

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0026.15
(145 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
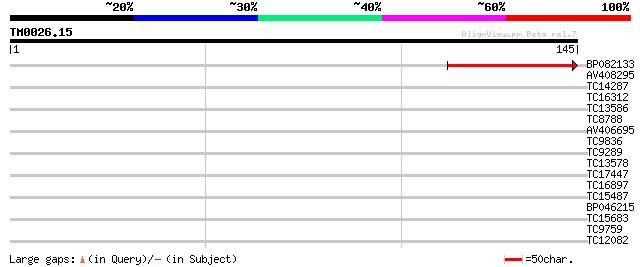
Score E
Sequences producing significant alignments: (bits) Value
BP082133 40 1e-04
AV408295 27 1.8
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 27 1.8
TC16312 similar to GB|AAM65771.1|21593804|AY088230 ribosomal pro... 26 2.4
TC13586 similar to UP|Q84WJ1 (Q84WJ1) At5g49570, partial (14%) 26 3.1
TC8788 UP|Q84LE3 (Q84LE3) Squalene synthase, complete 26 3.1
AV406695 26 3.1
TC9836 weakly similar to UP|Q9LXV0 (Q9LXV0) Glucosyltransferase-... 25 5.3
TC9289 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/MR... 25 5.3
TC13578 similar to UP|Q7RZL8 (Q7RZL8) Predicted protein, partial... 25 6.9
TC17447 weakly similar to UP|Q8L9I2 (Q8L9I2) Nodulin MtN21-like ... 25 6.9
TC16897 homologue to PIR|T09261|T09261 JUN kinase-activation-dom... 25 6.9
TC15487 similar to UP|O23144 (O23144) Proton pump interactor (AT... 25 6.9
BP046215 24 9.0
TC15683 weakly similar to UP|Q8GTM1 (Q8GTM1) WD40, partial (4%) 24 9.0
TC9759 24 9.0
TC12082 weakly similar to UP|Q40385 (Q40385) Pistil extensin-lik... 24 9.0
>BP082133
Length = 395
Score = 40.4 bits (93), Expect = 1e-04
Identities = 17/33 (51%), Positives = 27/33 (81%)
Frame = -1
Query: 113 MEEEELPPLEGELWNWEDPPDDELRARSFVGMA 145
++EEELPPL+GE N ++P +DE++A+SF +A
Sbjct: 386 VKEEELPPLQGES*NMKNPDEDEVKAQSFFRLA 288
>AV408295
Length = 426
Score = 26.6 bits (57), Expect = 1.8
Identities = 16/41 (39%), Positives = 23/41 (56%)
Frame = +3
Query: 84 KSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGE 124
K+E EK P E +++ E V+ + KP EEE+ EGE
Sbjct: 210 KTEEEKKPAE---EEKKEEVVEEE--KKPAEEEKKAEAEGE 317
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial (75%)
Length = 1999
Score = 26.6 bits (57), Expect = 1.8
Identities = 20/81 (24%), Positives = 38/81 (46%), Gaps = 4/81 (4%)
Frame = -3
Query: 18 EDRDRAWVNDEHQVRRIYVGSAAVRSFDQRGRWCETYLMNGRVCLPYAPRSFELPIVVGS 77
E R ++ + + ++R+ S+ +R + R WC ++G LP+ P L
Sbjct: 1682 EGRSHLFLTELNILQRV---SSRLRKLNHRHLWCHLKHLHGHCQLPFPPLLHPL----HP 1524
Query: 78 HQLLNMKSEPE----KDPLEL 94
H+ L + +PE + PL+L
Sbjct: 1523 HRELLLPRDPEIQLLRHPLKL 1461
>TC16312 similar to GB|AAM65771.1|21593804|AY088230 ribosomal protein S1
{Arabidopsis thaliana;}, partial (5%)
Length = 691
Score = 26.2 bits (56), Expect = 2.4
Identities = 12/36 (33%), Positives = 18/36 (49%)
Frame = -2
Query: 96 PDDENEHYVDSKLMSKPMEEEELPPLEGELWNWEDP 131
P+ E + MSK M + LE E+W+W +P
Sbjct: 327 PEKEKVARIAITAMSKQMRKRYRVVLEMEVWDWGNP 220
>TC13586 similar to UP|Q84WJ1 (Q84WJ1) At5g49570, partial (14%)
Length = 467
Score = 25.8 bits (55), Expect = 3.1
Identities = 7/37 (18%), Positives = 24/37 (63%)
Frame = +3
Query: 75 VGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSK 111
+ +++L++ + PE+DP++ + + N+ + +++ K
Sbjct: 33 LAAYELMSANNAPERDPMDWILEGSNDKGISWQVLDK 143
>TC8788 UP|Q84LE3 (Q84LE3) Squalene synthase, complete
Length = 1892
Score = 25.8 bits (55), Expect = 3.1
Identities = 8/16 (50%), Positives = 13/16 (81%)
Frame = +3
Query: 14 ILHDEDRDRAWVNDEH 29
ILH E+R+R W+N ++
Sbjct: 1386 ILHSEERERIWLNTDY 1433
>AV406695
Length = 421
Score = 25.8 bits (55), Expect = 3.1
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = -1
Query: 38 SAAVRSFDQRGRWCETYLMNG 58
+ AVR+FD R W +L+NG
Sbjct: 199 ATAVRTFDLRREWALLFLING 137
>TC9836 weakly similar to UP|Q9LXV0 (Q9LXV0) Glucosyltransferase-like
protein, partial (55%)
Length = 1286
Score = 25.0 bits (53), Expect = 5.3
Identities = 11/39 (28%), Positives = 19/39 (48%)
Frame = -2
Query: 82 NMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPP 120
++ S+P P++ P D + S L P + + LPP
