

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0020.6
(795 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
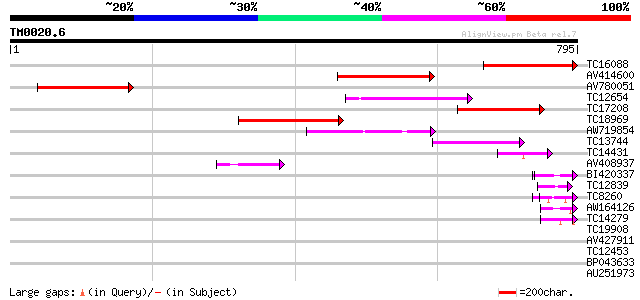
Score E
Sequences producing significant alignments: (bits) Value
TC16088 similar to UP|Q8PX92 (Q8PX92) Chemotaxis protein CheW, p... 286 7e-78
AV414600 186 2e-47
AV780051 140 8e-34
TC12654 similar to UP|Q94II2 (Q94II2) ERD4 protein (Fragment), p... 133 1e-31
TC17208 similar to UP|Q9MAV5 (Q9MAV5) F24O1.4, partial (13%) 122 2e-28
TC18969 similar to UP|Q94II2 (Q94II2) ERD4 protein (Fragment), p... 110 7e-25
AW719854 108 3e-24
TC13744 weakly similar to UP|Q39126 (Q39126) HYP1 protein (T4P13... 97 1e-20
TC14431 UP|BAC29902 (BAC29902) 16 days neonate thymus cDNA, RIKE... 59 2e-09
AV408937 55 3e-08
BI420337 45 3e-05
TC12839 similar to GB|BAB01914.1|9294011|AP001307 casein kinase-... 44 8e-05
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 44 1e-04
AW164126 42 4e-04
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 41 6e-04
TC19908 similar to GB|CAA60022.1|791150|VURNEXT26 extensin-like ... 40 0.001
AV427911 39 0.002
TC12453 weakly similar to GB|BAA88270.1|6573289|AB008023 RXW8 {A... 39 0.002
BP043633 38 0.007
AU251973 37 0.009
>TC16088 similar to UP|Q8PX92 (Q8PX92) Chemotaxis protein CheW, partial (8%)
Length = 769
Score = 286 bits (733), Expect = 7e-78
Identities = 131/131 (100%), Positives = 131/131 (100%)
Frame = +2
Query: 665 AFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFEVEEEL 724
AFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFEVEEEL
Sbjct: 2 AFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFEVEEEL 181
Query: 725 VEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP 784
VEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP
Sbjct: 182 VEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP 361
Query: 785 PHYAYQYQDEP 795
PHYAYQYQDEP
Sbjct: 362 PHYAYQYQDEP 394
>AV414600
Length = 414
Score = 186 bits (471), Expect = 2e-47
Identities = 84/136 (61%), Positives = 112/136 (81%)
Frame = +2
Query: 460 RKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIM 519
R++A++YY F VN+FLG+I+ G+AF+QL +F+H+S + P IG +IP+KA+FF+TYIM
Sbjct: 5 RRSASRYYLFNFVNIFLGNILAGSAFQQLDTFIHESANRYPIIIGSAIPIKASFFITYIM 184
Query: 520 VDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGI 579
VDGWAGI E+L LKPL+IYHLKN F+VKTE+DR +AMDPGS+ F P +QLYFLLG+
Sbjct: 185 VDGWAGIDAEVLMLKPLIIYHLKNFFLVKTEKDREEAMDPGSIGFNTGEPRIQLYFLLGL 364
Query: 580 VYAVVTPILLPFILVF 595
VYA VTP +LPFI++F
Sbjct: 365 VYAAVTPTVLPFIIIF 412
>AV780051
Length = 473
Score = 140 bits (353), Expect = 8e-34
Identities = 67/136 (49%), Positives = 93/136 (68%), Gaps = 1/136 (0%)
Frame = +3
Query: 39 FPKWYINGERTSPRSTGNFVGKFVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVF 98
F +WY+ G RT P G ++ KFVNL++++Y+ F NWMP ALRM E E+I+HAG DS V+
