

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0016.15
(1469 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
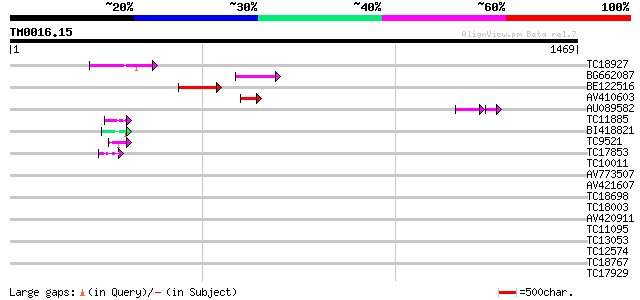
Sequences producing significant alignments: (bits) Value
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 135 6e-32
BG662087 92 6e-19
BE122516 88 1e-17
AV410603 75 7e-14
AU089582 50 7e-10
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 47 2e-05
BI418821 44 1e-04
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 43 3e-04
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 43 3e-04
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 41 0.002
AV773507 40 0.003
AV421607 40 0.003
TC18698 40 0.003
TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 38 0.010
AV420911 38 0.010
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 38 0.013
TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 34 0.19
TC12574 33 0.43
TC18767 32 0.73
TC17929 32 0.95
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 135 bits (339), Expect = 6e-32
Identities = 78/190 (41%), Positives = 98/190 (51%), Gaps = 15/190 (7%)
Frame = -2
Query: 207 KATEVELMKNRRLNRA-----GTGGPMRSGSQNFQNRGKFQNRRPYQRPAGRGFASGSYR 261
KA +E + N R A G+GGP RS F FQ ++P+QRP RG +SG Y
Sbjct: 560 KAKSIEAIDNLRSRPAFRPNQGSGGPNRSAPGRFDRNKSFQ-KKPFQRPQNRGTSSG-YS 387
Query: 262 PMVGAAGGSGDQTQNRELTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQCRVPKAEP 321
G Q+ E+ C +C K GHFA CPD C+NC K GH+ C PK E
Sbjct: 386 HSFGNFVPRPTQSDTSEIVCHRCSKKGHFANRCPDL--VCWNCQKTGHSGKDCTNPKVEA 213
Query: 322 TVNTA----------RGKRPAAKARVYTMDGDDTEGVDGLIRGDCEIDGNLLSVLFDSGA 371
N +GKRP A ARVYT+ G ++ DGLIR ++ L++LFDSGA
Sbjct: 212 ATNAIAARRPAPAANKGKRPVASARVYTVSGAESHRADGLIRSVGSVNCKPLTILFDSGA 33
Query: 372 THSFISRECA 381
THSFI CA
Sbjct: 32 THSFIDLACA 3
>BG662087
Length = 373
Score = 92.0 bits (227), Expect = 6e-19
Identities = 48/119 (40%), Positives = 68/119 (56%)
Frame = +1
Query: 584 GKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQK 643
GK R+ VDY LNK K+ YPLP ID L+D + S +D SGYHQI++ D K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 644 TAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEH 702
TAF T +Y Y +PFG+ NA A + M+R F + R + V++D++++ S H
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>BE122516
Length = 364
Score = 87.8 bits (216), Expect = 1e-17
Identities = 51/113 (45%), Positives = 68/113 (60%), Gaps = 1/113 (0%)
Frame = +2
Query: 438 LGMDWLSHYHCLLDCN*KRVVFPDSVLSEYLFANHISVSLNGGSQEYSVLLSLES-KGNP 496
+GM+WL+ L+C K V F S V + +L +LE+ K +
Sbjct: 14 VGMNWLTANDATLNCRKKTVTFGTSEGDAKRVKRTDKVGKASECESDVLLGALETDKSDT 193
Query: 497 EVDSIPVVKEFTDVFPGDVPGLPPVRDIEFAIDIVPGTGPISIAPYRMAPAEL 549
V+ IPVV+EF+DVFP +V LPP R++EF+ID VPGTGPISIAPYRM+ EL
Sbjct: 194 GVEGIPVVREFSDVFPEEVSELPPEREVEFSID*VPGTGPISIAPYRMSLVEL 352
>AV410603
Length = 162
Score = 75.1 bits (183), Expect = 7e-14
Identities = 32/53 (60%), Positives = 43/53 (80%)
Frame = +1
Query: 599 TVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYG 651
TVK+ +P+P +D+L+D+LRG+ FSK+DL+SGYHQI VK ED KT FRT +G
Sbjct: 4 TVKDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>AU089582
