

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.17
(341 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
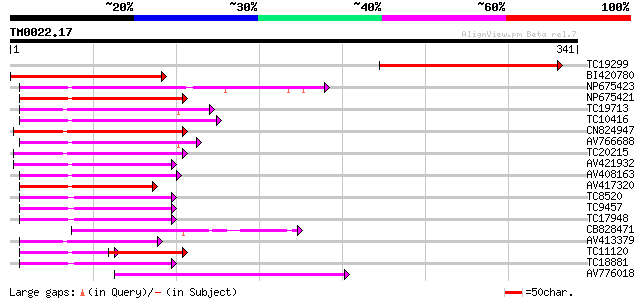
Score E
Sequences producing significant alignments: (bits) Value
TC19299 similar to UP|Q9FUK9 (Q9FUK9) MYB-related transcription ... 201 1e-52
BI420780 197 2e-51
NP675423 transcription factor MYB101 [Lotus japonicus] 97 3e-21
NP675421 transcription factor MYB103 [Lotus japonicus] 94 4e-20
TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%) 93 6e-20
TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial... 91 3e-19
CN824947 89 1e-18
AV766688 89 1e-18
TC20215 similar to AAS10104 (AAS10104) MYB transcription factor,... 87 5e-18
AV421932 82 2e-16
AV408163 80 4e-16
AV417320 77 3e-15
TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 75 1e-14
TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 75 1e-14
TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose... 73 8e-14
CB828471 72 1e-13
AV413379 72 1e-13
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 52 2e-13
TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose... 68 2e-12
AV776018 44 4e-05
>TC19299 similar to UP|Q9FUK9 (Q9FUK9) MYB-related transcription factor
PHAN1, partial (30%)
Length = 511
Score = 201 bits (511), Expect = 1e-52
Identities = 104/110 (94%), Positives = 105/110 (94%)
Frame = +1
Query: 223 HILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKA 282
HILMSELVGFS+ELEEG RALAAHKKEAEWRLRRLEL LESEKACRRRETVEEFEANIKA
Sbjct: 1 HILMSELVGFSQELEEGPRALAAHKKEAEWRLRRLELPLESEKACRRRETVEEFEANIKA 180
Query: 283 LQEEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLE 332
L EEQTAALNRIENACREQLGGLRRDAESKE KLAEKWTSK LRLTRLLE
Sbjct: 181 LPEEQTAALNRIENACREQLGGLRRDAESKEPKLAEKWTSKPLRLTRLLE 330
>BI420780
Length = 475
Score = 197 bits (502), Expect = 2e-51
Identities = 94/94 (100%), Positives = 94/94 (100%)
Frame = +1
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG
Sbjct: 193 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 372
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAK 94
SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAK
Sbjct: 373 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAK 474
>NP675423 transcription factor MYB101 [Lotus japonicus]
Length = 1047
Score = 97.4 bits (241), Expect = 3e-21
Identities = 62/192 (32%), Positives = 95/192 (49%), Gaps = 6/192 (3%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EEDA+L Y+Q++G W + + LNR KSC RW NYL+P IK+G T+EE
Sbjct: 49 WTPEEDAILVDYIQKHGHGSWRALPKLAG--LNRCGKSCRLRWTNYLRPDIKRGKFTEEE 222
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
++L+I L A GNKW IA +PGRT + +W + +K K+ G+ +
Sbjct: 223 EQLIINLHAVLGNKWSAIAGHLPGRTDNEIKNFWNTHLKK----KLLQMGLDPVTHRPRL 390
Query: 127 EH--MLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLP--SWLSNYDST--STPPS 180
+H +L + L + + SF SN A + + L L N + P S
Sbjct: 391 DHLNLLSNLQQFLAATNMVTSFTNTLDSNAALRLQSDATQLAKLQLLQNILQVLGTNPAS 570
Query: 181 SISVTLSLSPST 192
++ + L PS+
Sbjct: 571 NLELLNQLGPSS 606
>NP675421 transcription factor MYB103 [Lotus japonicus]
Length = 924
Score = 93.6 bits (231), Expect = 4e-20
