

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.11
(334 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
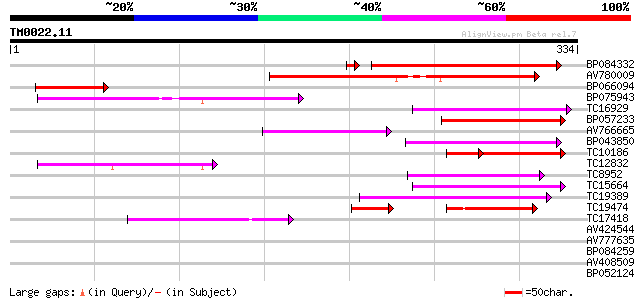
Score E
Sequences producing significant alignments: (bits) Value
BP084332 218 1e-57
AV780009 123 5e-29
BP066094 81 3e-16
BP075943 75 2e-14
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 68 2e-12
BP057233 65 2e-11
AV766665 61 3e-10
BP043850 61 3e-10
TC10186 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprot... 49 3e-10
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 59 1e-09
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 55 1e-08
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 55 2e-08
TC19389 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 52 1e-07
TC19474 weakly similar to UP|Q9FJA1 (Q9FJA1) Similarity to retro... 37 3e-06
TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partia... 47 6e-06
AV424544 37 0.004
AV777635 35 0.014
BP084259 33 0.067
AV408509 32 0.11
BP052124 31 0.33
>BP084332
Length = 368
Score = 218 bits (555), Expect(2) = 1e-57
Identities = 108/112 (96%), Positives = 109/112 (96%)
Frame = +2
Query: 214 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNISIYCDNTAAISLSKNLILHS 273
QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNI IYCDNTAAISLSKN ILHS
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS 211
Query: 274 RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFILKNL 325
RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIF+KPLAEDRF FILKNL
Sbjct: 212 RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNL 367
Score = 21.6 bits (44), Expect(2) = 1e-57
Identities = 8/8 (100%), Positives = 8/8 (100%)
Frame = +1
Query: 199 CQFLGSNL 206
CQFLGSNL
Sbjct: 4 CQFLGSNL 27
>AV780009
Length = 529
Score = 123 bits (308), Expect = 5e-29
Identities = 64/165 (38%), Positives = 102/165 (61%), Gaps = 6/165 (3%)
Frame = -1
Query: 154 RILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKR 213
R+L+Y+KG GL + S KL Y D+D+AG R+S G FLG++L+SW +K+
Sbjct: 517 RVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISWRTKK 338
Query: 214 QSTIALSTAEAEY--ISAAICSTQMLWMKHQLEDYQILESN----ISIYCDNTAAISLSK 267
Q+T++ S++EAEY ++A +C Q W+ + +Q L+ N + ++CDN +A+ ++
Sbjct: 337 QTTVSRSSSEAEYRALAATVCEVQ--WLSYL---FQFLKLNVPLPVPLFCDNQSALHIAH 173
Query: 268 NLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSKP 312
N H R KHIE+ H +R +Q G++ L + T HQ ADIF+KP
Sbjct: 172 NPTFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTKP 38
>BP066094
Length = 532
Score = 80.9 bits (198), Expect = 3e-16
Identities = 39/43 (90%), Positives = 40/43 (92%)
Frame = +2
Query: 16 QIYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQ 58
QIYVDDIIFGSAN LCKEFS+MMQAEFEM MMGELKYFLGIQ
Sbjct: 404 QIYVDDIIFGSANPSLCKEFSEMMQAEFEMRMMGELKYFLGIQ 532
>BP075943
Length = 547
Score = 74.7 bits (182), Expect = 2e-14
Identities = 47/162 (29%), Positives = 76/162 (46%), Gaps = 5/162 (3%)
Frame = -3
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTKE 76
+YVDD++ + + + +F + +GE KYFLG+++ + ++Q KY +
Sbjct: 515 LYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQRKYALQ 336
Query: 77 FLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKL-----YRGMIGSLLYLTASRPDIL 131
+ P + H T + + L YR ++G LLYL +RPDI
Sbjct: 335 LISDSGHFGFQP-RFYSHGT---NSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTTRPDIT 168
Query: 132 FSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS 173
F+V+ +QF S P + H + LKYL G+ GL Y SS
Sbjct: 167 FAVNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFYPASS 42
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 67.8 bits (164), Expect = 2e-12
Identities = 32/96 (33%), Positives = 56/96 (58%), Gaps = 2/96 (2%)
Frame = +1
Query: 238 WMKHQLEDYQI-LESNISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLL 296
W+ + L+D ++ E +YCDN +A ++ N + H R KHIE+ H +R+ +QKG++ L