Sbjct: 892 SLHSQPHSSPIQSSPPDPHA*PHTSSLSQPPHKHQALPP 776
>TC9289 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/MRP15_10),
partial (36%)
Length = 994
Score = 25.0 bits (53), Expect = 5.3
Identities = 16/53 (30%), Positives = 26/53 (48%), Gaps = 8/53 (15%)
Frame = +3
Query: 74 VVGSHQLLNMKSEPEKDPLELVPD-------DENEHYVDSK-LMSKPMEEEEL 118
V G + N+ +E EKD +PD D +E Y D+K + K E+++
Sbjct: 393 VKGQNLPANLTTEEEKDKTSNIPDREQYNKADFSEEYTDAKFFVIKSYSEDDI 551
>TC13578 similar to UP|Q7RZL8 (Q7RZL8) Predicted protein, partial (4%)
Length = 416
Score = 24.6 bits (52), Expect = 6.9
Identities = 9/24 (37%), Positives = 16/24 (66%)
Frame = -3
Query: 6 RTGHSWEIILHDEDRDRAWVNDEH 29
+T HS EI + R+++W+ +EH
Sbjct: 234 KTLHSTEIDIRTRVRNKSWIIEEH 163
>TC17447 weakly similar to UP|Q8L9I2 (Q8L9I2) Nodulin MtN21-like protein,
partial (24%)
Length = 590
Score = 24.6 bits (52), Expect = 6.9
Identities = 10/40 (25%), Positives = 21/40 (52%), Gaps = 3/40 (7%)
Frame = +1
Query: 18 EDRDRAWVND---EHQVRRIYVGSAAVRSFDQRGRWCETY 54
E + + W+ HQ++R+ + A++ Q+ +W E Y
Sbjct: 289 EKQQKKWMMSLGRNHQLQRMLLSCKAIKLSGQKRQWMELY 408
>TC16897 homologue to PIR|T09261|T09261 JUN kinase-activation-domain-binding
protein homolog - alfalfa {Medicago
sativa;}, partial (37%)
Length = 505
Score = 24.6 bits (52), Expect = 6.9
Identities = 7/22 (31%), Positives = 14/22 (62%)
Frame = +2
Query: 20 RDRAWVNDEHQVRRIYVGSAAV 41
R++ W ND H +R+ + + A+
Sbjct: 248 REKPWANDPHYFKRVKISALAL 313
>TC15487 similar to UP|O23144 (O23144) Proton pump interactor
(AT4g27500/F27G19_100), partial (16%)
Length = 1153
Score = 24.6 bits (52), Expect = 6.9
Identities = 16/53 (30%), Positives = 24/53 (45%), Gaps = 2/53 (3%)
Frame = +2
Query: 84 KSEPEKDPLELVPDDENEHYVDSKLMSKPMEE--EELPPLEGELWNWEDPPDD 134
K EP+ P E +P + L +KP ++ EE E E+ E PP +
Sbjct: 260 KEEPKPSPQETLPTQKESKNKVRDLKAKPDKKDLEEADEYEFEVPRKETPPKE 418
>BP046215
Length = 255
Score = 24.3 bits (51), Expect = 9.0
Identities = 7/12 (58%), Positives = 9/12 (74%)
Frame = +1
Query: 120 PLEGELWNWEDP 131
P EG+ W+W DP
Sbjct: 193 PEEGQSWSWNDP 228
>TC15683 weakly similar to UP|Q8GTM1 (Q8GTM1) WD40, partial (4%)
Length = 957
Score = 24.3 bits (51), Expect = 9.0
Identities = 10/38 (26%), Positives = 17/38 (44%)
Frame = -2
Query: 25 VNDEHQVRRIYVGSAAVRSFDQRGRWCETYLMNGRVCL 62
V+ E+Q R R + WC+ + +G +CL
Sbjct: 119 VSHENQKRERASSEQICRKVEPDDYWCQQQIFDGHLCL 6
>TC9759
Length = 584
Score = 24.3 bits (51), Expect = 9.0
Identities = 22/83 (26%), Positives = 32/83 (38%), Gaps = 8/83 (9%)
Frame = -2
Query: 45 DQRGRWCET--------YLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVP 96
D R R C T YLM + Y P+ +L +V S L ++ +D + L
Sbjct: 463 DTRARKCFTTLS*QWDKYLM*SHIMTVYIPKFTQLHVVTKSDSLKILQH**MQDTVHLCQ 284
Query: 97 DDENEHYVDSKLMSKPMEEEELP 119
H +SK+ S LP
Sbjct: 283 PLTRYHR*ESKMYSVDSPFSSLP 215
>TC12082 weakly similar to UP|Q40385 (Q40385) Pistil extensin-like protein,
partial (11%)
Length = 308
Score = 24.3 bits (51), Expect = 9.0
Identities = 7/17 (41%), Positives = 11/17 (64%)
Frame = -3
Query: 115 EEELPPLEGELWNWEDP 131
EE P ++ W+WE+P
Sbjct: 69 EESKPQMQCRFWSWENP 19
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.137 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,922,953
Number of Sequences: 28460
Number of extensions: 39688
Number of successful extensions: 207
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 207
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 207
length of query: 145
length of database: 4,897,600
effective HSP length: 82
effective length of query: 63
effective length of database: 2,563,880
effective search space: 161524440
effective search space used: 161524440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0026.15


