

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0024.11
(148 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
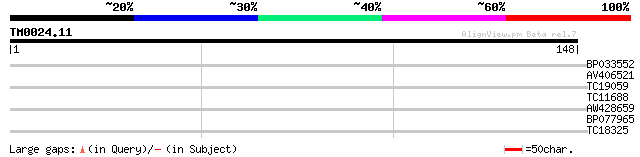
Score E
Sequences producing significant alignments: (bits) Value
BP033552 28 0.65
AV406521 26 3.2
TC19059 weakly similar to PIR|E86340|E86340 protein F2D10.32 [im... 25 4.2
TC11688 weakly similar to UP|Q9M269 (Q9M269) Serine/threonine-pr... 25 5.5
AW428659 25 5.5
BP077965 25 7.2
TC18325 24 9.4
>BP033552
Length = 505
Score = 28.1 bits (61), Expect = 0.65
Identities = 16/30 (53%), Positives = 19/30 (63%)
Frame = +2
Query: 20 FSTIKFRNHVHQGILREILASEFQNSKQIS 49
F +F V ILR +LA+EFQNSKQ S
Sbjct: 155 FLRFRFFLRVPLRILR*VLATEFQNSKQDS 244
>AV406521
Length = 267
Score = 25.8 bits (55), Expect = 3.2
Identities = 14/45 (31%), Positives = 22/45 (48%), Gaps = 3/45 (6%)
Frame = +3
Query: 15 KSDYDFSTIKFRNHVHQGILR---EILASEFQNSKQISKIWEKRY 56
K Y S +F N H +R E + EF+NS+ + +W+ Y
Sbjct: 126 KGYYRRSEDEFSNSSHTFCMRHLSESIGKEFKNSRLVHLLWKAAY 260
>TC19059 weakly similar to PIR|E86340|E86340 protein F2D10.32 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(26%)
Length = 438
Score = 25.4 bits (54), Expect = 4.2
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +2
Query: 98 KRIFKFSKDSKSHLKRSSKSEIAYS 122
+R FKF +SKS ++RSS S ++
Sbjct: 98 RRAFKFEFESKSQVQRSSYSSSTFT 172
>TC11688 weakly similar to UP|Q9M269 (Q9M269) Serine/threonine-protein
kinase-like protein, partial (6%)
Length = 445
Score = 25.0 bits (53), Expect = 5.5
Identities = 14/43 (32%), Positives = 22/43 (50%)
Frame = +1
Query: 91 TLVEGKIKRIFKFSKDSKSHLKRSSKSEIAYSGHARRKAANSC 133
+L+ KRI ++ + + HLK SS+S + K NSC
Sbjct: 61 SLLAADKKRILQYIRHLQFHLKSSSESSSKEADALAAK*INSC 189
>AW428659
Length = 281
Score = 25.0 bits (53), Expect = 5.5
Identities = 13/39 (33%), Positives = 21/39 (53%)
Frame = +2
Query: 105 KDSKSHLKRSSKSEIAYSGHARRKAANSCHAYTDSTRES 143
KDS SH K +S+SE G+ A + ++ S+ +S
Sbjct: 101 KDSSSHFKNASQSENQMVGNTGAGAGPTLYSSLGSSTDS 217
>BP077965
Length = 275
Score = 24.6 bits (52), Expect = 7.2
Identities = 13/46 (28%), Positives = 22/46 (47%)
Frame = +3
Query: 42 FQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPESEYYRAYSPEF 87
F Q +IW +Y + KR+ + V+ L + + +SPEF
Sbjct: 135 FLKKDQRKRIWYLKY*QFIKRYIKKDIFQVSSL*PIKGFCGHSPEF 272
>TC18325
Length = 513
Score = 24.3 bits (51), Expect = 9.4
Identities = 12/38 (31%), Positives = 19/38 (49%)
Frame = -2
Query: 58 KSQKRWFRSKEISVARLPESEYYRAYSPEFRNETLVEG 95
KS+ + + + L ES YY+ + N+TLV G
Sbjct: 284 KSKTKTVSALSTRICYLIESHYYQLVATRTGNQTLVSG 171
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,215,451
Number of Sequences: 28460
Number of extensions: 24113
Number of successful extensions: 131
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 131
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 131
length of query: 148
length of database: 4,897,600
effective HSP length: 82
effective length of query: 66
effective length of database: 2,563,880
effective search space: 169216080
effective search space used: 169216080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0024.11