Sbjct: 66 FQEWYLKGLRTDPVHGGAYMRKFVNLDWRSYIIFWNWMPAALRMPEPELIDHAGSDSVVY 245
Query: 99 LRIYTLGLKIFVPVTVVALLILIPVN-VSSGALFFLKRDLVVSDIDKLSISNVPSKSSRF 157
LRIY +GLKIFVP+ +A +L+PVN S+G +++ SDID +S+SNV S R+
Sbjct: 246 LRIYLIGLKIFVPIAFLAWAVLVPVNWTSTGLEGTGLKNITASDIDTISVSNVQRGSQRY 425
Query: 158 FVHIGLEYMFTLWICF 173
HI + Y FT W C+
Sbjct: 426 RAHIVVAYAFTFWTCY 473
Score = 37.0 bits (84), Expect = 0.012
Identities = 18/53 (33%), Positives = 26/53 (48%)
Frame = +2
Query: 21 FLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVNLNFKTYLTFL 73
F A+LR+QP NDRVYFP+ G + P + + + L + FL
Sbjct: 11 FSCGVAMLRLQPFNDRVYFPRMVPQGFKNRPCARRSVYAQVCQLRLEVIYHFL 169
>TC12654 similar to UP|Q94II2 (Q94II2) ERD4 protein (Fragment), partial
(24%)
Length = 556
Score = 133 bits (334), Expect = 1e-31
Identities = 59/177 (33%), Positives = 106/177 (59%)
Frame = +2
Query: 472 VNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEIL 531
+NVF+G + GT F + + + P ++ + S+P ATFF+TY+ + + G E+
Sbjct: 5 LNVFIGVTIGGTLFNTFKT-VEEHPNKLVSMLAASLPGNATFFLTYVALKFFVGYGLELS 181
Query: 532 RLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPF 591
R+ PL+IYHLK F+ K E + A PG + + +P L + + Y+++ P+++PF
Sbjct: 182 RIVPLIIYHLKRKFVCKNEAELKAAWAPGDLSYGTRVPGDMLIVTIVLCYSIIAPLIIPF 361
Query: 592 ILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKK 648
+++F L +L+ R+Q + VY YES WPH+++RI+A+L++ Q+ MLG ++
Sbjct: 362 GVLYFGLGWLILRNQALKVYVPSYESYGRMWPHINNRILAALILYQITMLGYFGVQQ 532
>TC17208 similar to UP|Q9MAV5 (Q9MAV5) F24O1.4, partial (13%)
Length = 682
Score = 122 bits (306), Expect = 2e-28
Identities = 62/123 (50%), Positives = 86/123 (69%)
Frame = +1
Query: 628 RIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAM 687
RII +L+ISQLL++GLLSTK+AA STPLL ILP+LT FH +C+ R+EPAF ++PL+EAM
Sbjct: 1 RIIFALVISQLLLMGLLSTKEAANSTPLLIILPVLTIWFHLFCKGRYEPAFVQHPLQEAM 180
Query: 688 AKDLLEKTTEPDLNIKAYLADAYLHPIFRSFEVEEELVEVRVDKQPETQVASPTLSEPSS 747
KD LE+ EP LN K +L +AY+HP+F+S E + V + + V + S ++
Sbjct: 181 MKDTLERAREPQLNYKEFLQNAYIHPVFKSDEDSDSDVMSQEFEDEPMLVQTKRQSRKNT 360
Query: 748 PSP 750
P P
Sbjct: 361 PLP 369
>TC18969 similar to UP|Q94II2 (Q94II2) ERD4 protein (Fragment), partial
(23%)
Length = 456
Score = 110 bits (276), Expect = 7e-25
Identities = 52/147 (35%), Positives = 90/147 (60%)
Frame = +2
Query: 321 AFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSV 380
A + FS+R A+ AQ+ ++ W APEPR + W NL I + +R+ V+ + V
Sbjct: 14 ALVFFSNRIAAASAAQSLHAQMVDTWSVFDAPEPRQLLWPNLKIKYFERELRQYVVYVIV 193
Query: 381 FALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILP 440
++ FYMIPI F+ ++ L+ L ++ PF++P++ + +++ L+ +LP LAL IFL +LP
Sbjct: 194 ALMILFYMIPITFISAVTTLKNLVKILPFIKPIVRITVLRTVLEAYLPQLALIIFLAMLP 373
Query: 441 AVLMIMSKIEGHIAKSTLERKTAAKYY 467
+L+ +SK+EG +S R + KY+
Sbjct: 374 KLLLFLSKLEGIPTESHAVRAASGKYF 454
>AW719854
Length = 532
Score = 108 bits (271), Expect = 3e-24