Length = 383
Score = 50.1 bits (118), Expect(2) = 7e-10
Identities = 34/75 (45%), Positives = 40/75 (53%)
Frame = +3
Query: 1156 NYQRFI*LRLYDFMECQQVLFPIETRSLLHIFGELFKRLWERG*D*VLLIIRRLMDRLRG 1215
N RF +RL+ M + F IE + HIFG LFK LWE * *V L I +LM RG
Sbjct: 18 NMLRFTWMRLFPCMVYLCL*FQIEELNSHHIFGGLFKLLWELD*K*VPLFILKLMVSPRG 197
Query: 1216 LSNHWRICYEHAC*T 1230
L RIC+ C T
Sbjct: 198 LFRS*RICFVLVCXT 242
Score = 31.6 bits (70), Expect(2) = 7e-10
Identities = 16/42 (38%), Positives = 20/42 (47%)
Frame = +1
Query: 1232 RAVGIAYCH*SSSHTTIVFIRP*VWHRMRLCMAASVEHLYVG 1273
R GI+ C * +SH I WH ++ CM HL VG
Sbjct: 247 RXAGISICL*WNSHIIIAIGLAFRWHHLKPCMVGDAGHL*VG 372
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 47.0 bits (110), Expect = 2e-05
Identities = 24/69 (34%), Positives = 34/69 (48%)
Frame = +2
Query: 246 PYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACPDTRPQCYNCN 305
PY+R + RGF+ + G + N + C CG PGH A C T+ C+NC
Sbjct: 299 PYRRDSRRGFSRDNLCKNCKRPGHFARECPNVAI-CHNCGLPGHIASECT-TKSLCWNCK 472
Query: 306 KLGHTAAQC 314
+ GH A+ C
Sbjct: 473 EPGHMASSC 499
>BI418821
Length = 614
Score = 44.3 bits (103), Expect = 1e-04
Identities = 28/86 (32%), Positives = 30/86 (34%), Gaps = 8/86 (9%)
Frame = +2
Query: 237 NRGKFQNRRPYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACPD 296
N Q R P G G G R G GG G C+ CG GH AR C
Sbjct: 281 NGAALQPTRKDSAPRGFGGWRGGERRNGGGGGGGGGG-------CYNCGDTGHLARDCHR 439
Query: 297 TR--------PQCYNCNKLGHTAAQC 314
+ CYNC GH A C
Sbjct: 440 SNNNGGGGGGAACYNCGDAGHLARDC 517
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 43.1 bits (100), Expect = 3e-04
Identities = 25/73 (34%), Positives = 34/73 (46%), Gaps = 12/73 (16%)
Frame = +1
Query: 255 FASGSYRPMVGAAGGSGDQTQNRELT----------CFKCGKPGHFARACP--DTRPQCY 302
FA G R G+ G G + ++RE CF CG GH+AR C D + +CY
Sbjct: 292 FAKGVPR---GSREGGGGRDRDREYMGRGPPPGSGRCFNCGIDGHWARDCKAGDWKNKCY 462
Query: 303 NCNKLGHTAAQCR 315
C + GH C+
Sbjct: 463 RCGERGHIEKNCK 501
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 43.1 bits (100), Expect = 3e-04
Identities = 24/67 (35%), Positives = 33/67 (48%)
Frame = +2
Query: 229 RSGSQNFQNRGKFQNRRPYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPG 288
R ++ RG+ ++ R R RG+ G+ G +G + CFKCGKPG
Sbjct: 311 RDDGDRYRERGRDRDDRG-DRDRSRGYG--------GSRGSNGGE-------CFKCGKPG 442
Query: 289 HFARACP 295
HFAR CP
Sbjct: 443 HFARECP 463
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 40.8 bits (94), Expect = 0.002
Identities = 16/37 (43%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +2
Query: 281 CFKCGKPGHFARACP--DTRPQCYNCNKLGHTAAQCR 315
CF CG GH+AR C D + +CY C GH C+
Sbjct: 371 CFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVERNCK 481
>AV773507
Length = 496
Score = 40.0 bits (92), Expect = 0.003
Identities = 18/45 (40%), Positives = 26/45 (57%)
Frame = +2
Query: 251 AGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACP 295
A +G+ G Y + G+GD+ + CF+CG+PGH AR CP
Sbjct: 266 ADQGYRGGGYSSGGRGSYGAGDRVGQDD--CFECGRPGHRARDCP 394
>AV421607
Length = 245
Score = 39.7 bits (91), Expect = 0.003
Identities = 16/35 (45%), Positives = 22/35 (62%)
Frame = +3
