

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0014.15
(766 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
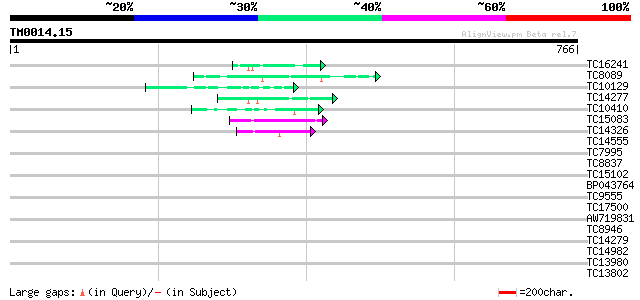
Sequences producing significant alignments: (bits) Value
TC16241 weakly similar to UP|Q91TH4 (Q91TH4) T122, partial (7%) 48 5e-06
TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinog... 47 1e-05
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 46 3e-05
TC14277 similar to UP|Q9SHY3 (Q9SHY3) F1E22.9 (AT1G65720/F1E22_1... 44 1e-04
TC10410 weakly similar to UP|Q8H5W8 (Q8H5W8) OJ1123_B01.10 prote... 42 4e-04
TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, p... 41 6e-04
TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific... 41 8e-04
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 40 0.001
TC7995 similar to PIR|S63686|S63686 sterol 24-C-methyltransferas... 40 0.001
TC8837 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein ki... 40 0.002
TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, ... 39 0.002
BP043764 39 0.003
TC9555 similar to UP|Q9MUE2 (Q9MUE2) S-adenosyl-L-methionine Mg-... 39 0.003
TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase t... 39 0.003
AW719831 39 0.004
TC8946 homologue to GB|AAM66972.1|21617922|AY088650 pyridoxine b... 39 0.004
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 39 0.004
TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%) 38 0.005
TC13980 similar to UP|Q9SGT0 (Q9SGT0) T6H22.18 protein (F14J16.3... 38 0.005
TC13802 similar to UP|Q9M1T1 (Q9M1T1) Sugar-phosphate isomerase-... 38 0.007
>TC16241 weakly similar to UP|Q91TH4 (Q91TH4) T122, partial (7%)
Length = 799
Score = 48.1 bits (113), Expect = 5e-06
Identities = 42/141 (29%), Positives = 56/141 (38%), Gaps = 15/141 (10%)
Frame = +1
Query: 301 SSNPLTPIPESQPAAQTTSPP--------HSPRS-------PFFQPSPTEAPLWNLLKNP 345
S P TP P +P + T+ PP HSP S P PSP+ P + L P
Sbjct: 7 SQPPYTPAP-IKPPSPTSQPPYIATPPNTHSPTSQPPNTQTPANTPSPSSQPPY--LSTP 177
Query: 346 TSRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDS 405
+S PTS + P + PI P P P T+ P+ + + S
Sbjct: 178 P-KSNSPTSQPPVAQTPRAVTPISSPSSSPTSNSPYPSTNP---------PSLAPSISTS 327
Query: 406 STAFTPVSFPINVIDSPPSNT 426
A TP S P+ SPPS +
Sbjct: 328 PPALTPASTPVPATASPPSQS 390
>TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (43%)
Length = 890
Score = 47.0 bits (110), Expect = 1e-05
Identities = 61/264 (23%), Positives = 99/264 (37%), Gaps = 11/264 (4%)
Frame = +2
Query: 249 PTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSNPLTPI 308
PT++ P + P R++R SS L P ++ S P +P
Sbjct: 122 PTQISPRQCSCSARQ-PWRLSRQRCSSSPWLLPPQT--------PTTSPAFSPSTPSSP- 271
Query: 309 PESQPAAQTTSPPHSPRSPFFQPSPTEAPLW-----NLLKNPTSRSEDPTSLLTIPYDPV 363
P + + TSPP S P +P+ P W N ++P SR+ P++ + P
Sbjct: 272 PSTTTSPSPTSPPRSTNGPQSPSAPSTTPPWTTSSQNTHQSPPSRTSSPSTSSSTTSAPR 451
Query: 364 SSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVID--- 420
SS P P Q +P + P +S P + +DS+ T P+ +
Sbjct: 452 SSTRSPTAPPSPPQC-TKPPAPPRAPPDSSTSPTSAAGKSDSAPRTTTAPSPLRSSNP*R 628