Identities = 52/180 (28%), Positives = 101/180 (55%)
Frame = +3
Query: 417 KFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFL 476
KF+ + G+LP L L++ L ++P ++ ++S I+G+I+ S +E K +F + NVF
Sbjct: 12 KFVTQVITGYLPSLILQMSLKVVPPIMRLLSTIQGYISHSNIEMSACRKVLWFTVWNVFF 191
Query: 477 GSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPL 536
++ +G+ + L L + + V +P +A+FF+TY++ GW I E+ ++ PL
Sbjct: 192 ATVFSGSVIDHLSVVLDLK--DLAGKLAVGVPAQASFFITYVVTIGWTSILSELFQIIPL 365
Query: 537 VIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFF 596
++ +K F + + + S+ + +P + + LLGI Y + P++LPF+LV+F
Sbjct: 366 ILSLIKRPFTRQEDE-----FEAPSMAYHRDVPRVLFFGLLGITYFFLAPLILPFLLVYF 530
>TC13744 weakly similar to UP|Q39126 (Q39126) HYP1 protein (T4P13.21
protein), partial (26%)
Length = 744
Score = 97.1 bits (240), Expect = 1e-20
Identities = 47/130 (36%), Positives = 71/130 (54%)
Frame = +2
Query: 593 LVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKS 652
L++F L Y+++R+Q++ VY +YE+ FWP VH I SL++ ++ +GL KK +
Sbjct: 2 LIYFCLGYIIFRNQLLKVYVPRYETGGQFWPTVHHSTIFSLILMHVIAIGLFGLKKLPLA 181
Query: 653 TPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLH 712
+ L LPILT F++YCQ RF P F+ YP E + KD + +ADAY
Sbjct: 182 SLLTLPLPILTLLFNEYCQTRFLPIFKDYPAECLIKKDRADPNDHNMSEFYDKMADAYND 361
Query: 713 PIFRSFEVEE 722
P + E
Sbjct: 362 PALNPIQYSE 391
>TC14431 UP|BAC29902 (BAC29902) 16 days neonate thymus cDNA, RIKEN
full-length enriched library, clone:A130064K11
product:high mobility group box 1, full insert sequence,
partial (5%)
Length = 804
Score = 59.3 bits (142), Expect = 2e-09
Identities = 33/80 (41%), Positives = 45/80 (56%), Gaps = 3/80 (3%)
Frame = +1
Query: 685 EAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSF---EVEEELVEVRVDKQPETQVASPT 741
EAM KD LE+ TEP+LN+K YL +AY+HP+F++ E +EE +V K V PT
Sbjct: 1 EAMMKDTLERATEPNLNLKGYLQNAYVHPVFKASMEDEEDEEEDDVMSHKWDSESVTVPT 180
Query: 742 LSEPSSPSPLHDHVHQPSPP 761
+ +PL V S P
Sbjct: 181 KRQSRRNTPLPSKVSGASSP 240
>AV408937
Length = 324
Score = 55.5 bits (132), Expect = 3e-08
Identities = 34/98 (34%), Positives = 51/98 (51%), Gaps = 3/98 (3%)
Frame = +3
Query: 291 YYSHT-IKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTD 349
Y SH I+ K+ +L R AF+ F +R+ A+ QQS +PT W+T+
Sbjct: 57 YLSHVVIRRTSKIGSLLEAR----------AAFVFFKARYAAATALHLQQSVDPTQWVTE 206
Query: 350 WAPEPRDIYWRNLAIPFVSLSIRKLVISL--SVFALVF 385
APEP D+YW + F+ I KLV+ + VF ++F
Sbjct: 207 LAPEPHDVYWPFFSKSFMRRWISKLVVVVLCIVFTIMF 320
Score = 34.7 bits (78), Expect = 0.060
Identities = 13/37 (35%), Positives = 23/37 (62%)
Frame = +3
Query: 214 SVSDSVESFFQTNHPNHYIGHQAVYNANKFAKLVRRR 250
S+SDS++SFF+ +P+ Y+ H + +K L+ R
Sbjct: 6 SISDSIDSFFKELYPSTYLSHVVIRRTSKIGSLLEAR 116
>BI420337
Length = 397
Score = 45.4 bits (106), Expect = 3e-05
Identities = 24/64 (37%), Positives = 26/64 (40%), Gaps = 1/64 (1%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYH-PPPPPHYAYQY 791
