+S S
Sbjct: 76 TRPKTAPTSSKNAPRSTPTKPHGTSTP*TGASAATKSTASPSPPSSNNTPTASRSSSSTT 255
Query: 303 NKPLHKSLNLEPSNP---DPAPLPTLRKS 328
+ PSN P P +L KS
Sbjct: 256 PTARSDPTRISPSNTHTHPPRPFSSLTKS 342
>BP041908
Length = 487
Score = 32.7 bits (73), Expect = 0.14
Identities = 32/109 (29%), Positives = 44/109 (40%), Gaps = 10/109 (9%)
Frame = +3
Query: 230 KVASVNRESNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSR 289
K S + S+ T + L AS+IS PAS+ P TS +T + P
Sbjct: 105 KSNSFSGPSSLTSTSPNTFLSTLTASKISLTLPHDPASSTPPPATSTPTTTSTSPPGEPP 284
Query: 290 KPITSSEESKKVANK------PLHKSLNLEPSNPD----PAPLPTLRKS 328
T+S SKK + PL+ S PS+P P P P + S
Sbjct: 285 PASTNSATSKKETSS*SSSPTPLNSS---SPSSPHLTSAPWPPPQIPSS 422
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.302 0.120 0.317
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,585,140
Number of Sequences: 28460
Number of extensions: 80046
Number of successful extensions: 639
Number of sequences better than 10.0: 153
Number of HSP's better than 10.0 without gapping: 604
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 633
length of query: 492
length of database: 4,897,600
effective HSP length: 94
effective length of query: 398
effective length of database: 2,222,360
effective search space: 884499280
effective search space used: 884499280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 17 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0028a.11