Query: 421 -SP-PSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYP-GPRPERLVDPDEPILANPI 477
SP S +S S+R F P P++ P P P + P P+ + +
Sbjct: 629 KSPTTSPSSRSVRFF------------------PPPQQKPLHPHPLSRISP--PLCPSTV 748
Query: 478 QEADPLVQQAHPVPDQQEPIQPDP 501
+ P + P ++ QP P
Sbjct: 749 ARSSPTLS-----PPRRTRTQPSP 805
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 45.8 bits (107), Expect = 3e-05
Identities = 56/207 (27%), Positives = 77/207 (37%)
Frame = +1
Query: 184 LPPAPEFPSPPKKKTKKKMVLEESSEESDVPLVKKTKSKPDDGDDDDSEDGPPKKKQKKV 243
+P P+ PSP + + + S P V P + PPK
Sbjct: 28 IPTPPKTPSPGNQPPHTPIPPKTPSPSYSPPNVPSPPKAPSPNNHPPYTPTPPKTPS--- 198
Query: 244 RIVVKPTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSN 303
PT P + +T +P +++ + P+ S PIS +
Sbjct: 199 -----PTSQPPYIPLPPKTPSP--ISQPPHVPTPPSTPS----------PISQPPYTPTP 327
Query: 304 PLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPV 363
P TP P SQP T +PP++P SP QP T P K P+ ++ P IP P
Sbjct: 328 PKTPSPTSQP-PHTPTPPNTP-SPISQPPYTPTP----PKTPSPTNQPP----HIPSPPN 477
Query: 364 SSEPIIHEQPEPNQTEPQPRTSDHSVP 390
SS P QP PN P S S P
Sbjct: 478 SSSPT--SQP-PNTPSPPKTPSPTSQP 549
Score = 36.6 bits (83), Expect = 0.015
Identities = 39/138 (28%), Positives = 53/138 (38%), Gaps = 13/138 (9%)
Frame = +1
Query: 304 PLTPIPESQPA---------AQTTSPPHSPRSPFFQPSPTEAPLWNLL--KNPTSRSEDP 352
P TP P SQP + + PPH P P PSP P + K P+ S+ P
Sbjct: 184 PKTPSPTSQPPYIPLPPKTPSPISQPPHVPTPP-STPSPISQPPYTPTPPKTPSPTSQPP 360
Query: 353 TSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPV 412
+ P P + PI QP T P+ + + P + P + +P
Sbjct: 361 HT----PTPPNTPSPI--SQPPYTPTPPKTPSPTNQPPHIPSPP----------NSSSPT 492
Query: 413 SFPINVIDSP--PSNTSE 428
S P N P PS TS+
Sbjct: 493 SQPPNTPSPPKTPSPTSQ 546
>TC14277 similar to UP|Q9SHY3 (Q9SHY3) F1E22.9 (AT1G65720/F1E22_13), partial
(33%)
Length = 941
Score = 43.5 bits (101), Expect = 1e-04
Identities = 51/177 (28%), Positives = 71/177 (39%), Gaps = 16/177 (9%)
Frame = +3
Query: 282 SDDDLNLFDALPISALLQHSSNPLTPIPESQPAAQTTSPP---------HSPRSPFFQPS 332
SD D L + + SS+P P P PAA ++SPP SP SP PS
Sbjct: 54 SDSDPWLLSSAASRSPPSSSSSPSPPPPPHAPAAASSSPPTPSATLPPTPSPPSPRSDPS 233
Query: 333 P----TEAPLWNLLKNP--TSRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSD 386
P T P + P T+ +P + P S P++ P+ T PRTS
Sbjct: 234 PLSTSTTNPTTSSSTTPKTTTNKSNPNTARLSHAAP*GSPPLLTTTSPPSVT--APRTSS 407
Query: 387 HS-VPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSESIRKFMEVRKEKVS 442
S +P +SA AA + T+ P S P+ P + S R+ R+ S
Sbjct: 408 ASRLPSSSALDAALSPPL-PCTSSGPSSPPVTTTAPPCTAISPMTRRSTAPRRPATS 575
>TC10410 weakly similar to UP|Q8H5W8 (Q8H5W8) OJ1123_B01.10 protein, partial
(12%)
Length = 431
Score = 42.0 bits (97), Expect = 4e-04
Identities = 47/182 (25%), Positives = 70/182 (37%), Gaps = 3/182 (1%)
Frame = +3
Query: 246 VVKPTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSNPL 305
++ P AAA R T P++ S+P L P S+L Q SS+
Sbjct: 3 LMDPQPPATAAATARSTSEPSQ--------SQPPLQ-----------PPSSLPQTSSSTT 125
