

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.7
(1703 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
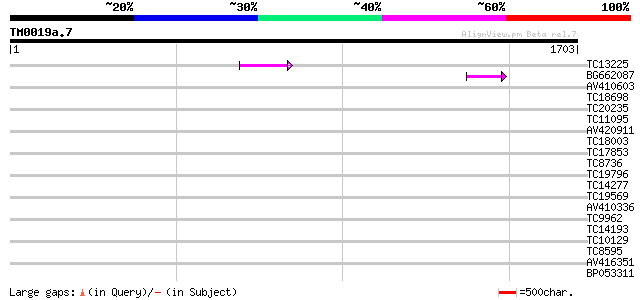
Score E
Sequences producing significant alignments: (bits) Value
TC13225 123 2e-28
BG662087 67 3e-11
AV410603 41 0.002
TC18698 40 0.004
TC20235 39 0.009
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 38 0.012
AV420911 37 0.026
TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 37 0.026
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 36 0.044
TC8736 similar to PIR|T06396|T06396 isoprenylated protein - soyb... 36 0.044
TC19796 35 0.076
TC14277 similar to UP|Q9SHY3 (Q9SHY3) F1E22.9 (AT1G65720/F1E22_1... 35 0.076
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 35 0.13
AV410336 34 0.17
TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial... 28 0.18
TC14193 similar to UP|Q8S2S7 (Q8S2S7) Ethylene responsive elemen... 34 0.22
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 34 0.22
TC8595 similar to PIR|T09390|T09390 21K protein precursor - alfa... 34 0.22
AV416351 34 0.22
BP053311 31 1.9
>TC13225
Length = 513
Score = 123 bits (309), Expect = 2e-28
Identities = 62/162 (38%), Positives = 95/162 (58%), Gaps = 2/162 (1%)
Frame = +3
Query: 689 VKTDEEGNPIMEDGKFIS--DAVNSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDF 746
+ T E+ + I+E K DAV +LI+TI HF+GDPS+ ++++ L+NL C ++ D+
Sbjct: 3 IMTPEDKSTILEKKKENGEEDAVATLIYTIILHFIGDPSIFRERASSQLANLYCPTMSDY 182
Query: 747 RWYKDTFLTRVYTREDSQQAFWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQ 806
RWYKDTF ++V RED AFWKE+F+AGLP+ KV + L N G + + +LS+GQ
Sbjct: 183 RWYKDTFFSKVTLREDGNSAFWKERFIAGLPRLMQSKVLDNLSLFNQGNPVNFGSLSFGQ 362
Query: 807 LIAIIQRVALKICQDDKIQQQLTKEKSQNRRDLGTFCEQFGI 848
L I ++ + +SQ R +FCE +G+
Sbjct: 363 LHNTIVHTGIQFVLTSNFKTNAKDMQSQKRS--SSFCEXYGV 482
>BG662087
Length = 373
Score = 66.6 bits (161), Expect = 3e-11
Identities = 38/119 (31%), Positives = 65/119 (53%), Gaps = 1/119 (0%)
Frame = +1
Query: 1373 GTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYK 1432
G R+ ++Y LN+A YP+P+ L+ D ++ S D SG+ QI++ D+ K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 1433 TAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIF-NPYSKLTIVYIDDVLIFSQTLDQH 1490
TAF Y + +PFGLKNA + +Q +M+ +F + + VY+D++++ S H
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>AV410603
Length = 162
Score = 40.8 bits (94), Expect = 0.002
Identities = 17/48 (35%), Positives = 30/48 (62%)
Frame = +1
Query: 1393 YPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFG 1440
+P+P +LL ++ FSK D++SG+ QI ++ +D++KT F G
Sbjct: 19 FPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>TC18698
Length = 808
Score = 39.7 bits (91), Expect = 0.004
Identities = 21/56 (37%), Positives = 34/56 (60%), Gaps = 1/56 (1%)
Frame = -2
Query: 1430 KYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYS-KLTIVYIDDVLIFS 1484