Query: 265 GAAGGSGDQTQNRELTCFKCGKPGHFARACPDTRP 299
G+ GSG T C+KCG+PGH++R CP + P
Sbjct: 21 GSGSGSGTATG-----CYKCGRPGHWSRDCPSSAP 110
>TC18698
Length = 808
Score = 39.7 bits (91), Expect = 0.003
Identities = 20/69 (28%), Positives = 36/69 (51%)
Frame = -2
Query: 642 QKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVE 701
+KT + +Y Y VMP G+ N + M++ FH + + V V+++D+++ S
Sbjct: 804 KKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*F 625
Query: 702 HEGHLRQVL 710
H G L + L
Sbjct: 624 HRGDLSRDL 598
>TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (42%)
Length = 587
Score = 38.1 bits (87), Expect = 0.010
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = +1
Query: 265 GAAGGSGDQTQNRELTCFKCGKPGHFARAC 294
G G G +L C++CG+PGHFAR C
Sbjct: 118 GGGRGGGRGRGGEDLKCYECGEPGHFAREC 207
>AV420911
Length = 418
Score = 38.1 bits (87), Expect = 0.010
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = +2
Query: 265 GAAGGSGDQTQNRELTCFKCGKPGHFARAC 294
G G G +L C++CG+PGHFAR C
Sbjct: 326 GGGRGGGRGRGGEDLKCYECGEPGHFAREC 415
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 37.7 bits (86), Expect = 0.013
Identities = 18/46 (39%), Positives = 22/46 (47%)
Frame = +1
Query: 249 RPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARAC 294
R G G G G GG D + C++CG+PGHFAR C
Sbjct: 301 RGGGGGGGGGRGGGRSGGGGGGSD------MKCYECGEPGHFAREC 420
>TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (9%)
Length = 450
Score = 33.9 bits (76), Expect = 0.19
Identities = 13/36 (36%), Positives = 18/36 (49%)
Frame = +3
Query: 279 LTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQC 314
+ C C + GH +R C C+NC GH A +C
Sbjct: 3 VVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 110
>TC12574
Length = 325
Score = 32.7 bits (73), Expect = 0.43
Identities = 13/33 (39%), Positives = 23/33 (69%)
Frame = +2
Query: 669 FMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVE 701
F + +N F +F + F++VFI+DIL Y+++ E
Sbjct: 2 FKNSVNHIFESFFEHFMIVFINDILSYTEDKEE 100
>TC18767
Length = 1004
Score = 32.0 bits (71), Expect = 0.73
Identities = 11/35 (31%), Positives = 20/35 (56%)
Frame = +2
Query: 300 QCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAK 334
+C+NC H+ +C P+ VN+AR +R + +
Sbjct: 164 RCFNCGSYNHSLRECSRPRDNVAVNSARKQRKSRR 268
>TC17929
Length = 791
Score = 31.6 bits (70), Expect = 0.95
Identities = 13/31 (41%), Positives = 18/31 (57%), Gaps = 6/31 (19%)
Frame = +2
Query: 271 GDQTQNR------ELTCFKCGKPGHFARACP 295
G++T NR TC++CG+ GH R CP
Sbjct: 14 GEETSNRPNDSKFRQTCYRCGESGHKMRNCP 106
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.345 0.152 0.510
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,729,595
Number of Sequences: 28460
Number of extensions: 375680
Number of successful extensions: 3281
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 3185
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3267
length of query: 1469
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1367
effective length of database: 1,994,680
effective search space: 2726727560
effective search space used: 2726727560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0016.15