Query: 306 TPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSS 365
PI ++PPH P++P P+P AP NP P +
Sbjct: 126 PPI---------SAPPHPPQNP--NPTPIPAP------NPNPN----------PIQIQNP 224
Query: 366 EPIIHEQPEPNQTEPQPR---TSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSP 422
PI + P P T PQPR T + + P +P+ + + + P P + +S
Sbjct: 225 NPIHNPNPRPTPTPPQPRPPPTFNRAPPPQQQQPSHFSHFSSLPPSSAPSPSPASSFNST 404
Query: 423 PS 424
PS
Sbjct: 405 PS 410
Score = 33.5 bits (75), Expect = 0.13
Identities = 45/192 (23%), Positives = 67/192 (34%)
Frame = +3
Query: 368 IIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTS 427
++ QP TS+ S + +P + T SST TP PI+ PP N +
Sbjct: 3 LMDPQPPATAAATARSTSEPSQSQPPLQPPSSLPQTSSST--TP---PISAPPHPPQNPN 167
Query: 428 ESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEADPLVQQA 487
P+P P P P NPIQ +Q
Sbjct: 168 ------------------------PTPIPAPNPNP------------NPIQ-----IQNP 224
Query: 488 HPVPDQQEPIQPDPEPEQSVSNQSSVRSPHPLVETSDHHRGTSEPNAPMINIGSPQGASE 547
+P+ + +P P P Q + R+P P + H S + P + SP AS
Sbjct: 225 NPIHNPNP--RPTPTPPQPRPPPTFNRAPPPQQQQPSHFSHFS--SLPPSSAPSPSPASS 392
Query: 548 AHSSNHQASPEP 559
+S+ + P P
Sbjct: 393 FNSTPSPSIPAP 428
>TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, partial (7%)
Length = 1367
Score = 41.2 bits (95), Expect = 6e-04
Identities = 37/133 (27%), Positives = 58/133 (42%), Gaps = 1/133 (0%)
Frame = +2
Query: 298 LQHSSNPLTP-IPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLL 356
L SS TP + + P+A +PP S + PSPT +PT+ S P +
Sbjct: 185 LNQSSETSTPNMAQLSPSASAPAPPSSSPT---APSPTVPSFKTPPYSPTALSPSPPAKS 355
Query: 357 TIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPI 416
+ P S+ P++ + P+ R S V S +P + T SSTA P+ P
Sbjct: 356 SPPTSTTSALPLMAQPGVPSDATSPARCSTPPV*SPSHKP-GNGSSTRSSTA*NPLP-PN 529
Query: 417 NVIDSPPSNTSES 429
++ PP + S +
Sbjct: 530 QILTPPPLSLSSA 568
>TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific
extensin-like protein precursor (PELP), partial (11%)
Length = 630
Score = 40.8 bits (94), Expect = 8e-04
Identities = 30/113 (26%), Positives = 47/113 (41%), Gaps = 6/113 (5%)
Frame = +2
Query: 307 PIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDP---- 362
P P QP + SPP P+F P P +AP ++K+P ++ P + + P P
Sbjct: 221 PTPPPQPPVKAPSPPIVKSPPYF-PPPVKAPSPPIVKSPPVKAPTPPIVKSPPSYPPPVK 397
Query: 363 VSSEPIIHEQPEPNQTEPQPRTSDHSVP--RASARPAARTTDTDSSTAFTPVS 413
S PI+ P T P ++ + P +A P +T PV+
Sbjct: 398 APSPPIVKSPPVKAPTPPIVKSPPYYPPPVKAPTPPQVKTPPPHPPVVKPPVA 556
Score = 33.5 bits (75), Expect = 0.13
Identities = 24/79 (30%), Positives = 37/79 (46%)
Frame = +2
Query: 304 PLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPV 363
P PI +S P T P +SP P P +AP ++K+P ++ P + + PY P
Sbjct: 308 PSPPIVKSPPVKAPTPP--IVKSPPSYPPPVKAPSPPIVKSPPVKAPTPPIVKSPPYYP- 478
Query: 364 SSEPIIHEQPEPNQTEPQP 382
P+ + P P Q + P
Sbjct: 479 --PPV--KAPTPPQVKTPP 523
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 40.4 bits (93), Expect = 0.001
Identities = 48/168 (28%), Positives = 59/168 (34%), Gaps = 9/168 (5%)