Sbjct: 4 WLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIHL 183
Query: 297 KFVDTDHQWADIFSKPLAEDRFKFILKNLNM-DLCS 331
+ + ADI++K L+ F I L + ++CS
Sbjct: 184 LPISSSEPLADIYTKALSPQNFHQICAKLGLINICS 291
>BP057233
Length = 473
Score = 65.1 bits (157), Expect = 2e-11
Identities = 29/73 (39%), Positives = 48/73 (65%)
Frame = -2
Query: 255 IYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSKPLA 314
++CDN +A +L+ N +LH+R+KHIE+ H+IRD V + +++ +V T Q AD +KPL+
Sbjct: 445 LWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQIADCLTKPLS 266
Query: 315 EDRFKFILKNLNM 327
RF + L +
Sbjct: 265 HTRFSQLRDKLGV 227
>AV766665
Length = 601
Score = 60.8 bits (146), Expect = 3e-10
Identities = 32/76 (42%), Positives = 43/76 (56%)
Frame = +3
Query: 150 TTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSW 209
T RI YLK + G +++K + G+ ADY G +R ST G FL NLV+W
Sbjct: 360 TFTTRIF*YLKANSRRGPLFQKEGKSSMDGFTYADYLGSIVDRLSTMGYYMFLSGNLVTW 539
Query: 210 ASKRQSTIALSTAEAE 225
SK+Q+ IA S+ EAE
Sbjct: 540 RSKQQNIIARSSGEAE 587
>BP043850
Length = 515
Score = 60.8 bits (146), Expect = 3e-10
Identities = 30/93 (32%), Positives = 53/93 (56%), Gaps = 1/93 (1%)
Frame = -1
Query: 234 TQMLWMKHQLEDYQILESNIS-IYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKG 292
++++W++ L + L+S + ++ DNT+AI ++ N + H +HIEV H +R+ +
Sbjct: 515 SEIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRR 336
Query: 293 VLLLKFVDTDHQWADIFSKPLAEDRFKFILKNL 325
V+ L V T Q ADI +K L R F++ L
Sbjct: 335 VITLPHVSTSVQIADILTKSLTRQRHNFLVSKL 237
>TC10186 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein,
partial (4%)
Length = 528
Score = 48.9 bits (115), Expect(2) = 3e-10
Identities = 25/51 (49%), Positives = 33/51 (64%)
Frame = +3
Query: 277 HIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFILKNLNM 327
+IE KYHF+RD V KG + LK TD Q ADI +K L +RF+ + LN+
Sbjct: 69 NIETKYHFLRDQVTKGKISLKHCGTDLQVADIMTKGLKTERFRNMRAMLNV 221
Score = 31.6 bits (70), Expect(2) = 3e-10
Identities = 13/22 (59%), Positives = 17/22 (77%)
Frame = +2
Query: 258 DNTAAISLSKNLILHSRAKHIE 279
DN +AI L+KN + H R+KHIE
Sbjct: 5 DNKSAIDLAKNPVSHGRSKHIE 70
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 58.5 bits (140), Expect = 1e-09
Identities = 30/114 (26%), Positives = 56/114 (48%), Gaps = 8/114 (7%)
Frame = +2
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQV--DQQPEATYIHQSKYT 74
+YVDD++ N +E + EF+M +G LG+Q+ D++ ++ Q Y
Sbjct: 101 LYVDDMLVVGPNKDRVQELKAQLAREFDMKDLGPANKILGMQIHRDRKDRRIWLSQKNYL 280
Query: 75 KEFLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKL------YRGMIGSLLY 122
++ L++FNM +C P TP+ L + + +++ Y +GSL+Y
Sbjct: 281 QKVLRRFNMQDCNPISTPLPVNYKLSSSMIPSSEAERMEMSRVPYASAVGSLMY 442
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 55.5 bits (132), Expect = 1e-08
Identities = 28/82 (34%), Positives = 48/82 (58%), Gaps = 1/82 (1%)
Frame = +3
Query: 235 QMLWMKHQLEDYQILESN-ISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGV 293
+ LW+ + L D +I + IYCDN +A+ L+ N + H R ++IE+ H + V G+
Sbjct: 21 EALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFGI 200
Query: 294 LLLKFVDTDHQWADIFSKPLAE 315
L L V + Q AD+F+K +++
Sbjct: 201 LHLLHVPSSDQVADVFTKTISQ 266
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 55.1 bits (131), Expect = 2e-08
Identities = 26/91 (28%), Positives = 50/91 (54%), Gaps = 1/91 (1%)
Frame = +1
Query: 238 WMKHQLEDYQI-LESNISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLL 296
W+ + L+D ++ S +YCD+ +A ++ N + H R KH+++ H +R+ +Q + L
Sbjct: 13 WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAKLFHL 192
Query: 297 KFVDTDHQWADIFSKPLAEDRFKFILKNLNM 327
+ + Q ADI +KPL F ++ L +
Sbjct: 193 LPISSVDQTADILTKPLESGPFSHLVSKLGV 285
>TC19389 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (6%)
Length = 498
Score = 52.0 bits (123), Expect = 1e-07
Identities = 34/114 (29%), Positives = 60/114 (51%), Gaps = 1/114 (0%)
Frame = +3
Query: 207 VSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNISIYCDNTAAI-SL 265