P P S P P PSPP V +Y P P YYY PPPPP Y+Y
Sbjct: 145 PPMHSPPPPYHYSSPPPPPKKPYKYPSPPPPVYKYKSPPPRTPAYYYKSPPPPPKNPYKY 324
Query: 792 QDEP 795
P
Sbjct: 325 PSPP 336
Score = 41.2 bits (95), Expect = 6e-04
Identities = 20/60 (33%), Positives = 26/60 (43%)
Frame = +1
Query: 736 QVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
Q+++ S P P + H P PP H PP +Y PPPPP Y+Y P
Sbjct: 73 QISANNYIYTSPPPPKPYYYHSPPPPMH-------SPPPPYHYSSPPPPPKKPYKYPSPP 231
Score = 34.3 bits (77), Expect = 0.079
Identities = 16/50 (32%), Positives = 22/50 (44%)
Frame = +1
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAY 789
P + + SP P + SPP YP+ PP Y + PPP + Y
Sbjct: 232 PPVYKYKSPPPRTPAYYYKSPPPPPKNPYKYPSPPPPVYKYKSPPPPFKY 381
Score = 32.7 bits (73), Expect = 0.23
Identities = 16/42 (38%), Positives = 22/42 (52%), Gaps = 7/42 (16%)
Frame = +1
Query: 761 PQHVNQYSHYPTSPPG---YYYHPPPPPHYA----YQYQDEP 795
P ++ ++ TSPP YYYH PPPP ++ Y Y P
Sbjct: 67 PIQISANNYIYTSPPPPKPYYYHSPPPPMHSPPPPYHYSSPP 192
>TC12839 similar to GB|BAB01914.1|9294011|AP001307 casein kinase-like
protein {Arabidopsis thaliana;} , partial (5%)
Length = 806
Score = 44.3 bits (103), Expect = 8e-05
Identities = 22/49 (44%), Positives = 25/49 (50%)
Frame = -2
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
P++S P PLH PS Q + SH P P YYY PPPPH A
Sbjct: 592 PSVSPQPLPYPLHPDSLLPSSAQQLR--SHRPQPPAAYYYSSPPPPHAA 452
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 43.5 bits (101), Expect = 1e-04
Identities = 23/63 (36%), Positives = 28/63 (43%)
Frame = +3
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQ 792
P + P P P P H PSPP ++Y H P P Y+ PPPPP Y+Y
Sbjct: 228 PPYEKPHPVYHSPPPPPP-HKPYKYPSPPPPPHKYPH-PHPHPVYHSPPPPPPKKHYKYS 401
Query: 793 DEP 795
P
Sbjct: 402 SPP 410
Score = 41.2 bits (95), Expect = 6e-04
Identities = 25/65 (38%), Positives = 29/65 (44%), Gaps = 12/65 (18%)
Frame = +3
Query: 743 SEPSSPSPLHDH-----VHQPSPPQHVNQYSHYPTSPPGY-----YYH--PPPPPHYAYQ 790
S P P P H VH P PP + + Y + PP Y YH PPPPPH Y+
Sbjct: 120 SPPPPPKPYSYHSPPPPVHSPPPP-YEKPHPVYHSPPPPYEKPHPVYHSPPPPPPHKPYK 296
Query: 791 YQDEP 795
Y P
Sbjct: 297 YPSPP 311
Score = 39.3 bits (90), Expect = 0.002
Identities = 23/70 (32%), Positives = 28/70 (39%), Gaps = 20/70 (28%)
Frame = +3
Query: 746 SSPSPLHDHVHQ-------PSPPQHVNQYSHY-PTSPPGYYY------------HPPPPP 785
S P P+H + H P PP H Y + P PP + Y H PPPP
Sbjct: 453 SPPPPVHTYPHPHPVYHSPPPPPPHEKPYKYASPPPPPVHTYPHPVYHSPPPPVHSPPPP 632
Query: 786 HYAYQYQDEP 795
HY Y+ P
Sbjct: 633 HYYYKSPPPP 662
Score = 36.6 bits (83), Expect = 0.016
Identities = 17/44 (38%), Positives = 21/44 (47%), Gaps = 4/44 (9%)
Frame = +3
Query: 748 PSPLHDH----VHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P P+H + H P PP H PP YYY PPPP++
Sbjct: 558 PPPVHTYPHPVYHSPPPPVH-------SPPPPHYYYKSPPPPYH 668
Score = 35.0 bits (79), Expect = 0.046
Identities = 19/62 (30%), Positives = 24/62 (38%), Gaps = 15/62 (24%)