Frame = +2
Query: 254 PAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSN-------PLT 306
PAA+ + P + H+ L L L + S Q SN P T
Sbjct: 62 PAASPGAQVNQPREMKMMDTHNMHVVLV----LGLICIVIASVGAQSPSNSPTTSPAPPT 229
Query: 307 PIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSE 366
P PAAQ++ PP P Q SP A P ++S P PVSS
Sbjct: 230 PTTPQPPAAQSSPPPAQSSPPPVQSSPPPAST-----PPPAQSSPP---------PVSSP 367
Query: 367 PIIHE--QPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPV 412
P + P P T P S P A+ P A T A TPV
Sbjct: 368 PPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAAT----PPPALTPV 499
Score = 28.5 bits (62), Expect = 4.2
Identities = 16/60 (26%), Positives = 25/60 (41%)
Frame = +2
Query: 364 SSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPP 423
S P P+P + P + S P + P +T + ++ PVS P V +PP
Sbjct: 212 SPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPVSSPPPVQSTPP 391
Score = 28.1 bits (61), Expect = 5.5
Identities = 21/77 (27%), Positives = 29/77 (37%), Gaps = 1/77 (1%)
Frame = +2
Query: 261 RTEAPARVTRSSAHSSKPALASDDDLNLFDA-LPISALLQHSSNPLTPIPESQPAAQTTS 319
++ P T A SS P ++S + P S S P +P P + P T
Sbjct: 299 QSSPPPASTPPPAQSSPPPVSSPPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATP 478
Query: 320 PPHSPRSPFFQPSPTEA 336
PP P P+P A
Sbjct: 479 PPALTPVPATSPAPAPA 529
Score = 27.3 bits (59), Expect = 9.4
Identities = 41/166 (24%), Positives = 59/166 (34%), Gaps = 10/166 (6%)
Frame = +2
Query: 182 SRLPPAPEFPSPPKKKTKKKMVLEESSEESDVPLVKKTKSKPDDGDDDDSEDGPPKKKQK 241
S PP P P PP ++ +S P V+ S P ++ PP
Sbjct: 212 SPAPPTPTTPQPPAAQSSPPPA------QSSPPPVQS--SPPPASTPPPAQSSPP----- 352
Query: 242 KVRIVVKPTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHS 301
P P P + T A +S P AS F P + +
Sbjct: 353 -------PVSSPP----------PVQSTPPPAPASTPPPASPPP---FSPPPATPPPPAA 472
Query: 302 SNP--LTPIPESQPA-----AQTTSPPHSPR---SPFFQPSPTEAP 337
+ P LTP+P + PA ++ SP +P +P SP+EAP
Sbjct: 473 TPPPALTPVPATSPAPAPAKVKSKSPAPAPAPALAPVVSLSPSEAP 610
>TC7995 similar to PIR|S63686|S63686 sterol 24-C-methyltransferase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (98%)
Length = 1524
Score = 40.0 bits (92), Expect = 0.001
Identities = 40/147 (27%), Positives = 55/147 (37%)
Frame = +3
Query: 254 PAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSNPLTPIPESQP 313
P+ + +PA T SSA S+ P ++ A PIS P P P +
Sbjct: 153 PSLSSAPELSSPAAYTGSSAFSAPPNKRAN-------APPIS--------PADPSPPRKS 287
Query: 314 AAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQP 373
TTS S +P P +P+ + S PTS V + P I P
Sbjct: 288 RTTTTSTGPSSAAPRRSKPPRRSPI----SSTPSTISSPTST-----SGVGASPSISPLP 440
Query: 374 EPNQTEPQPRTSDHSVPRASARPAART 400
P PR S S+P +RP+ T
Sbjct: 441 SPGNLTATPRASTRSMPSI*SRPSRVT 521
>TC8837 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(15%)
Length = 606
Score = 39.7 bits (91), Expect = 0.002
Identities = 48/176 (27%), Positives = 64/176 (36%)
Frame = -3
Query: 248 KPTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSNPLTP 307
KPT P A+ +R R+T++ L+ AL A L+HSS P
