

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0014.18
(181 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
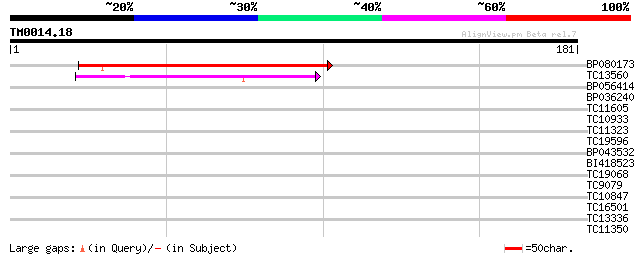
Score E
Sequences producing significant alignments: (bits) Value
BP080173 68 1e-12
TC13560 44 2e-05
BP056414 37 0.003
BP036240 35 0.006
TC11605 35 0.007
TC10933 similar to UP|Q9LNS0 (Q9LNS0) F1L3.4, partial (6%) 34 0.013
TC11323 31 0.14
TC19596 30 0.18
BP043532 30 0.18
BI418523 30 0.24
TC19068 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltra... 27 1.6
TC9079 weakly similar to UP|Q8W4G1 (Q8W4G1) UDP-glucose glucosyl... 27 1.6
TC10847 similar to UP|Q8RX88 (Q8RX88) AT4g01320/F2N1_21, partial... 26 3.5
TC16501 homologue to UP|Q8LJS9 (Q8LJS9) WD-repeat protein GhTTG4... 26 4.5
TC13336 similar to UP|Q8VZH0 (Q8VZH0) AT3g58640/F14P22_230, part... 25 5.9
TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3,... 25 7.7
>BP080173
Length = 382
Score = 67.8 bits (164), Expect = 1e-12
Identities = 31/82 (37%), Positives = 51/82 (61%), Gaps = 1/82 (1%)
Frame = -2
Query: 23 WPLDAV-WVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILC 81
WP+ W+K+N DG++ S++ AAC G++ D F+ + RLG Y AELWGI
Sbjct: 345 WPVPMEGWIKINIDGSFAPSSNCAACGGVMWDHEGTFITGASVRLGCYFINHAELWGIFH 166
Query: 82 GARLLQERDY*RVLIETNLSIL 103
GA++ Q R + ++L+E++ S +
Sbjct: 165 GAKIAQARGFKKILVESDSSFV 100
>TC13560
Length = 571
Score = 43.9 bits (102), Expect = 2e-05
Identities = 24/81 (29%), Positives = 43/81 (52%), Gaps = 3/81 (3%)
Frame = +2
Query: 22 WWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQ---AELWG 78
W DA WVK+N DGA +S+ A+CRG++R+ +++ + G++S+ EL
Sbjct: 146 WIKPDAGWVKINVDGA-VSNCLQASCRGVLRNADGSWLKGFRWNFGTFSSANVFLTELMA 322
Query: 79 ILCGARLLQERDY*RVLIETN 99
+ D RV++E++
Sbjct: 323 VKTAVEAAMSLDLARVIVESD 385
>BP056414
Length = 606
Score = 36.6 bits (83), Expect = 0.003
Identities = 24/86 (27%), Positives = 40/86 (45%), Gaps = 3/86 (3%)
Frame = -3
Query: 17 APAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGS---YSTLQ 73
A + W W K+N DGA +S A+C G++RD ++ +AR LG +
Sbjct: 577 ASSYHWTRPPEGWFKVNVDGA-LSGDLQASCGGVVRDAAGSWSKGFARSLGVLRWHXAFF 401
Query: 74 AELWGILCGARLLQERDY*RVLIETN 99
EL + + D +V+IE++
Sbjct: 400 VELMAVQTAVDFIMSWDIPQVIIESD 323
>BP036240
Length = 567
Score = 35.4 bits (80), Expect = 0.006
Identities = 21/81 (25%), Positives = 38/81 (45%), Gaps = 3/81 (3%)
Frame = -3
Query: 22 WWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGS---YSTLQAELWG 78
W D W K + DGA +S +A+C G++RD ++ + R +G ++ EL
Sbjct: 562 WTAPDEGWTKFDVDGA-LSRDLLASCGGVLRDSRRHWIKGFCRNMGDLTYHNVFLVELSA 386
Query: 79 ILCGARLLQERDY*RVLIETN 99
I + D V++E++
Sbjct: 385 IQSAVEIALSMDLQHVIVESD 323
>TC11605
Length = 546
Score = 35.0 bits (79), Expect = 0.007
Identities = 20/59 (33%), Positives = 32/59 (53%)
Frame = +3
Query: 22 WWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGIL 80
W P +A +K N DG+ S ++ GI+R+ SA + +++ LG QAE+ IL
Sbjct: 348 WCPPEANILKFNVDGSARGSPGISGAGGILRNAESAVLGRFSKPLGVLCAYQAEVKAIL 524
>TC10933 similar to UP|Q9LNS0 (Q9LNS0) F1L3.4, partial (6%)
Length = 596
Score = 34.3 bits (77), Expect = 0.013
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +2
Query: 21 RWWPLDAVWVKLNSDGAYISSADMAACRGI 50
RW P WVKLN+DG++ + + C G+
Sbjct: 104 RWTPPSVGWVKLNTDGSFTRGSSLMGCGGL 193
>TC11323
Length = 657
Score = 30.8 bits (68), Expect = 0.14
Identities = 20/73 (27%), Positives = 36/73 (48%)
Frame = +2
Query: 31 KLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQERD 90
KLN DG+ + + A G++RD S F+ ++ L ++ L IL G + +
Sbjct: 32 KLNFDGSSLGNRGNAGGGGLLRDGSSNFIFGFSIFLAVAQIMKLSLCAILEGLLVCKPLG 211
Query: 91 Y*RVLIETNLSIL 103
Y + IE + +I+
Sbjct: 212 YDGIGIECDSNIV 250
>TC19596
Length = 499
Score = 30.4 bits (67), Expect = 0.18