Frame = +3
Query: 745 PSSPSPLHDH--------VHQPSPPQHVNQYSHYPTSPPGY-------YYHPPPPPHYAY 789
PS P P H + H P PP Y + PP + YH PPPP + Y
Sbjct: 300 PSPPPPPHKYPHPHPHPVYHSPPPPPPKKHYKYSSPPPPVHTYPHPHPVYHSPPPPVHTY 479
Query: 790 QY 791
+
Sbjct: 480 PH 485
Score = 34.7 bits (78), Expect = 0.060
Identities = 20/64 (31%), Positives = 24/64 (37%), Gaps = 1/64 (1%)
Frame = +3
Query: 733 PETQVASPTLSEPSSP-SPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQY 791
P + P P P H H P PP Y YP+ PP + +P P PH Y
Sbjct: 186 PPYEKPHPVYHSPPPPYEKPHPVYHSPPPPPPHKPYK-YPSPPPPPHKYPHPHPHPVYHS 362
Query: 792 QDEP 795
P
Sbjct: 363 PPPP 374
Score = 32.7 bits (73), Expect = 0.23
Identities = 21/62 (33%), Positives = 23/62 (36%), Gaps = 6/62 (9%)
Frame = +3
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP------PHYAYQYQD 793
P P P P H SPP V+ Y H P Y+ PPPP PH Y
Sbjct: 348 PVYHSPPPPPP-KKHYKYSSPPPPVHTYPH----PHPVYHSPPPPVHTYPHPHPVYHSPP 512
Query: 794 EP 795
P
Sbjct: 513 PP 518
Score = 28.9 bits (63), Expect = 3.3
Identities = 11/19 (57%), Positives = 11/19 (57%)
Frame = +3
Query: 767 YSHYPTSPPGYYYHPPPPP 785
YS P P Y YH PPPP
Sbjct: 114 YSSPPPPPKPYSYHSPPPP 170
>AW164126
Length = 387
Score = 42.0 bits (97), Expect = 4e-04
Identities = 22/55 (40%), Positives = 25/55 (45%), Gaps = 4/55 (7%)
Frame = +2
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP----PHYAYQYQDEP 795
P SPSP + P PP P+ PP YYY PPP PH Y Y+ P
Sbjct: 149 PPSPSPKPYYYKSPPPPS--------PSPPPPYYYKSPPPPSPVPHPPYYYKSPP 289
Score = 36.6 bits (83), Expect = 0.016
Identities = 23/60 (38%), Positives = 24/60 (39%), Gaps = 3/60 (5%)
Frame = +2
Query: 739 SPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP---PHYAYQYQDEP 795
SP PS P P + P PP V PP YY PPPP PH Y Y P
Sbjct: 185 SPPPPSPSPPPPYY--YKSPPPPSPV-------PHPPYYYKSPPPPSPIPHPPYYYSSPP 337
Score = 33.9 bits (76), Expect = 0.10
Identities = 17/46 (36%), Positives = 23/46 (49%), Gaps = 1/46 (2%)
Frame = +2
Query: 745 PSSPSPLHDHVHQ-PSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAY 789
P SP P + ++ P PP + +Y +SPP PPPP Y Y
Sbjct: 242 PPSPVPHPPYYYKSPPPPSPIPHPPYYYSSPPPPVKSPPPPTPYYY 379
Score = 33.5 bits (75), Expect = 0.13
Identities = 14/29 (48%), Positives = 16/29 (54%), Gaps = 4/29 (13%)
Frame = +3
Query: 771 PTSPPGYYYHPPPP----PHYAYQYQDEP 795
P+ PP YYY PPP PH Y Y+ P
Sbjct: 18 PSPPPPYYYKSPPPPSPVPHPPYYYKSPP 104
Score = 31.6 bits (70), Expect = 0.51
Identities = 16/31 (51%), Positives = 16/31 (51%), Gaps = 3/31 (9%)
Frame = +3
Query: 758 PSPPQHVNQYSHYPTSP---PGYYYHPPPPP 785
PSPP S P SP P YYY PPPP
Sbjct: 18 PSPPPPYYYKSPPPPSPVPHPPYYYKSPPPP 110
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 41.2 bits (95), Expect = 6e-04
Identities = 24/64 (37%), Positives = 30/64 (46%), Gaps = 13/64 (20%)
Frame = +1
Query: 745 PSSPSPLHDHVHQ-PSPPQHVNQYSHY--------PTSPPGYYYHPPPPPHYA----YQY 791