Sbjct: 451 KPTTNYPVPAMTKR----CRLTKAL-------------LDPHTALYTPACLRHSSKTPPP 323
Query: 308 IPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEP 367
P PA PP SP SP PSP P S P P SS
Sbjct: 322 PP---PAPPQIPPPSSPPSPSSPPSP------------------PASAP*TPSSP*SSNS 206
Query: 368 IIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPP 423
P+ + P P S + S+ ++D +TA T +S D+PP
Sbjct: 205 ASSPPPQASPPPPTPTVSR*TSTATSSSSQTNSSDQTRATA-TRLSQTRCTTDAPP 41
>TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, partial
(68%)
Length = 1097
Score = 39.3 bits (90), Expect = 0.002
Identities = 32/120 (26%), Positives = 51/120 (41%), Gaps = 8/120 (6%)
Frame = +1
Query: 311 SQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNP--------TSRSEDPTSLLTIPYDP 362
S P +++T+PP S + P+ P + P T +S++ TSL T P P
Sbjct: 220 SPPESRSTTPPSSEPTCLLAPASPPTPKSSHSSTPPPSRSSSTTPKSKNATSL-TAPSCP 396
Query: 363 VSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSP 422
+ P P P++T P P T+ + T + SS +P + PI+ SP
Sbjct: 397 PPASPAA-TAPAPSKTPPNPPTNSSKPSSCKSSLPNTTPSSLSSAPPSPTTSPISTPSSP 573
>BP043764
Length = 479
Score = 38.9 bits (89), Expect = 0.003
Identities = 34/112 (30%), Positives = 49/112 (43%), Gaps = 11/112 (9%)
Frame = +3
Query: 286 LNLFDALPISALLQHS---SNPLTPIPES--------QPAAQTTSPPHSPRSPFFQPSPT 334
L L + L+H+ S+ L P P+S +P TT+PP +P+ P +P P
Sbjct: 48 LTLIHSPQTPLFLRHTTTVSSSLKPPPDSNNSAAPFRRPLKTTTTPPSNPK-PTPKPKPK 224
Query: 335 EAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSD 386
AP N LKNP S T+ L + +S P P P P R ++
Sbjct: 225 PAPTSNPLKNPFKSSHALTNKLWLT-SKLSPPPPPPPPPPPPPLPPPERETE 377
>TC9555 similar to UP|Q9MUE2 (Q9MUE2) S-adenosyl-L-methionine
Mg-protoporphyrin IX methyltranserase , partial (34%)
Length = 586
Score = 38.9 bits (89), Expect = 0.003
Identities = 30/94 (31%), Positives = 41/94 (42%), Gaps = 12/94 (12%)
Frame = +2
Query: 317 TTSPPHSPRSPFFQPSPTEAPLWNLLKN---PTSR---SEDPTSLLTIPYDPVSSE---- 366
+TS PHS R P Q + P W + P+S +E SL ++P P+ +
Sbjct: 65 STSTPHSHR-PVTQTGTKKKPKWRSHRR*DLPSSLQILTEPTVSLDSLPLPPIPNSHSQP 241
Query: 367 --PIIHEQPEPNQTEPQPRTSDHSVPRASARPAA 398
P H QP+P T P T HS+ A A A
Sbjct: 242 HSPFPHSQPQPPPTSPAQSTGPHSL*SAEASSPA 343
Score = 31.2 bits (69), Expect = 0.65
Identities = 14/34 (41%), Positives = 18/34 (52%)
Frame = +2
Query: 290 DALPISALLQHSSNPLTPIPESQPAAQTTSPPHS 323
D+LP+ + S P +P P SQP TSP S
Sbjct: 197 DSLPLPPIPNSHSQPHSPFPHSQPQPPPTSPAQS 298
>TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase type 2C,
complete
Length = 1325
Score = 38.9 bits (89), Expect = 0.003
Identities = 43/174 (24%), Positives = 64/174 (36%), Gaps = 9/174 (5%)
Frame = -3
Query: 300 HSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEA--------PLWNLLKNPTSRSED 351
H P P E+ + SPP P S QP P +A P L ++ +
Sbjct: 657 HYPPPHRPYYENTLPSLPPSPPPPPHSRTSQPPPPQADTAAPGTPPPPELRRDRQKQRSS 478
Query: 352 PTSLLTI-PYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFT 410
TS T+ P P P P+PN+T + R ASA T S+ T
Sbjct: 477 RTSSRTLSPRSPRPPSPSCSPPPKPNRTAKRRRL-------ASAPDPPPMMMTTKSSPGT 319