Identities = 20/57 (35%), Positives = 30/57 (52%), Gaps = 1/57 (1%)
Frame = +1
Query: 31 KLNSDGAYISSADMAACRGIIRDRFSAFVIA-YARRLGSYSTLQAELWGILCGARLL 86
+++SDGA+ D GI+RD A++ YA LG L+AE+ + G LL
Sbjct: 4 RMDSDGAFKHDDDRMGMGGIVRDAHGAWISGFYAGSLGG-DALRAEIAALKHGLTLL 171
>BP043532
Length = 465
Score = 30.4 bits (67), Expect = 0.18
Identities = 14/34 (41%), Positives = 21/34 (61%)
Frame = -2
Query: 66 LGSYSTLQAELWGILCGARLLQERDY*RVLIETN 99
+G S L AELWG++ G R R + ++LIE +
Sbjct: 464 VGVCSILHAELWGLVHGLRFALGRGFSKILIEAD 363
>BI418523
Length = 496
Score = 30.0 bits (66), Expect = 0.24
Identities = 22/105 (20%), Positives = 48/105 (44%), Gaps = 6/105 (5%)
Frame = +1
Query: 4 VAQVTRVVSPVRDAPAIR------WWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSA 57
+ Q+ + + +R P ++ W P A ++K N DGA + + G+ RD
Sbjct: 115 IYQIMQNLGDIRITPKLKPPRCWSWSPPSAGFIKCNVDGASQGNPGPSGVGGVFRDANRK 294
Query: 58 FVIAYARRLGSYSTLQAELWGILCGARLLQERDY*RVLIETNLSI 102
+ ++ G+ +A++ IL +Q+ +LIE++ ++
Sbjct: 295 ILGYFSLNSGNGWAYEAKVRSILNALVFIQKFLLKNILIESDSTV 429
>TC19068 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltransferase
85A8, partial (26%)
Length = 565
Score = 27.3 bits (59), Expect = 1.6
Identities = 8/16 (50%), Positives = 12/16 (75%)
Frame = +3
Query: 18 PAIRWWPLDAVWVKLN 33
P WW LD++W++LN
Sbjct: 93 PPFNWWFLDSLWMELN 140
>TC9079 weakly similar to UP|Q8W4G1 (Q8W4G1) UDP-glucose glucosyltransferase,
partial (65%)
Length = 1585
Score = 27.3 bits (59), Expect = 1.6
Identities = 8/16 (50%), Positives = 12/16 (75%)
Frame = +3
Query: 18 PAIRWWPLDAVWVKLN 33
P WW LD++W++LN
Sbjct: 1095 PPFSWWILDSLWMELN 1142
>TC10847 similar to UP|Q8RX88 (Q8RX88) AT4g01320/F2N1_21, partial (24%)
Length = 720
Score = 26.2 bits (56), Expect = 3.5
Identities = 13/41 (31%), Positives = 24/41 (57%)
Frame = +3
Query: 90 DY*RVLIETNLSILGVVLVLIPVLLLLTK*SRLFSRVSPRL 130
DY R + + + ILG +L IP+L LL +L +++ ++
Sbjct: 189 DYRRRICQL*IQILGTLLTTIPILPLLKDWKQLTNQIKKQI 311
>TC16501 homologue to UP|Q8LJS9 (Q8LJS9) WD-repeat protein GhTTG4, partial
(23%)
Length = 611
Score = 25.8 bits (55), Expect = 4.5
Identities = 18/40 (45%), Positives = 24/40 (60%), Gaps = 2/40 (5%)
Frame = -1
Query: 78 GILCGARLLQERDY*RVLIETN--LSILGVVLVLIPVLLL 115
GIL R+ + +L ETN L +LG+ LVLIP LL+
Sbjct: 485 GILQ*TRM*ETIQTINILPETNQYLFLLGIFLVLIPGLLI 366
>TC13336 similar to UP|Q8VZH0 (Q8VZH0) AT3g58640/F14P22_230, partial (7%)
Length = 584
Score = 25.4 bits (54), Expect = 5.9
Identities = 12/28 (42%), Positives = 17/28 (59%), Gaps = 2/28 (7%)
Frame = -3
Query: 3 RVAQVTRV-VSPVRDAPA-IRWWPLDAV 28
++ +VTRV PV P+ + WWP D V
Sbjct: 129 KIMEVTRVDAGPVAQRPSNVSWWPSDFV 46
>TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3, P1 clone:
MIG10, partial (18%)
Length = 1118
Score = 25.0 bits (53), Expect = 7.7
Identities = 19/78 (24%), Positives = 33/78 (41%)
Frame = -3
Query: 22 WWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILC 81
W P KLNSD ++ + + +IRD + A R + S + +E +
Sbjct: 795 WRPPPLGVYKLNSDASWKAPSPSCTVGVVIRDNLGLLISGTASRPLAPSPIVSEALALRE 616
Query: 82 GARLLQERDY*RVLIETN 99
G L + R+L E++
Sbjct: 615 GLILARSLGLDRILSESD 562
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.336 0.145 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,116,196
Number of Sequences: 28460
Number of extensions: 39067
Number of successful extensions: 285
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 283
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 284
length of query: 181
length of database: 4,897,600
effective HSP length: 85
effective length of query: 96
effective length of database: 2,478,500
effective search space: 237936000
effective search space used: 237936000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0014.18