Identities = 44/101 (43%), Positives = 62/101 (60%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
WS EED +L Y+Q++G W + Q LNR KSC RW NYL+P IK+G + EE
Sbjct: 49 WSQEEDKVLVDYIQKHGHGSWRALPQLAG--LNRCGKSCRLRWTNYLRPDIKRGKFSDEE 222
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
++L+I L A+ GNKW IA+ +PGRT + W + +K+
Sbjct: 223 EQLIINLHASLGNKWATIASHLPGRTDNEIKNLWNTHLKKK 345
>TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%)
Length = 532
Score = 93.2 bits (230), Expect = 6e-20
Identities = 45/119 (37%), Positives = 71/119 (58%), Gaps = 2/119 (1%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L Y+ +G WN +++ + L R KSC RW NYL+P +++G++T EE
Sbjct: 120 WTIEEDLILINYIANHGEGVWNTLAK--SAGLKRTGKSCRLRWLNYLRPDVRRGNITPEE 293
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW--EVYKEKQQREKIEINGIVSPISD 123
Q L++ L A +GN+W KIA +PGRT + +W + K +Q E ++ S I+D
Sbjct: 294 QLLIMELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQKHIKQAESLQQQQSSSEIND 470
>TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial (41%)
Length = 1046
Score = 90.9 bits (224), Expect = 3e-19
Identities = 45/121 (37%), Positives = 69/121 (56%)
Frame = +2
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W++EED L AYV +YG W + + L R KSC RW NYL+P IK+G+ T+EE
Sbjct: 140 WTAEEDKKLIAYVTRYGCWNWRQLPKFAG--LARCGKSCRLRWLNYLRPNIKRGNFTQEE 313
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
+ ++I + GN+W IAAE+PGRT + W ++ E+ + + +S++K
Sbjct: 314 EEIIIRMHKKLGNRWSTIAAELPGRTDSEVKNHWHTTLKRTVHEQNTVTNEETKVSNSKN 493
Query: 127 E 127
E
Sbjct: 494 E 496
>CN824947
Length = 738
Score = 88.6 bits (218), Expect = 1e-18
Identities = 40/105 (38%), Positives = 65/105 (61%)
Frame = +1
Query: 3 ERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSL 62
+R RW++EED LL Y+Q G W + + N L R KSC RW NYL+ +K+G++
Sbjct: 187 KRGRWTAEEDELLTKYIQASGEGSWRSLPK--NAGLLRCGKSCRLRWINYLRGDLKRGNI 360
Query: 63 TKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
+ EE+ L++ L A++GN+W IA+ +PGRT + +W + ++
Sbjct: 361 SDEEESLIVKLHASFGNRWSLIASHMPGRTDNEIKNYWNSHLSRK 495
>AV766688
Length = 608
Score = 88.6 bits (218), Expect = 1e-18
Identities = 45/111 (40%), Positives = 66/111 (58%), Gaps = 2/111 (1%)
Frame = +2
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W++EED L AYV +YG W + + L R KSC RW NYL+P IK+G T+EE
Sbjct: 125 WTAEEDRKLIAYVTRYGCWNWRQLPKFAG--LARCGKSCRLRWLNYLRPDIKRGHFTQEE 298
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW--EVYKEKQQREKIEIN 115
+ ++I + N GN+W IAAE+PGRT + W + + QQ E+ +++
Sbjct: 299 EEIIIRMHKNLGNRWSIIAAELPGRTDNEVKNHWHTSLKRRVQQNEEAKVS 451
>TC20215 similar to AAS10104 (AAS10104) MYB transcription factor, partial
(35%)
Length = 464
Score = 86.7 bits (213), Expect = 5e-18
Identities = 39/105 (37%), Positives = 62/105 (58%)
Frame = +2
Query: 3 ERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSL 62
+R RWS EED +L +++Q G W + + N L R KSC RW NYL+ +K+G++
Sbjct: 65 KRGRWSEEEDKILTDFIKQNGEGSWKSLPK--NAGLLRCGKSCRLRWINYLREDVKRGNI 238
Query: 63 TKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
T EE+ +++ L A GN+W IA +PGRT + +W + ++
Sbjct: 239 TSEEEEIIVKLHAALGNRWSVIAGHLPGRTDNEIKNYWNSHLRRK 373