Query: 411 PVSFPINVIDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPER 464
+S + S PS T E++R+ ++ E+ + P P+R
Sbjct: 318 VISAQTHESPSAPSPTEENLRRRR*SSPGSCTSFAEFSASSGEHSSTPSRSPQR 157
>AW719831
Length = 436
Score = 38.5 bits (88), Expect = 0.004
Identities = 42/146 (28%), Positives = 62/146 (41%), Gaps = 1/146 (0%)
Frame = +1
Query: 286 LNLFDALPISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEA-PLWNLLKN 344
+N LP+ L Q+ ++ +PIP Q TT+ P+P A PL L
Sbjct: 4 VNNHTPLPLKILHQNPNSQKSPIPFHQQWRTTTTTT--------LPTPILAIPL--LAPA 153
Query: 345 PTSRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTD 404
P + PT T P S+ P + P P+ R+S + A + PA TT +
Sbjct: 154 PM*STPPPTMTPTTTTTPPSTNPHLLLPPPPH-----ARSSSCAATAAKSSPAPTTTTSP 318
Query: 405 SSTAFTPVSFPINVIDSPPSNTSESI 430
+S A TP S P S P ++ S+
Sbjct: 319 TSPA-TPRSSPSTASSSSPPSSPSSL 393
>TC8946 homologue to GB|AAM66972.1|21617922|AY088650 pyridoxine
biosynthesis protein-like {Arabidopsis thaliana;} ,
partial (45%)
Length = 503
Score = 38.5 bits (88), Expect = 0.004
Identities = 33/134 (24%), Positives = 60/134 (44%), Gaps = 16/134 (11%)
Frame = +2
Query: 294 ISALLQHSSNPLTPIPESQPAAQ--------------TTSPPHSPRSPFFQPSPTEAPLW 339
+ + L ++ P +P P + P+ T+S P+ P SP P AP W
Sbjct: 92 LESSLSTATAPQSPRPRNPPSPSKSASLRCSAAASSWTSSTPNKPASP---KKPAPAPSW 262
Query: 340 NLLKNPTSRSEDPTSLL--TIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPA 397
+P + + SL T+ S++P P+ + P+P ++ S PR S++P+
Sbjct: 263 PSSVSPPTSAPKAASLA*ATLSSSRKSNKP------SPSPSWPKPASAISSKPR-SSKPS 421
Query: 398 ARTTDTDSSTAFTP 411
A T T + ++ +P
Sbjct: 422 ASITSTRARSSLSP 463
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 38.5 bits (88), Expect = 0.004
Identities = 52/225 (23%), Positives = 65/225 (28%), Gaps = 2/225 (0%)
Frame = +1
Query: 320 PPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSS--EPIIHEQPEPNQ 377
PP P+ P++ SP P + P S S P P P S P ++ P P
Sbjct: 31 PPPVPK-PYYYKSPPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSHSPPPPYYYKSPPPPS 207
Query: 378 TEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSESIRKFMEVR 437
P P S P S P S +P+ P SPP TS
Sbjct: 208 PSPPPPYYYKSPPPPSPSPPP-PYYYKSPPPPSPIPHPPYYYKSPPPPTS---------- 354
Query: 438 KEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEADPLVQQAHPVPDQQEPI 497
S Y+ P P P P P P P + P+
Sbjct: 355 ----SPPPPYHYVSPPPPSPSPPPPYHYASPPPP---------SPSPAPTYIYKSPPPPV 495
Query: 498 QPDPEPEQSVSNQSSVRSPHPLVETSDHHRGTSEPNAPMINIGSP 542
+ P P S SP P T P P+ SP
Sbjct: 496 KLPPPPYHYTSPPPPSPSPAPTYIYKSPPPPTKSPPPPVYIYASP 630
>TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%)
Length = 1036
Score = 38.1 bits (87), Expect = 0.005
Identities = 40/139 (28%), Positives = 52/139 (36%), Gaps = 3/139 (2%)
Frame = +3
Query: 247 VKPTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSNPLT 306
+ P+ PAAA + + + SS+ SS LAS S+ P T
Sbjct: 153 LNPSTAAPAAAASAPSASG*SSSSSSSSSSSAVLASPST--------------SSTAPTT 290
Query: 307 PIPESQPAAQ--TTSPPHSPRSP-FFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPV 363
