

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.4
(310 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
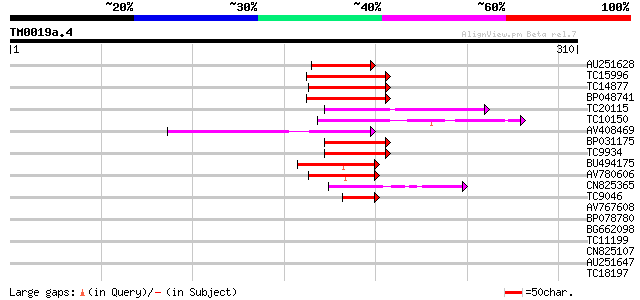
Sequences producing significant alignments: (bits) Value
AU251628 57 3e-09
TC15996 similar to UP|BAC98491 (BAC98491) AG-motif binding prote... 57 4e-09
TC14877 similar to GB|AAP37701.1|30725358|BT008342 At4g32890 {Ar... 57 5e-09
BP048741 55 1e-08
TC20115 similar to PIR|T52104|T52104 GATA-binding transcription ... 55 1e-08
TC10150 similar to UP|BAC98494 (BAC98494) AG-motif binding prote... 55 1e-08
AV408469 55 2e-08
BP031175 54 3e-08
TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protei... 54 3e-08
BU494175 48 2e-06
AV780606 47 4e-06
CN825365 45 2e-05
TC9046 homologue to UP|Q9LRH6 (Q9LRH6) ZIM, partial (10%) 39 9e-04
AV767608 38 0.002
BP078780 29 0.89
BG662098 29 0.89
TC11199 28 2.0
CN825107 27 3.4
AU251647 27 3.4
TC18197 27 3.4
>AU251628
Length = 350
Score = 57.4 bits (137), Expect = 3e-09
Identities = 22/35 (62%), Positives = 26/35 (73%)
Frame = +3
Query: 166 NNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNAC 200
++ S + C+DC TS TPLWR GP GPKSLCNAC
Sbjct: 246 SSGSEQKKTCADCGTSKTPLWRGGPAGPKSLCNAC 350
>TC15996 similar to UP|BAC98491 (BAC98491) AG-motif binding protein-1,
partial (19%)
Length = 555
Score = 57.0 bits (136), Expect = 4e-09
Identities = 23/46 (50%), Positives = 28/46 (60%)
Frame = +2
Query: 163 RNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
R ++ S R C C + TP WR GP GPK+LCNACG+R R R
Sbjct: 131 RFSSQGSAVPRKCMHCEVTKTPQWREGPMGPKTLCNACGVRYRSGR 268
>TC14877 similar to GB|AAP37701.1|30725358|BT008342 At4g32890 {Arabidopsis
thaliana;}, partial (25%)
Length = 1039
Score = 56.6 bits (135), Expect = 5e-09
Identities = 22/45 (48%), Positives = 30/45 (65%)
Frame = +3
Query: 164 NNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
+NN + R C+ C + TP WR+GP GPK+LCNACG+R + R
Sbjct: 441 SNNGQNPMPRRCTHCLSQRTPQWRAGPLGPKTLCNACGVRYKSGR 575
>BP048741
Length = 558
Score = 55.5 bits (132), Expect = 1e-08
Identities = 22/46 (47%), Positives = 29/46 (62%)
Frame = -1
Query: 163 RNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
RN+ + R CS C TP WR+GP+G K+LCNACG+R + R
Sbjct: 492 RNSCSDQAAPRRCSHCGVQKTPQWRTGPHGAKTLCNACGVRFKSGR 355
>TC20115 similar to PIR|T52104|T52104 GATA-binding transcription factor
homolog 2 [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (31%)
Length = 635
Score = 55.5 bits (132), Expect = 1e-08
Identities = 28/90 (31%), Positives = 43/90 (47%)
Frame = +3
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVH 232
R CS C + TP WR+GP GPK+LCNACG+R + R + A+P S + H
Sbjct: 99 RRCSHCASEKTPQWRTGPLGPKTLCNACGVRFKSGR--LVPEYRPAASPTFVLSQHSNSH 272
Query: 233 HNKEKKPRANHFAQFKNKSKSTSSNAGSSQ 262
+ R + + + + ++ A Q
Sbjct: 273 RKVMELRRQKELIRHQQQPQQPAAAASQDQ 362
>TC10150 similar to UP|BAC98494 (BAC98494) AG-motif binding protein-4,
partial (22%)
Length = 552
Score = 55.5 bits (132), Expect = 1e-08