K KT + Y + VMP GLKN + +QR+M++IF+ K VY++D+++ S
Sbjct: 807 KKKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKS 640
>TC20235
Length = 543
Score = 38.5 bits (88), Expect = 0.009
Identities = 26/103 (25%), Positives = 51/103 (49%), Gaps = 2/103 (1%)
Frame = +1
Query: 61 CFPDLTVSLEDKNILDVLFLNIKLHGLDMKEDSI--PISLIYRVQYKVMNSIKSYFLRTN 118
C+P VSL D ++ L L++K+ +M D P+S++YR YKV + ++ + +
Sbjct: 1 CYPSYPVSLRDPHVTKCLDLDVKIPN-EMFVDGAPPPVSIMYRFCYKVTDYLRGVKAQRS 177
Query: 119 CEKSGETTLFLTDYDKANVIVPKTIQWSDITLPEKWSLEKATP 161
ET + T + + + ++ I PE+W +++P
Sbjct: 178 -NPVNETDVIETHIALYSSEITRILKHDRIAYPEEWLPGRSSP 303
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 38.1 bits (87), Expect = 0.012
Identities = 12/22 (54%), Positives = 17/22 (76%)
Frame = +1
Query: 903 GKDVTCYNCGKPGHISRYCRLK 924
G D+ CY CG+PGH +R CR++
Sbjct: 364 GSDMKCYECGEPGHFARECRMR 429
>AV420911
Length = 418
Score = 37.0 bits (84), Expect = 0.026
Identities = 12/20 (60%), Positives = 16/20 (80%)
Frame = +2
Query: 903 GKDVTCYNCGKPGHISRYCR 922
G+D+ CY CG+PGH +R CR
Sbjct: 359 GEDLKCYECGEPGHFARECR 418
>TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (42%)
Length = 587
Score = 37.0 bits (84), Expect = 0.026
Identities = 12/20 (60%), Positives = 16/20 (80%)
Frame = +1
Query: 903 GKDVTCYNCGKPGHISRYCR 922
G+D+ CY CG+PGH +R CR
Sbjct: 151 GEDLKCYECGEPGHFARECR 210
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 36.2 bits (82), Expect = 0.044
Identities = 20/68 (29%), Positives = 32/68 (46%)
Frame = +2
Query: 854 KPKPRKQDPPPKQQWRKRSSRNDHRKPKPRSKPQSSQIPKNPPETRPSQGKDVTCYNCGK 913
+P+ +D ++R+R D R + RS+ +R S G + C+ CGK
Sbjct: 287 QPQGSSRDRDDGDRYRERGRDRDDRGDRDRSRGYGG--------SRGSNGGE--CFKCGK 436
Query: 914 PGHISRYC 921
PGH +R C
Sbjct: 437 PGHFAREC 460
>TC8736 similar to PIR|T06396|T06396 isoprenylated protein - soybean
(fragment) {Glycine max;}, partial (84%)
Length = 1268
Score = 36.2 bits (82), Expect = 0.044
Identities = 15/54 (27%), Positives = 26/54 (47%)
Frame = +2
Query: 852 PKKPKPRKQDPPPKQQWRKRSSRNDHRKPKPRSKPQSSQIPKNPPETRPSQGKD 905
PK P+P K+ K + + + D KPK ++P+ + + P P + KD
Sbjct: 467 PKPPEPEKKKEADKPKAAEHEKKKDGEKPKAAAEPEKKKDSEKPKAAEPEKKKD 628
>TC19796
Length = 522
Score = 35.4 bits (80), Expect = 0.076
Identities = 18/60 (30%), Positives = 33/60 (55%), Gaps = 2/60 (3%)
Frame = -1
Query: 843 CEQFGIQGCPKKPKPRKQDPPPKQQWRKRS--SRNDHRKPKPRSKPQSSQIPKNPPETRP 900
C Q +G K+P+ + P P+++ KR ++ +++PK + KPQ + K ET+P
Sbjct: 225 CRQSH*RGEDKEPESLPKTPAPRKENEKRDIGTKPRNKRPKRQRKPQKNHTAKQEGETKP 46
>TC14277 similar to UP|Q9SHY3 (Q9SHY3) F1E22.9 (AT1G65720/F1E22_13), partial
(33%)
Length = 941
Score = 35.4 bits (80), Expect = 0.076
Identities = 16/44 (36%), Positives = 24/44 (54%), Gaps = 3/44 (6%)
Frame = +3
Query: 506 IPETPEPPET---PEPTPEAVSNPTITAPPPPKTRTSTRNTSTS 546
+P TP PP P P + +NPT ++ PKT T+ N +T+
