

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.15
(313 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
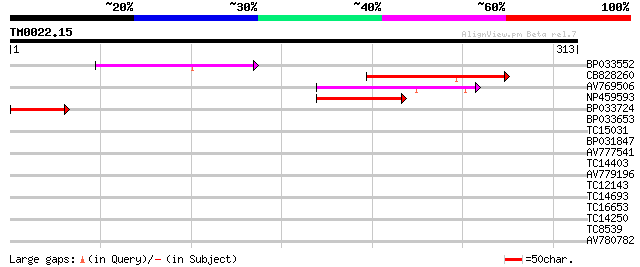
BP033552 94 3e-20
CB828260 79 1e-15
AV769506 62 1e-10
NP459593 Pge1 protein [Lotus japonicus] 60 6e-10
BP033724 49 8e-07
BP033653 36 0.007
TC15031 weakly similar to UP|Q69107 (Q69107) LAT protein, partia... 32 0.18
BP031847 28 2.0
AV777541 27 3.4
TC14403 homologue to UP|Q8LJR4 (Q8LJR4) Syntaxin, partial (38%) 27 3.4
AV779196 27 4.5
TC12143 27 4.5
TC14693 similar to GB|BAB01116.1|11994113|AB026658 CND41, chloro... 26 7.6
TC16653 similar to UP|Q8W1A1 (Q8W1A1) Adenosine 5'-phosphosulfat... 26 9.9
TC14250 homologue to UP|Q8GTQ9 (Q8GTQ9) Succinyl-CoA ligase alph... 26 9.9
TC8539 similar to UP|Q94KE3 (Q94KE3) AT3g52990/F8J2_160 (Pyruvat... 26 9.9
AV780782 26 9.9
>BP033552
Length = 505
Score = 94.0 bits (232), Expect = 3e-20
Identities = 59/112 (52%), Positives = 66/112 (58%), Gaps = 22/112 (19%)
Frame = +1
Query: 48 SSPLLLLKLTGDGKMRENENFAILIFPAPSSVNSKIGFGDRIPKLKRGFLEL-------- 99
S L+ G GK R+N+ FAI IFPA SS +SKIG GDRIP+LK GFLEL
Sbjct: 97 SKNLIFELFIGFGKKRKNDFFAIPIFPARSSAHSKIGSGDRIPELKTGFLELNQKIQNRH 276
Query: 100 --------------SGCRKRKLPSEERLETSREMGVRVWNLHLKLRIEKVFI 137
S R RKLPSEERL RE+G R WNL L+LRIEKVFI
Sbjct: 277 KSV*VGLSRKTTFCSDRRMRKLPSEERLGMMREIGERGWNLLLQLRIEKVFI 432
>CB828260
Length = 557
Score = 78.6 bits (192), Expect = 1e-15
Identities = 39/84 (46%), Positives = 55/84 (65%), Gaps = 5/84 (5%)
Frame = +3
Query: 198 SDNGGL-KRKTFMILGCERCDKYIPYKEVLKHQSTGTKKCYCPFRLRAR----GTKVGWK 252
SD+GG RK +++LGCE+ KY+PY++ + T T+K CPFRL+ R GT W+
Sbjct: 150 SDSGGNGSRKAYVMLGCEKHGKYVPYRDPDLVEGTRTQKTECPFRLKGRPMKNGTDRDWR 329
Query: 253 VSVMNGYHIHERSETLLGHPYVGR 276
+ VM G H HE + +LLGH +VGR
Sbjct: 330 LKVMEGAHNHEPARSLLGHNFVGR 401
>AV769506
Length = 345
Score = 62.4 bits (150), Expect = 1e-10
Identities = 33/102 (32%), Positives = 57/102 (55%), Gaps = 11/102 (10%)
Frame = +1
Query: 170 KVFASRDEILEWARNLGKQHGFIIVITRSDNGGLK-RKTFMILGCERCDKYIPYK----- 223
++F +RD+++ W + ++G++ +IT+SD GG + RK +++LGCE+ KY+PY+
Sbjct: 37 QIFPTRDDLINWVHGIAIENGYVTLITKSDYGGNESRKAYVMLGCEKHGKYVPYRDPDLV 216
Query: 224 --EVLKHQSTGTKKCYCPFRLRARGTKVG---WKVSVMNGYH 260
E QST C C +G+K W S ++G H
Sbjct: 217 ADEDPSAQSTLEPACDCLNHHHKQGSKPSPQLWPPSTVDGPH 342
>NP459593 Pge1 protein [Lotus japonicus]
Length = 633
Score = 59.7 bits (143), Expect = 6e-10
Identities = 23/50 (46%), Positives = 37/50 (74%)
Frame = +1
Query: 170 KVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKY 219
+ FAS ++++WAR +GK++G+++++ RSD G KRK + LGCER KY