P SPSP + +Q P PP H +Y P+ PP YYY PPPP + Y Y
Sbjct: 103 PPSPSPPPPYYYQSPPPPSHSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYY 282
Query: 792 QDEP 795
+ P
Sbjct: 283 KSPP 294
Score = 40.4 bits (93), Expect = 0.001
Identities = 21/55 (38%), Positives = 26/55 (47%), Gaps = 4/55 (7%)
Frame = +1
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP----PHYAYQYQDEP 795
P SPSP + ++ PP P+ PP YYY PPP PH Y Y+ P
Sbjct: 199 PPSPSPPPPYYYKSPPPPS-------PSPPPPYYYKSPPPPSPIPHPPYYYKSPP 342
Score = 37.7 bits (86), Expect = 0.007
Identities = 20/59 (33%), Positives = 26/59 (43%), Gaps = 8/59 (13%)
Frame = +1
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSH----YPTSPPGYYYHPPPPPHYA----YQYQDEP 795
P P P+ + SPP Y P+ PP YYY PPPP ++ Y Y+ P
Sbjct: 22 PPPPPPVPKPYYYKSPPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPPPPYYYKSPP 198
Score = 37.0 bits (84), Expect = 0.012
Identities = 20/55 (36%), Positives = 23/55 (41%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P SP PS P P + P PP + +Y SPP PPPP HY
Sbjct: 220 PPYYYKSPPPPSPSPPPPYY--YKSPPPPSPIPHPPYYYKSPPPPTSSPPPPYHY 378
Score = 34.7 bits (78), Expect = 0.060
Identities = 20/55 (36%), Positives = 22/55 (39%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P SP PS P P H P PP ++ SPP PPPP HY
Sbjct: 364 PPYHYVSPPPPSPSPPPPYH--YASPPPPSPSPAPTYIYKSPPPPVKLPPPPYHY 522
Score = 34.7 bits (78), Expect = 0.060
Identities = 19/54 (35%), Positives = 24/54 (44%), Gaps = 1/54 (1%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQH-VNQYSHYPTSPPGYYYHPPPPP 785
P + + P S P P P PP H V+ P+ PP Y+Y PPPP
Sbjct: 295 PPSPIPHPPYYYKSPPPP----TSSPPPPYHYVSPPPPSPSPPPPYHYASPPPP 444
Score = 33.9 bits (76), Expect = 0.10
Identities = 19/50 (38%), Positives = 23/50 (46%), Gaps = 3/50 (6%)
Frame = +1
Query: 741 TLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTS---PPGYYYHPPPPPHY 787
T P SPSP ++++ PP PT PP Y Y PPPP Y
Sbjct: 523 TSPPPPSPSPAPTYIYKSPPP---------PTKSPPPPVYIYASPPPPIY 645
Score = 33.9 bits (76), Expect = 0.10
Identities = 22/62 (35%), Positives = 29/62 (46%), Gaps = 9/62 (14%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQ-PSPPQHVNQYSHYPTSPPG--------YYYHPPP 783
P ASP P SPSP ++++ P PP + ++ TSPP Y Y PP
Sbjct: 412 PPYHYASPP---PPSPSPAPTYIYKSPPPPVKLPPPPYHYTSPPPPSPSPAPTYIYKSPP 582
Query: 784 PP 785
PP
Sbjct: 583 PP 588
Score = 30.0 bits (66), Expect = 1.5
Identities = 20/69 (28%), Positives = 22/69 (30%), Gaps = 12/69 (17%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPP---------- 782
P SP PS P P + P P Y + PP HPP
Sbjct: 172 PPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHPPYYYKSPPPPT 351
Query: 783 --PPPHYAY 789
PPP Y Y
Sbjct: 352 SSPPPPYHY 378
Score = 28.1 bits (61), Expect = 5.7
Identities = 19/73 (26%), Positives = 25/73 (34%), Gaps = 10/73 (13%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGY------YYHPPPPPH 786
P V P + P P + P P Y + PP + YY PPPP