>AV421932
Length = 402
Score = 81.6 bits (200), Expect = 2e-16
Identities = 40/98 (40%), Positives = 57/98 (57%)
Frame = +3
Query: 3 ERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSL 62
+R WS EED L Y+ +G W+ V ++ L R KSC RW NYL+P I++G
Sbjct: 114 KRGLWSPEEDEKLIRYITTHGYGCWSEVPEKAG--LQRCGKSCRLRWINYLRPDIRRGRF 287
Query: 63 TKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW 100
T EE++L+I L GN+W IA+ +PGRT + +W
Sbjct: 288 TPEEEKLIISLHGVVGNRWAHIASHLPGRTDNEIKNYW 401
>AV408163
Length = 425
Score = 80.5 bits (197), Expect = 4e-16
Identities = 38/97 (39%), Positives = 56/97 (57%)
Frame = +2
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED L +Y++ +G W + + L R KSC RW NYL+P +K+G+ T+EE
Sbjct: 125 WTKEEDDRLISYIRAHGEGCWRSLPKAAG--LLRCGKSCRLRWINYLRPDLKRGNFTEEE 298
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVY 103
L+I L + GNKW IA +PGRT + +W +
Sbjct: 299 DELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTH 409
>AV417320
Length = 369
Score = 77.4 bits (189), Expect = 3e-15
Identities = 36/83 (43%), Positives = 53/83 (63%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+SEED +L +Y+Q++G W + + L R KSC RW NYL+P IK+G+ T EE
Sbjct: 126 WTSEEDQILISYIQKHGHGNWRALPKHAG--LLRCGKSCRLRWINYLRPDIKRGNFTAEE 299
Query: 67 QRLVILLQANYGNKWKKIAAEVP 89
+ L+I + GN+W IAA++P
Sbjct: 300 EELIIKMHELLGNRWSAIAAKLP 368
>TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (36%)
Length = 1778
Score = 75.5 bits (184), Expect = 1e-14
Identities = 37/94 (39%), Positives = 51/94 (53%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
WS EED L V+++GPR W+L+S+ + R KSC RW N L P ++ + T EE
Sbjct: 193 WSPEEDEALQKLVERHGPRNWSLISKSIP---GRSGKSCRLRWCNQLSPQVEHRAFTPEE 363
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW 100
+I A +GNKW IA + GRT + W
Sbjct: 364 DDTIIRAHARFGNKWATIARLLSGRTDNAIKNHW 465
>TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (43%)
Length = 860
Score = 75.5 bits (184), Expect = 1e-14
Identities = 37/94 (39%), Positives = 51/94 (53%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
WS EED L V+++GPR W+L+S+ + R KSC RW N L P ++ + T EE
Sbjct: 87 WSPEEDEALQRLVEKHGPRNWSLISKSIP---GRSGKSCRLRWCNQLSPQVEHRAFTAEE 257
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW 100
+I A +GNKW IA + GRT + W
Sbjct: 258 DDTIIRAHARFGNKWATIARLLSGRTDNAIKNHW 359
>TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (50%)
Length = 487
Score = 72.8 bits (177), Expect = 8e-14
Identities = 34/94 (36%), Positives = 52/94 (55%)
Frame = +2
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
WS+EED +L V+++G R W+L+S+ + R KSC RW N L P ++ + +E
Sbjct: 179 WSAEEDRILTRLVERHGARNWSLISRYIK---GRSGKSCRLRWCNQLSPTVEHRPFSVQE 349
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW 100
+I A +GN+W IA +PGRT + W
Sbjct: 350 DETIIAAHARFGNRWAAIARLLPGRTDNAVKNHW 451
>CB828471
Length = 525
Score = 72.0 bits (175), Expect = 1e-13
Identities = 47/146 (32%), Positives = 75/146 (51%), Gaps = 7/146 (4%)
Frame = +1
Query: 38 LNRDTKSCLERWKNYLKPGIKKGSLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLG 97
L R KSC RW NYL+P IK+G+ ++EE+ +I L GN+W IAA +PGRT +