Sbjct: 463 EAFASHTDLIDWARCVGKENGYVVIVIRSDYGSAKRKPLITLGCERGGKY 612
>BP033724
Length = 486
Score = 49.3 bits (116), Expect = 8e-07
Identities = 24/33 (72%), Positives = 28/33 (84%)
Frame = +3
Query: 1 MNLRAKTTEFHHNQSETQHSILTKVTRSSIYSE 33
+NL AKTTEF+HNQSETQ + +TKVTRS I SE
Sbjct: 60 LNL*AKTTEFNHNQSETQQTTITKVTRSIINSE 158
>BP033653
Length = 516
Score = 36.2 bits (82), Expect = 0.007
Identities = 20/51 (39%), Positives = 32/51 (62%), Gaps = 8/51 (15%)
Frame = -2
Query: 13 NQSETQHSILTKVTRSSIYSE--------QLSEQRSSKKRRKSSSPLLLLK 55
N+SETQH+I T+V+R+ I SE Q+ E+R+ +SS PL +++
Sbjct: 515 NESETQHTISTRVSRNIINSEVRQSNALRQVFEKRNPPPLMRSSRPLFIIR 363
>TC15031 weakly similar to UP|Q69107 (Q69107) LAT protein, partial (27%)
Length = 997
Score = 31.6 bits (70), Expect = 0.18
Identities = 22/64 (34%), Positives = 32/64 (49%)
Frame = +3
Query: 27 RSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENENFAILIFPAPSSVNSKIGFG 86
RSS S S RS ++RRK P L + + + E+ +F+I PS + S FG
Sbjct: 759 RSSSRSSSSSTHRSRRRRRKKPLPSLASRASPSSRFTESSSFSI-----PSGIASPGRFG 923
Query: 87 DRIP 90
R+P
Sbjct: 924 -RLP 932
>BP031847
Length = 121
Score = 28.1 bits (61), Expect = 2.0
Identities = 12/19 (63%), Positives = 16/19 (84%)
Frame = +1
Query: 61 KMRENENFAILIFPAPSSV 79
+M++NENFAI IFP S+V
Sbjct: 25 EMQKNENFAIPIFPTDSNV 81
>AV777541
Length = 466
Score = 27.3 bits (59), Expect = 3.4
Identities = 17/41 (41%), Positives = 21/41 (50%)
Frame = +1
Query: 27 RSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENEN 67
R IY EQ+ E+ + R SSPLL L DG+ R N
Sbjct: 346 RRKIYLEQIRERANL---RDQSSPLLRRSLNKDGQGRSTPN 459
>TC14403 homologue to UP|Q8LJR4 (Q8LJR4) Syntaxin, partial (38%)
Length = 727
Score = 27.3 bits (59), Expect = 3.4
Identities = 21/90 (23%), Positives = 37/90 (40%)
Frame = -3
Query: 39 RSSKKRRKSSSPLLLLKLTGDGKMRENENFAILIFPAPSSVNSKIGFGDRIPKLKRGFLE 98
+S + +S+ P+L+L L +GK + N+ ILI P + + R L
Sbjct: 608 KSRSAQEESAQPILVLHLQVEGK*K---NYYILILKKPE---------ENLHHKSRHILR 465
Query: 99 LSGCRKRKLPSEERLETSREMGVRVWNLHL 128
R L ++ +R+WN H+
Sbjct: 464 TPNTNSRTLTPHNKIT------IRLWNYHM 393
>AV779196
Length = 509
Score = 26.9 bits (58), Expect = 4.5
Identities = 13/25 (52%), Positives = 18/25 (72%)
Frame = -3
Query: 19 HSILTKVTRSSIYSEQLSEQRSSKK 43
H +L+ VTR S++S SEQR+S K
Sbjct: 390 HVLLSPVTRWSVFSS*PSEQRNSTK 316
>TC12143
Length = 428
Score = 26.9 bits (58), Expect = 4.5
Identities = 21/79 (26%), Positives = 37/79 (46%)
Frame = +2
Query: 3 LRAKTTEFHHNQSETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKM 62
L +KT ++H+ ETQ + TK+ + I + E R++ K S+ + L + +