Identities = 35/117 (29%), Positives = 54/117 (45%), Gaps = 3/117 (2%)
Frame = +3
Query: 169 SNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTK 228
+ + R CS C TP WR+GP G K+LCNACG+R + R L + A +
Sbjct: 51 AQSQRRCSHCQVQKTPQWRTGPLGAKTLCNACGVRYKSGR---------LCSEYRPACSP 203
Query: 229 T---RVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRNNST 282
T +H N +K + + + + +AG + C K F+I +R NS+
Sbjct: 204 TFCGEIHSNSHRK-----VLEIRRRKEVVGPDAGLTPTQMVPTC*K-FSI*IRGNSS 356
>AV408469
Length = 429
Score = 55.1 bits (131), Expect = 2e-08
Identities = 33/115 (28%), Positives = 49/115 (41%), Gaps = 1/115 (0%)
Frame = +2
Query: 87 NPSSSTCDPNLSFSKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKANNSTTIPIS- 145
+P+SS N + ++ S + S+ + K MM A+ + I +
Sbjct: 113 SPASSFIGENNTMKPNYHHHVRTSSDSENFAESQPVITKMMMPKQASGEPKKKKKIKVPL 292
Query: 146 PRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNAC 200
P + G+ N N S R C C + TP WR+GP GPK+LCNAC
Sbjct: 293 PLAPSDGN----------NIQNGSQPVRKCMHCEITKTPQWRAGPMGPKTLCNAC 427
>BP031175
Length = 537
Score = 54.3 bits (129), Expect = 3e-08
Identities = 21/36 (58%), Positives = 25/36 (69%)
Frame = -2
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
R C C T TP WR+GP GPK+LCNACG+R + R
Sbjct: 524 RRCLHCATDKTPQWRTGPMGPKTLCNACGVRYKSGR 417
>TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protein-4,
partial (37%)
Length = 1050
Score = 54.3 bits (129), Expect = 3e-08
Identities = 21/36 (58%), Positives = 25/36 (69%)
Frame = +2
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
R CS C TP WR+GP GPK+LCNACG+R + R
Sbjct: 569 RRCSHCQVQKTPQWRAGPLGPKTLCNACGVRFKSGR 676
>BU494175
Length = 475
Score = 47.8 bits (112), Expect = 2e-06
Identities = 21/47 (44%), Positives = 30/47 (63%), Gaps = 2/47 (4%)
Frame = +2
Query: 158 SQRSPRNNNNSSNTTRVCSDCNTS--STPLWRSGPNGPKSLCNACGI 202
+Q S +N +S + R C C S +TP R GP+GP++LCNACG+
Sbjct: 101 AQSSGQNGTPNSESLRRCQHCGVSENNTPAMRRGPDGPRTLCNACGL 241
>AV780606
Length = 585
Score = 47.0 bits (110), Expect = 4e-06
Identities = 20/41 (48%), Positives = 27/41 (65%), Gaps = 2/41 (4%)
Frame = -1
Query: 164 NNNNSSNTTRVCSDCNTSS--TPLWRSGPNGPKSLCNACGI 202
+ + S C+ C TSS TP+ R GP+GP+SLCNACG+
Sbjct: 468 SGQDDSQAETSCTHCGTSSKSTPMMRRGPSGPRSLCNACGL 346
>CN825365
Length = 511
Score = 44.7 bits (104), Expect = 2e-05
Identities = 28/76 (36%), Positives = 38/76 (49%)
Frame = +3
Query: 175 CSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHN 234
C C +STPLWR+GP LCNACG R R AN TP++ S V ++
Sbjct: 165 CYHCGVTSTPLWRNGPPEKPVLCNACGSRW----RTKGTLAN--YTPLH--SRADNVDND 320
Query: 235 KEKKPRANHFAQFKNK 250
++ R + + KNK
Sbjct: 321 DQRVSRLKNMSLNKNK 368
>TC9046 homologue to UP|Q9LRH6 (Q9LRH6) ZIM, partial (10%)
Length = 629
Score = 39.3 bits (90), Expect = 9e-04
Identities = 14/20 (70%), Positives = 18/20 (90%)
Frame = +3
Query: 183 TPLWRSGPNGPKSLCNACGI 202
TP+ R GP+GP+SLCNACG+
Sbjct: 3 TPMMRRGPSGPRSLCNACGL 62
>AV767608
Length = 398
Score = 37.7 bits (86), Expect = 0.002
Identities = 16/49 (32%), Positives = 27/49 (54%)