Sbjct: 28 PPPPVPKPYYYKSPPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPPPPYYYKSPPPPS 207
Query: 787 YA----YQYQDEP 795
+ Y Y+ P
Sbjct: 208 PSPPPPYYYKSPP 246
>TC19908 similar to GB|CAA60022.1|791150|VURNEXT26 extensin-like protein
{Vigna unguiculata;} , partial (49%)
Length = 472
Score = 40.4 bits (93), Expect = 0.001
Identities = 22/56 (39%), Positives = 28/56 (49%), Gaps = 5/56 (8%)
Frame = +2
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPP-----PHYAYQYQDEP 795
P SPSP ++++ PP SH P PP Y Y PPP PH++Y Y P
Sbjct: 86 PPSPSPPPPYIYKSPPPP-----SHSP--PPPYVYKSPPPPSSSHPHHSYLYNSPP 232
Score = 34.3 bits (77), Expect = 0.079
Identities = 21/58 (36%), Positives = 26/58 (44%), Gaps = 7/58 (12%)
Frame = +2
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYY-------HPPPPPHYAYQYQDEP 795
P SPSP +V++ PP P+ PP Y Y H PPPP Y Y+ P
Sbjct: 38 PPSPSPPPPYVYKSPPPPS-------PSPPPPYIYKSPPPPSHSPPPP---YVYKSPP 181
>AV427911
Length = 348
Score = 39.3 bits (90), Expect = 0.002
Identities = 19/43 (44%), Positives = 20/43 (46%), Gaps = 1/43 (2%)
Frame = +1
Query: 754 HVHQPSPPQHVNQYSHYPTSPPGYYYH-PPPPPHYAYQYQDEP 795
H H P PP Y PP YYY PPPPP Y+Y P
Sbjct: 133 HYHSPPPPV-------YSPPPPKYYYKSPPPPPKKPYKYPSPP 240
Score = 36.6 bits (83), Expect = 0.016
Identities = 21/67 (31%), Positives = 27/67 (39%), Gaps = 4/67 (5%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTS----PPGYYYHPPPPPHYA 788
P SP S P P + + P PP+ +Y P P Y+ PPPPP
Sbjct: 124 PPYHYHSPPPPVYSPPPPKYYYKSPPPPPKKPYKYPSPPPPVHVYPKPTYHSPPPPPKKP 303
Query: 789 YQYQDEP 795
Y+Y P
Sbjct: 304 YKYPSPP 324
Score = 35.4 bits (80), Expect = 0.035
Identities = 22/64 (34%), Positives = 29/64 (44%), Gaps = 5/64 (7%)
Frame = +1
Query: 737 VASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA-----YQY 791
+A +L+ PS S P PP+ YP P Y+YH PPPP Y+ Y Y
Sbjct: 28 LAIVSLTLPSQISADGYIYSSPPPPKE------YPPVSPPYHYHSPPPPVYSPPPPKYYY 189
Query: 792 QDEP 795
+ P
Sbjct: 190 KSPP 201
Score = 33.5 bits (75), Expect = 0.13
Identities = 20/50 (40%), Positives = 22/50 (44%), Gaps = 6/50 (12%)
Frame = +1
Query: 746 SSPSPLHDHVHQPSPPQHVNQYS----HYPTSPP--GYYYHPPPPPHYAY 789
S P P PSPP V+ Y H P PP Y Y PPPP + Y
Sbjct: 193 SPPPPPKKPYKYPSPPPPVHVYPKPTYHSPPPPPKKPYKYPSPPPPVHVY 342
Score = 31.6 bits (70), Expect = 0.51
Identities = 18/47 (38%), Positives = 22/47 (46%), Gaps = 5/47 (10%)
Frame = +1
Query: 745 PSSPSPLHDH-----VHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPH 786
P SP P H H V+ P PP++ + P P Y PPPP H
Sbjct: 115 PVSP-PYHYHSPPPPVYSPPPPKYYYKSPPPPPKKPYKYPSPPPPVH 252
Score = 27.3 bits (59), Expect = 9.6
Identities = 14/41 (34%), Positives = 20/41 (48%), Gaps = 4/41 (9%)
Frame = +1
Query: 745 PSSPSPLHDH----VHQPSPPQHVNQYSHYPTSPPGYYYHP 781
PS P P+H + H P PP + YP+ PP + +P
Sbjct: 229 PSPPPPVHVYPKPTYHSPPPPP--KKPYKYPSPPPPVHVYP 345
>TC12453 weakly similar to GB|BAA88270.1|6573289|AB008023 RXW8 {Arabidopsis