Sbjct: 16 LLRCGKSCRLRWANYLRPDIKRGNFSREEEDEIINLHELLGNRWSAIAARLPGRTDNEIK 195
Query: 98 KWWEVY-------KEKQQREKIEINGIVSPISDTKYEHMLEGFAEKLVKEHTLPSFAMAA 150
W + + KQ++ ++++ S D K EH + + P ++
Sbjct: 196 NVWHTHLKKRLQPQTKQKQTNLDLDASKSN-KDAKKEHQED-------EGPRSPQQCSSS 351
Query: 151 SSNEAFLHTNSSAMLPSWLSNYDSTS 176
+S+ TNSS++ PS +N+D S
Sbjct: 352 NSDNMSSLTNSSSIAPS--TNHDDMS 423
>AV413379
Length = 416
Score = 72.0 bits (175), Expect = 1e-13
Identities = 36/86 (41%), Positives = 51/86 (58%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED L AY++ +G W + + + L R KSC RW NYL+P +K+G+ T +E
Sbjct: 150 WTKEEDDRLIAYIRAHGEGCWRSLPK--SAGLLRCGKSCRLRWINYLRPDLKRGNFTPQE 323
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRT 92
L+I L + GNKW IA + GRT
Sbjct: 324 DELIIKLHSLLGNKWSLIAGRLAGRT 401
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 52.4 bits (124), Expect(2) = 2e-13
Identities = 25/60 (41%), Positives = 35/60 (57%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED L ++Q++G W + + LNR KSC RW NYL+P IK+G + EE
Sbjct: 115 WTPEEDQKLVEHIQKHGHGSWRALPKLAG--LNRCGKSCRLRWTNYLRPDIKRGKFSPEE 288
Score = 39.3 bits (90), Expect(2) = 2e-13
Identities = 14/48 (29%), Positives = 29/48 (60%)
Frame = +2
Query: 60 GSLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
G+ +++++ ++ L + GNKW IA +PGRT + +W + +K+
Sbjct: 269 GNFLQKKEQTILHLHSILGNKWSTIATHLPGRTDNEIKNFWNTHLKKK 412
>TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose-responsive
element binding factor, partial (28%)
Length = 346
Score = 68.2 bits (165), Expect = 2e-12
Identities = 36/94 (38%), Positives = 46/94 (48%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
WS EED L VQ +GP+ W +S+ + R KSC RW N L P ++ T EE
Sbjct: 64 WSPEEDEALTRLVQVHGPKNWTAISKSIP---GRSGKSCRLRWCNQLSPEVEHRPFTPEE 234
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW 100
+I A GNKW IA + GRT + W
Sbjct: 235 DSTIIRAHARCGNKWATIARILNGRTDNAVKNHW 336
>AV776018
Length = 476
Score = 43.9 bits (102), Expect = 4e-05
Identities = 35/141 (24%), Positives = 59/141 (41%)
Frame = +1
Query: 64 KEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISD 123
KEE L+I L A GN+W + A + PGRT + W +K+ R++ P+S+
Sbjct: 19 KEEDNLIIELHAVLGNRWSQDATQWPGRTDNEIKNLWNSCLKKKLRQRGIDPVTHKPLSE 198
Query: 124 TKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTSTPPSSIS 183
+ + +++ V + + S+ A NSS++ P + S+ S
Sbjct: 199 VENGKEDKNKSQEKVSNELNLLKSESFKSDAASYEQNSSSLAPKAYTTEMEGSSSSDCSS 378
Query: 184 VTLSLSPSTVATPRGLENNAP 204
L L T G+ N P
Sbjct: 379 KDLLLDRFMSQTQNGMGNYYP 441
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.127 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,914,368
Number of Sequences: 28460
Number of extensions: 78358
Number of successful extensions: 583
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 559
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 568
length of query: 341
length of database: 4,897,600
effective HSP length: 91
effective length of query: 250
effective length of database: 2,307,740
effective search space: 576935000
effective search space used: 576935000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0022.17