V+W S+ Q +ALSTAEAE+I+A ++LWMK+ L++ I T A+ +L
Sbjct: 27 VAWPSRLQKCVALSTAEAEFIAATEACHELLWMKNFLQNAWFHSHPILCCIVITKALFTL 206
Query: 266 SKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSKPLAEDRFK 319
++ L+ + +RD + +L L+ + TD AD+ +K L ++ +
Sbjct: 207 ARILLFIQDPSTLMFVIIGLRDVLNSKLLELEKIHTDDDGADMMTKSLPREKLE 368
>TC19474 weakly similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement pol
polyprotein, partial (3%)
Length = 517
Score = 37.4 bits (85), Expect(2) = 3e-06
Identities = 20/54 (37%), Positives = 33/54 (61%)
Frame = -2
Query: 258 DNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSK 311
DN AA+ + K + RAKHIE+ +HF+ D + +FV+++ Q D+F+K
Sbjct: 354 DNKAALHMPK-I*FSMRAKHIEIDFHFL*DRRLYLDISTRFVNSNDQLTDVFTK 196
Score = 29.6 bits (65), Expect(2) = 3e-06
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = -1
Query: 202 LGSNLVSWASKRQSTIALSTAEAEY 226
L NL+SW SK +A S AEAE+
Sbjct: 517 LSDNLISWKSKETIIVARSRAEAEF 443
>TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partial (20%)
Length = 739
Score = 46.6 bits (109), Expect = 6e-06
Identities = 32/99 (32%), Positives = 49/99 (49%), Gaps = 1/99 (1%)
Frame = -1
Query: 70 QSKYTKEFLKKFNMTECTPAKTPMHPTCI-LEKEDVSAKVCQKLYRGMIGSLLYLTASRP 128
Q Y + LKKF MT T + I + + +V Y+ +IGS+ YL R
Sbjct: 739 QKIYVDDILKKFKMTNSKYISTTIGGKEIEAGRRNGGKRVDSTYYKSLIGSVRYLNTVRS 560
Query: 129 DILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGL 167
DI+ V L ++F +P + H +R L+Y+KGT G+
Sbjct: 559 DIVCGVGLRSRFM-EP*DCH*QGAQRSLRYIKGTLKDGI 446
>AV424544
Length = 276
Score = 37.4 bits (85), Expect = 0.004
Identities = 17/57 (29%), Positives = 30/57 (51%)
Frame = +3
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKY 73
+YVDD+I + + + +F + +G LKYFLG++V + ++ Q KY
Sbjct: 105 VYVDDLILAGNDLNEIQCVKNKLDIQFRIKDLGTLKYFLGLEVARSSCGLFLSQRKY 275
>AV777635
Length = 382
Score = 35.4 bits (80), Expect = 0.014
Identities = 18/37 (48%), Positives = 24/37 (64%)
Frame = -2
Query: 138 AQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS* 174
+QF DP E HL RIL+YLK + GL+++KS *
Sbjct: 381 SQFMHDPHERHLD---RILQYLKASPGRGLLFRKSE* 280
>BP084259
Length = 385
Score = 33.1 bits (74), Expect = 0.067
Identities = 15/53 (28%), Positives = 28/53 (52%), Gaps = 1/53 (1%)
Frame = -3
Query: 233 STQMLWMKHQLEDYQILESNIS-IYCDNTAAISLSKNLILHSRAKHIEVKYHF 284
+ + LW++ L++ Q + I+ +N A + NL+ HS KH+ + HF
Sbjct: 161 AAEFLWLQQLLKELQFSLPKVPCIFSNNINATYVCLNLVFHSHMKHVALDCHF 3
>AV408509
Length = 428
Score = 32.3 bits (72), Expect = 0.11
Identities = 31/110 (28%), Positives = 48/110 (43%), Gaps = 5/110 (4%)
Frame = -3
Query: 19 VDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQV-----DQQPEATYIHQSKY 73
V D I+ N L +E K +FE +G +KYF G QV + E + +++
Sbjct: 324 VIDFIY-MTNDELLEE*KKTTMCQFERLDLGLMKYFDGHQVMAYK*IKNMEKCFFYKASM 148
Query: 74 TKEFLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYL 123
+ + + F + K L +ED KV LYR + SL+YL
Sbjct: 147 LQIYYRSFALCRMGVNKN-------LLREDGKEKVDDTLYRKLGQSLIYL 19
>BP052124
Length = 467
Score = 30.8 bits (68), Expect = 0.33
Identities = 13/39 (33%), Positives = 24/39 (61%)
Frame = -2
Query: 256 YCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVL 294
+CDN +A++L+ I HSR H EV + ++ + G++
Sbjct: 454 FCDNNSALTLAPRPI*HSRPVHFEVDCPYPKEKLGTGLI 338
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.137 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,778,258
Number of Sequences: 28460
Number of extensions: 75886
Number of successful extensions: 494
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 489
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 490
length of query: 334
length of database: 4,897,600
effective HSP length: 91
effective length of query: 243
effective length of database: 2,307,740
effective search space: 560780820
effective search space used: 560780820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0022.11