P S P+ +TSPP P P PSP + P S PT L PY P
Sbjct: 291 PPSPSPPSNSPTSTSPPPQPSPPNSTSPSPPQTP-------TRKTSHSPTYPLPSPYSPA 449
Query: 364 SSEPIIHEQPEPNQTEPQP 382
+S + TEP P
Sbjct: 450 TS---------TSATEPYP 479
Score = 30.8 bits (68), Expect = 0.85
Identities = 51/216 (23%), Positives = 73/216 (33%)
Frame = +3
Query: 308 IPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEP 367
+P P QT PP + SP +P+ T PT+LLT +P ++
Sbjct: 48 LPPKPPPPQTAPPPPTQLSPPQRPNSTA----------------PTALLT-ALNPSTA-- 170
Query: 368 IIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTS 427
P + P S +S+ A + + SSTA T P SPPSN+
Sbjct: 171 ----APAAAASAPSASG*SSSSSSSSSSSAVLASPSTSSTAPTTPPSP-----SPPSNSP 323
Query: 428 ESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEADPLVQQA 487
S T P P+ P P +P P + +
Sbjct: 324 TS--------------------TSPPPQ------------PSPPNSTSPSPPQTPTRKTS 407
Query: 488 HPVPDQQEPIQPDPEPEQSVSNQSSVRSPHPLVETS 523
H P P P P S + +S P+PL T+
Sbjct: 408 H------SPTYPLPSP-YSPATSTSATEPYPLSITA 494
>TC13980 similar to UP|Q9SGT0 (Q9SGT0) T6H22.18 protein (F14J16.33), partial
(12%)
Length = 565
Score = 38.1 bits (87), Expect = 0.005
Identities = 41/140 (29%), Positives = 54/140 (38%)
Frame = +2
Query: 272 SAHSSKPALASDDDLNLFDALPISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQP 331
S HS P + DL L P S + IP + P + + S SP SP
Sbjct: 179 SPHSQFPETHTVHDLELQWLPPASTTPSATRASSLGIP-NPPTSSSASKTPSPCSP---- 343
Query: 332 SPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPR 391
PT + +PT S TS P+ P +P P P P T+ S PR
Sbjct: 344 -PTSS-------SPTGSSSHSTS--NPPHPPPQQQPF---SPPPPPPPPPSPTAPSSHPR 484
Query: 392 ASARPAARTTDTDSSTAFTP 411
A PAA + S ++ TP
Sbjct: 485 LPAAPAAGRISSASKSSTTP 544
>TC13802 similar to UP|Q9M1T1 (Q9M1T1) Sugar-phosphate isomerase-like
protein, partial (41%)
Length = 522
Score = 37.7 bits (86), Expect = 0.007
Identities = 34/128 (26%), Positives = 51/128 (39%)
Frame = +2
Query: 300 HSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIP 359
HS+ P S +TTS SP S +PSP+ AP + L+ P+S + +
Sbjct: 65 HSTLTKPPSQISSNPNKTTSTTSSPTSTTPKPSPSPAPS-STLRAPSSSPASASPASSPT 241
Query: 360 YDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVI 419
P S P P P TS+ S S+ +A S +A + + + +
Sbjct: 242 KSPKPSSPSASAPPSSTPLTPSTATSESSPTATSSSSSASPAPPTSFSALSR-ALELKAL 418
Query: 420 DSPPSNTS 427
S PS S
Sbjct: 419 ASSPSPPS 442
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.128 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,047,375
Number of Sequences: 28460
Number of extensions: 219057
Number of successful extensions: 2553
Number of sequences better than 10.0: 315
Number of HSP's better than 10.0 without gapping: 2099
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2416
length of query: 766
length of database: 4,897,600
effective HSP length: 97
effective length of query: 669
effective length of database: 2,136,980
effective search space: 1429639620
effective search space used: 1429639620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0014.15