Sbjct: 189 LPPTPSPPSPRSDPSPLSTSTTNPTTSSSTTPKTTTNKSNPNTA 320
Score = 33.5 bits (75), Expect = 0.29
Identities = 20/67 (29%), Positives = 25/67 (36%), Gaps = 3/67 (4%)
Frame = +3
Query: 486 ESDNELLQKVVEQLQLLKQVIPETPEPPETPEPTPEAVSNPTITAPP---PPKTRTSTRN 542
+SD LL + P P PP P + P+ T PP PP R+
Sbjct: 57 DSDPWLLSSAASRSPPSSSSSPSPPPPPHAPAAASSSPPTPSATLPPTPSPPSPRSDPSP 236
Query: 543 TSTSLNN 549
STS N
Sbjct: 237 LSTSTTN 257
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 34.7 bits (78), Expect = 0.13
Identities = 22/73 (30%), Positives = 29/73 (39%), Gaps = 4/73 (5%)
Frame = +1
Query: 509 TPEPPETPEPTPEAVSNPTITAPPPPKTRTSTRN----TSTSLNNIEDGKVESDSDIQSM 564
+P PP + +PT ++PT T PPP T N STS + E S
Sbjct: 70 SPPPPPSQQPTHSPSTSPTSTTTPPPSPTKVTENPPTAPSTSTKSPTSSASEEPSPPNPS 249
Query: 565 KSDVQSIKSIPVN 577
S Q K P +
Sbjct: 250 TSTTQQPKHSPTS 288
>AV410336
Length = 358
Score = 34.3 bits (77), Expect = 0.17
Identities = 20/46 (43%), Positives = 24/46 (51%), Gaps = 7/46 (15%)
Frame = +1
Query: 507 PETPEPPETPEPTPEAVSNPTITAPPP-------PKTRTSTRNTST 545
P P PP TP P + S+PT T+P P P T T T +TST
Sbjct: 67 PHAPPPP-TPNPKSPSSSSPTPTSPSPHSGRNSSPATTTFTTSTST 201
>TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial (21%)
Length = 585
Score = 27.7 bits (60), Expect(2) = 0.18
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = +3
Query: 518 PTPEAVSNPTITAPPPPK 535
PTP+ V++PT APP P+
Sbjct: 267 PTPKPVTHPTTPAPPQPQ 320
Score = 25.0 bits (53), Expect(2) = 0.18
Identities = 11/28 (39%), Positives = 11/28 (39%)
Frame = +1
Query: 493 QKVVEQLQLLKQVIPETPEPPETPEPTP 520
QK L L P PP TP P P
Sbjct: 73 QKTTSMLPLTSTTSTRNP*PPRTPPPPP 156
>TC14193 similar to UP|Q8S2S7 (Q8S2S7) Ethylene responsive element binding
factor 4-like protein, partial (50%)
Length = 1128
Score = 33.9 bits (76), Expect = 0.22
Identities = 18/44 (40%), Positives = 22/44 (49%), Gaps = 1/44 (2%)
Frame = +1
Query: 507 PETPEPPETPEPTPEAVSNPTITAP-PPPKTRTSTRNTSTSLNN 549
P P P TP P A P T+P PP+T T TS+SL +
Sbjct: 274 PRKPLEPTTPPPASSAALKPKPTSPNHPPRTTTPDPTTSSSLTS 405
Score = 28.5 bits (62), Expect = 9.3
Identities = 21/83 (25%), Positives = 34/83 (40%), Gaps = 2/83 (2%)
Frame = +1
Query: 503 KQVIPETPEPPETP--EPTPEAVSNPTITAPPPPKTRTSTRNTSTSLNNIEDGKVESDSD 560
K + P TP P + +P P + ++P T P P T +S + T
Sbjct: 280 KPLEPTTPPPASSAALKPKPTSPNHPPRTTTPDPTTSSSLTSRPTRAQARAAPWNPQPRS 459
Query: 561 IQSMKSDVQSIKSIPVNTVIHNS 583
+ S + V S + P + IHN+
Sbjct: 460 VTSRAAVVWSSTASPSSRCIHNN 528
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 33.9 bits (76), Expect = 0.22
Identities = 19/46 (41%), Positives = 22/46 (47%), Gaps = 12/46 (26%)
Frame = +1
Query: 507 PETPEPPETPEP------------TPEAVSNPTITAPPPPKTRTST 540
P TP PP+TP P TP +S P T P PPKT + T
Sbjct: 310 PYTPTPPKTPSPTSQPPHTPTPPNTPSPISQPPYT-PTPPKTPSPT 444
Score = 32.3 bits (72), Expect = 0.64
Identities = 19/45 (42%), Positives = 22/45 (48%), Gaps = 11/45 (24%)
