

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
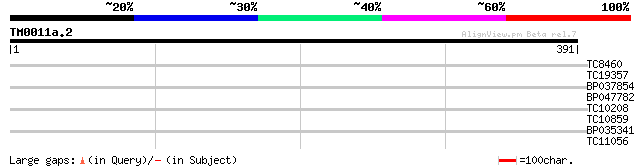
Query= TM0011a.2
(391 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F... 38 0.003
TC19357 weakly similar to UP|Q9FHY5 (Q9FHY5) Emb|CAB72466.1 (At5... 36 0.013
BP037854 28 2.6
BP047782 28 2.6
TC10208 weakly similar to UP|Q8TUQ7 (Q8TUQ7) TraB family protein... 28 3.4
TC10859 27 5.8
BP035341 27 7.6
TC11056 similar to UP|O81390 (O81390) Calcium-dependent protein ... 26 10.0
>TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;}, partial (36%)
Length = 880
Score = 38.1 bits (87), Expect = 0.003
Identities = 34/132 (25%), Positives = 54/132 (40%), Gaps = 1/132 (0%)
Frame = +3
Query: 29 TNPVYTEEQFQRRYRMRKHMFLRIVEAISNTDDYFQMRNDATGRMGLSPLQKQTAAIRML 88
++P + EE+F+R +RM K F E I D + + R + Q+ I L
Sbjct: 357 SHPDFPEEEFRRSFRMSKATF----EMICRELDSAVTKKNTMLREAIPVRQRVAVCIWRL 524
Query: 89 AYGSSTDSVDEYVCIGESTARKCMARFVKGINEVFGPEYLRKTNNKDI-ARLMELGKARG 147
A G V + +G ST K + I V P++LR + + A E G
Sbjct: 525 ATGDPLRLVAKRFGLGISTCHKLVLEVCSAIKTVLMPKFLRWPDEAAMTAAKSEFEALSG 704
Query: 148 FPGMLGSIDCMH 159
P + G++ H
Sbjct: 705 IPNIGGAMYTTH 740
>TC19357 weakly similar to UP|Q9FHY5 (Q9FHY5) Emb|CAB72466.1 (At5g41980),
partial (16%)
Length = 577
Score = 35.8 bits (81), Expect = 0.013
Identities = 21/60 (35%), Positives = 34/60 (56%)
Frame = +2
Query: 267 RKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDER 326
++LF R + R+F VL +RF I+ PA + I +DI+ A +LHN+I ++R
Sbjct: 5 KELFNHRHYFLRGAILRSFTVLKARFPILI-PAPQYSFQIQRDIVIAACVLHNIIRREDR 181
>BP037854
Length = 462
Score = 28.1 bits (61), Expect = 2.6
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -1
Query: 183 IILEAIASQDLWIWHAFFGVA 203
+I ++ + DLW W FFG A
Sbjct: 282 LIFSSVLALDLWFWELFFGTA 220
>BP047782
Length = 479
Score = 28.1 bits (61), Expect = 2.6
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = +3
Query: 89 AYGSSTDSVDEYVCIGESTARKCMA 113
AY S +DS+ YV +GES ++ MA
Sbjct: 105 AYLSLSDSISSYVSVGESLCKQLMA 179
>TC10208 weakly similar to UP|Q8TUQ7 (Q8TUQ7) TraB family protein, partial
(4%)
Length = 566
Score = 27.7 bits (60), Expect = 3.4
Identities = 11/29 (37%), Positives = 19/29 (64%)
Frame = +1
Query: 320 MIVEDEREMCDGNIDFSYDQLENDASTPE 348
+I+ ERE+ + +DFS+ +N S+PE
Sbjct: 304 IILTSEREL*ERLVDFSFSAFQNSMSSPE 390
>TC10859
Length = 533
Score = 26.9 bits (58), Expect = 5.8
Identities = 10/30 (33%), Positives = 19/30 (63%)
Frame = +3
Query: 320 MIVEDEREMCDGNIDFSYDQLENDASTPEV 349
+I+ ERE+ + +DF + +N S+PE+
Sbjct: 342 VILSSEREL*ERLVDFRFSSFQNSMSSPEL 431
>BP035341
Length = 523
Score = 26.6 bits (57), Expect = 7.6
Identities = 14/47 (29%), Positives = 25/47 (52%)
Frame = -3
Query: 219 NDVLQGKAPEVQFTLNGTTYNMGYYLADEIYPEWATFVKTISMPQGE 265
NDV +A NG+T +MGY+L + ++P A +++ G+
Sbjct: 479 NDVFASQAH------NGSTDSMGYFLGEVLFPYHAESEVELNLSAGD 357
>TC11056 similar to UP|O81390 (O81390) Calcium-dependent protein kinase ,
partial (22%)
Length = 675
Score = 26.2 bits (56), Expect = 10.0
Identities = 14/51 (27%), Positives = 23/51 (44%)
Frame = -2
Query: 290 SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDEREMCDGNIDFSYDQL 340
+ F IV P +H+ +Y C I H+M +C G++ F Y +
Sbjct: 347 TEFLIVYSPIIVC-IHLRNYFLYRCFITHSM-------LCHGSLQFIYSNM 219
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,664,113
Number of Sequences: 28460
Number of extensions: 88138
Number of successful extensions: 434
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 433
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 434
length of query: 391
length of database: 4,897,600
effective HSP length: 92
effective length of query: 299
effective length of database: 2,279,280
effective search space: 681504720
effective search space used: 681504720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0011a.2