Sbjct: 98 LVSKTNHYYHHTIETQIKL*TKIYQFFISGIKDRE*RNTMPEEKESTSIPLSQAEDNASC 277
Query: 63 RENENFAILIFPAPSSVNS 81
+ E+ PA S NS
Sbjct: 278 DDPED------PAKSPPNS 316
>TC14693 similar to GB|BAB01116.1|11994113|AB026658 CND41, chloroplast
nucleoid DNA binding protein-like {Arabidopsis
thaliana;} , partial (58%)
Length = 2067
Score = 26.2 bits (56), Expect = 7.6
Identities = 14/41 (34%), Positives = 24/41 (58%), Gaps = 3/41 (7%)
Frame = -3
Query: 87 DRIPKL---KRGFLELSGCRKRKLPSEERLETSREMGVRVW 124
++IP++ +R EL G R+R+ R + +RE+G R W
Sbjct: 229 EKIPQVECYRRELGELGGGRRRRRGRRWRTQKNRELGWRGW 107
>TC16653 similar to UP|Q8W1A1 (Q8W1A1) Adenosine 5'-phosphosulfate
reductase, partial (69%)
Length = 1242
Score = 25.8 bits (55), Expect = 9.9
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = +2
Query: 76 PSSVNSKIGFGDRIPKLK 93
P ++N K+G G R+PKL+
Sbjct: 929 PGNMNGKVGGGGRMPKLR 982
>TC14250 homologue to UP|Q8GTQ9 (Q8GTQ9) Succinyl-CoA ligase alpha subunit
, partial (89%)
Length = 1395
Score = 25.8 bits (55), Expect = 9.9
Identities = 10/31 (32%), Positives = 21/31 (67%)
Frame = +2
Query: 102 CRKRKLPSEERLETSREMGVRVWNLHLKLRI 132
CR ++ + +L+ SR++G ++W+ LKL +
Sbjct: 905 CRVARVLHKTKLKPSRKLGSQLWSPLLKLEL 997
>TC8539 similar to UP|Q94KE3 (Q94KE3) AT3g52990/F8J2_160 (Pyruvate
kinase-like protein), partial (81%)
Length = 1430
Score = 25.8 bits (55), Expect = 9.9
Identities = 25/99 (25%), Positives = 42/99 (42%)
Frame = +2
Query: 6 KTTEFHHNQSETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMREN 65
KT + H+ S TQH+ + ++ + +R ++ SS LLL + + E
Sbjct: 23 KTRDTAHSLSTTQHATTLQPPPP*LFLSDSAARRRAQPNTMPSSHLLLEEPIRMASILEP 202
Query: 66 ENFAILIFPAPSSVNSKIGFGDRIPKLKRGFLELSGCRK 104
+ FPA + + +G PK R +SGC K
Sbjct: 203 SKAS--FFPAMTKIVGTLG-----PK-SRSIEAISGCLK 295
>AV780782
Length = 511
Score = 25.8 bits (55), Expect = 9.9
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = -2
Query: 174 SRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDK 218
SR I+ WA +LG ++ + + S+ G K +LGC+ C +
Sbjct: 387 SRGGIIFWADSLGSEYIYSRLEKWSELYGEFFKPCALLGCKSCQR 253
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,625,101
Number of Sequences: 28460
Number of extensions: 76702
Number of successful extensions: 418
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 414
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 415
length of query: 313
length of database: 4,897,600
effective HSP length: 90
effective length of query: 223
effective length of database: 2,336,200
effective search space: 520972600
effective search space used: 520972600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0022.15