thaliana;} , partial (10%)
Length = 528
Score = 39.3 bits (90), Expect = 0.002
Identities = 20/56 (35%), Positives = 31/56 (54%)
Frame = +3
Query: 165 YMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVE 220
Y+ TL C LLY EY +I +RL + F+++VRS+P S S ++V+
Sbjct: 144 YIITLAACTLLYFEYKSITNLRLLHIIGSPPNPSHFSILVRSIPWSSEESYCETVK 311
>BP043633
Length = 542
Score = 37.7 bits (86), Expect = 0.007
Identities = 29/111 (26%), Positives = 40/111 (35%), Gaps = 25/111 (22%)
Frame = +1
Query: 710 YLHPIFRSFEVEEELVEV-RVDKQPETQVAS------------PTLSEPSSPSPLHDHVH 756
+L+P SF +E + + + + ET + S P P P P H H
Sbjct: 121 FLNPSLPSFSIETQRISIITIQINSETPMDSYQQPHGYMRPPQPPPPPPPPPDPNHHHHF 300
Query: 757 QPSPPQHVNQYSHY------------PTSPPGYYYHPPPPPHYAYQYQDEP 795
PP H Q Y P+ PP + PPPPP + Y P
Sbjct: 301 PQIPPPHPPQAPWYSPQFQYHHPSQTPSPPPQWGPPPPPPPPPSNPYSYHP 453
Score = 28.9 bits (63), Expect = 3.3
Identities = 20/56 (35%), Positives = 25/56 (43%), Gaps = 2/56 (3%)
Frame = +1
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTS--PPGYYYHPPPPPH 786
P+ Q P+ + PS P P PP N YS++P PP PP PPH
Sbjct: 346 PQFQYHHPSQT-PSPPPQWGPPPPPPPPPS--NPYSYHPKQFPPPPPPPRPPHPPH 504
>AU251973
Length = 205
Score = 37.4 bits (85), Expect = 0.009
Identities = 19/56 (33%), Positives = 22/56 (38%)
Frame = +2
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
P S P P + + P PP Y + PP Y Y PPPP Y Y P
Sbjct: 50 PPYKYSSPPPPPYKYSSPPPPP-----YKYSSPPPPPYKYSSPPPPPYKYSSPPPP 202
Score = 33.9 bits (76), Expect = 0.10
Identities = 17/37 (45%), Positives = 18/37 (47%)
Frame = +2
Query: 759 SPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
SPP +YS P PP Y Y PPPP Y Y P
Sbjct: 8 SPPPPPYKYSSPP--PPPYKYSSPPPPPYKYSSPPPP 112
Score = 32.0 bits (71), Expect = 0.39
Identities = 18/46 (39%), Positives = 21/46 (45%)
Frame = +2
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPP 785
P S P P + + P PP +YS P PP Y PPPPP
Sbjct: 80 PPYKYSSPPPPPYKYSSPPPPPY---KYSS-PPPPPYKYSSPPPPP 205
Score = 31.6 bits (70), Expect = 0.51
Identities = 20/56 (35%), Positives = 24/56 (42%), Gaps = 6/56 (10%)
Frame = +2
Query: 746 SSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA------YQYQDEP 795
SSP P P PP + +YS P P Y PPPP Y+ Y+Y P
Sbjct: 5 SSPPPPPYKYSSPPPPPY--KYSSPPPPPYKYSSPPPPPYKYSSPPPPPYKYSSPP 166
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,836,842
Number of Sequences: 28460
Number of extensions: 258789
Number of successful extensions: 4126
Number of sequences better than 10.0: 246
Number of HSP's better than 10.0 without gapping: 3115
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3672
length of query: 795
length of database: 4,897,600
effective HSP length: 98
effective length of query: 697
effective length of database: 2,108,520
effective search space: 1469638440
effective search space used: 1469638440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0020.6