Frame = +1
Query: 507 PETPEPPETPEPT--------PEAVSNPTI---TAPPPPKTRTST 540
P TP PP+TP PT P S+PT P PPKT + T
Sbjct: 406 PYTPTPPKTPSPTNQPPHIPSPPNSSSPTSQPPNTPSPPKTPSPT 540
Score = 31.2 bits (69), Expect = 1.4
Identities = 16/34 (47%), Positives = 18/34 (52%)
Frame = +1
Query: 507 PETPEPPETPEPTPEAVSNPTITAPPPPKTRTST 540
P P PP TP P +S P T P PPKT + T
Sbjct: 262 PHVPTPPSTPSP----ISQPPYT-PTPPKTPSPT 348
Score = 30.0 bits (66), Expect = 3.2
Identities = 17/46 (36%), Positives = 22/46 (46%), Gaps = 8/46 (17%)
Frame = +1
Query: 507 PETPEPPETPEPTPEAVSNPTITAPP--------PPKTRTSTRNTS 544
P TP PP+TP P + S P + +PP PP T T + S
Sbjct: 70 PHTPIPPKTPSP---SYSPPNVPSPPKAPSPNNHPPYTPTPPKTPS 198
Score = 29.6 bits (65), Expect = 4.2
Identities = 16/30 (53%), Positives = 18/30 (59%)
Frame = +1
Query: 507 PETPEPPETPEPTPEAVSNPTITAPPPPKT 536
P TP PP+TP PT + P I P PPKT
Sbjct: 166 PYTPTPPKTPSPTSQP---PYI--PLPPKT 240
>TC8595 similar to PIR|T09390|T09390 21K protein precursor - alfalfa
{Medicago sativa;}, partial (93%)
Length = 811
Score = 33.9 bits (76), Expect = 0.22
Identities = 22/81 (27%), Positives = 31/81 (38%), Gaps = 1/81 (1%)
Frame = +2
Query: 504 QVIPETPEPPETPEPTPEAVSNPTITAPPPP-KTRTSTRNTSTSLNNIEDGKVESDSDIQ 562
Q P + PP P T + SNP+ PPP KT TS+ + + E + S
Sbjct: 98 QTPPTSSNPPAAPPNTQLSASNPSPPTPPPSNKTHTSSSKLHSPSPSTEPSPPKPSSQDA 277
Query: 563 SMKSDVQSIKSIPVNTVIHNS 583
D + P T + S
Sbjct: 278 KTSGDSNPRSTPPSTTALRRS 340
>AV416351
Length = 428
Score = 33.9 bits (76), Expect = 0.22
Identities = 18/83 (21%), Positives = 35/83 (41%), Gaps = 7/83 (8%)
Frame = +3
Query: 500 QLLKQVIPETPEPPETPEPTPEAVSNPTITAPPPPKTRTSTRNTSTSLNNI-------ED 552
+++ +P +P P P + NP PPPP + +S ++ +N++ +
Sbjct: 18 KIVSTSLPSSPLPKAPPFSSSSPPRNPRPPPPPPPSSSSSPEKKNSPINHVLPSPTPHQS 197
Query: 553 GKVESDSDIQSMKSDVQSIKSIP 575
K SD+ M S + + P
Sbjct: 198 PKTIRQSDLSPMNSPKSKLSARP 266
>BP053311
Length = 529
Score = 30.8 bits (68), Expect = 1.9
Identities = 17/72 (23%), Positives = 29/72 (39%), Gaps = 11/72 (15%)
Frame = +2
Query: 836 RRDLGTFCEQFGIQGCP----------KKPKPRKQDPPPKQQWRKRSSRN-DHRKPKPRS 884
+ D+ +F Q G+ +K KP + P+ + ++ N H P P S
Sbjct: 29 KSDIASFASQLGLSTSQSYSGFNDVDFRKTKPNTEKATPQNTQKPKNDTNRPHEHPNPNS 208
Query: 885 KPQSSQIPKNPP 896
KP+ P+ P
Sbjct: 209 KPKXKXRPRXXP 244
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,714,154
Number of Sequences: 28460
Number of extensions: 427485
Number of successful extensions: 5610
Number of sequences better than 10.0: 218
Number of HSP's better than 10.0 without gapping: 3822
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5162
length of query: 1703
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1600
effective length of database: 1,966,220
effective search space: 3145952000
effective search space used: 3145952000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0019a.7