Frame = -1
Query: 190 PNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKK 238
P+GPK+LCNACG+R + R + P ++ + + +H N +K
Sbjct: 398 PHGPKTLCNACGVRFKSGRLVLEN------RPASSPTFRAELHSNSPRK 270
>BP078780
Length = 516
Score = 29.3 bits (64), Expect = 0.89
Identities = 12/25 (48%), Positives = 13/25 (52%)
Frame = -1
Query: 163 RNNNNSSNTTRVCSDCNTSSTPLWR 187
R N R DCN+S TPLWR
Sbjct: 432 RRGTNYCPAAR*LGDCNSSGTPLWR 358
>BG662098
Length = 466
Score = 29.3 bits (64), Expect = 0.89
Identities = 17/65 (26%), Positives = 39/65 (59%), Gaps = 2/65 (3%)
Frame = +1
Query: 67 SSSSNHLYNSTF--ISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVHGSAKWMSSKMRLM 124
SSS +H +S+ ++ P +P+S++C P+L F++ + + + HG+ +S++ L
Sbjct: 100 SSSQSHTQSSSLLRVNSSPRRVSPTSASCSPSLVFTQSKPQPL---HGTP--TTSRLHLA 264
Query: 125 KKMMS 129
+ ++S
Sbjct: 265 EPLVS 279
>TC11199
Length = 743
Score = 28.1 bits (61), Expect = 2.0
Identities = 27/113 (23%), Positives = 45/113 (38%), Gaps = 4/113 (3%)
Frame = +1
Query: 139 STTIPISPRIQNQG---HENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLW-RSGPNGPK 194
S+ P SP Q+ + +SPR+ ++ TTR S +T S P S P+ P
Sbjct: 163 SSARPSSPASQSHSTTTSATSLAPKSPRSKRSARRTTRTSSSPSTISDPSSPTSNPSSPP 342
Query: 195 SLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQF 247
SL + + L++P +T S+K ++ P + F
Sbjct: 343 SL----------TPTPSSSPSLALSSPPSTPSSKRGMYPGTSTSPSTRSESAF 471
>CN825107
Length = 254
Score = 27.3 bits (59), Expect = 3.4
Identities = 19/52 (36%), Positives = 27/52 (51%)
Frame = -2
Query: 64 SSSSSSSNHLYNSTFISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVHGSAK 115
SSSS SS+ ++T+ S + +PSSST S +E +KN AK
Sbjct: 187 SSSSPSSSFSPSTTYFSSETCNTSPSSST-------SSIESLPLKNFKKGAK 53
>AU251647
Length = 390
Score = 27.3 bits (59), Expect = 3.4
Identities = 18/62 (29%), Positives = 31/62 (49%)
Frame = +3
Query: 126 KMMSTPATDKANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPL 185
+++ PAT + +ST+ P P + G +N S ++NN S ++ DC S+ L
Sbjct: 78 RVLIKPATPSSGSSTS-PREPSRISLGGKNSNSGGGGDHDNNKSFSSEEDEDCEVVSSNL 254
Query: 186 WR 187
R
Sbjct: 255 QR 260
>TC18197
Length = 521
Score = 27.3 bits (59), Expect = 3.4
Identities = 12/51 (23%), Positives = 24/51 (46%)
Frame = -3
Query: 123 LMKKMMSTPATDKANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTR 173
++++ + TT + R N+ Y +R NNNNSS++++
Sbjct: 387 ILREQEEEEEAHRRRTQTTTQTNHRTTNEWQTVSYQKRRRNNNNNSSSSSK 235
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.125 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,070,389
Number of Sequences: 28460
Number of extensions: 95746
Number of successful extensions: 630
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 608
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 626
length of query: 310
length of database: 4,897,600
effective HSP length: 90
effective length of query: 220
effective length of database: 2,336,200
effective search space: 513964000
effective search space used: 513964000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0019a.4


