

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.6
(493 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
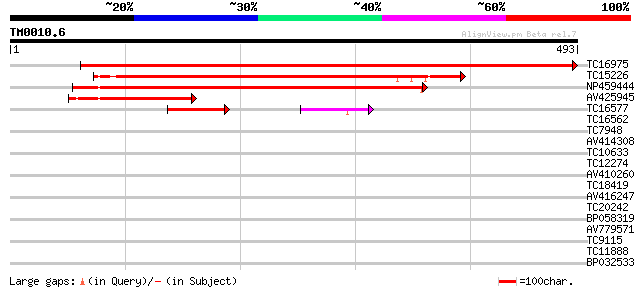
Score E
Sequences producing significant alignments: (bits) Value
TC16975 UP|O82458 (O82458) Rac GTPase activating protein 1, part... 852 0.0
TC15226 UP|O82459 (O82459) Rac GTPase activating protein 2 (Frag... 375 e-104
NP459444 rac GTPase activating protein 3 [Lotus japonicus] 356 5e-99
AV425945 137 5e-33
TC16577 similar to UP|O82458 (O82458) Rac GTPase activating prot... 91 6e-19
TC16562 similar to GB|AAM78073.1|21928089|AY125563 AT4g24660/F22... 32 0.31
TC7948 31 0.53
AV414308 29 1.5
TC10633 29 1.5
TC12274 similar to PIR|G88642|G88642 protein C54E4.4 [imported] ... 29 2.0
AV410260 28 2.6
TC18419 28 2.6
AV416247 28 3.4
TC20242 similar to UP|Q9LXK4 (Q9LXK4) Guanine nucleotide-exchang... 28 3.4
BP058319 28 4.5
AV779571 28 4.5
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 27 7.6
TC11888 similar to UP|O22636 (O22636) Poly(A) polymerase (Pepti... 27 9.9
BP032533 27 9.9
>TC16975 UP|O82458 (O82458) Rac GTPase activating protein 1, partial (88%)
Length = 1686
Score = 852 bits (2201), Expect = 0.0
Identities = 429/432 (99%), Positives = 430/432 (99%)
Frame = +1
Query: 62 LSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLP 121
LSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLP
Sbjct: 1 LSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLP 180
Query: 122 VEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIF 181
VEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIF
Sbjct: 181 VEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIF 360
Query: 182 RINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSE 241
RINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSE
Sbjct: 361 RINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSE 540
Query: 242 EECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTAL 301
EECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTAL
Sbjct: 541 EECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTAL 720
Query: 302 MYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDC 361
MYAVQVM+FLKTLVVKTLR REESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDC
Sbjct: 721 MYAVQVMNFLKTLVVKTLRVREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDC 900
Query: 362 SDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCNVVSQ 421
SDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCNVVSQ
Sbjct: 901 SDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCNVVSQ 1080
Query: 422 LCSFAIGLQDSSIATGQAKISRSKSLQMSTSDIDKSFKNVIEFPVVGPAEKNRGTAIIGR 481
LCSFAIGLQDSSIATGQAKISRSKSL MSTSDIDKSFKNVIEFPVVGPAEKNRGTAIIGR
Sbjct: 1081LCSFAIGLQDSSIATGQAKISRSKSLPMSTSDIDKSFKNVIEFPVVGPAEKNRGTAIIGR 1260
Query: 482 INSRTELTEAWR 493
INSRTELTEAWR
Sbjct: 1261INSRTELTEAWR 1296
>TC15226 UP|O82459 (O82459) Rac GTPase activating protein 2 (Fragment),
partial (92%)
Length = 1436
Score = 375 bits (962), Expect = e-104
Identities = 202/341 (59%), Positives = 246/341 (71%), Gaps = 18/341 (5%)
Frame = +3
Query: 74 KSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPP 133
KSL+ +CS D SS++I WP+ VRHV+HVTFDRF+GFLGLP E +PEVP++ P
Sbjct: 3 KSLV-TCSVDREDV-----SSLDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEVPQKVP 164
Query: 134 SASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELV 193
SAS VFGVS +SMQ S+D RGNSVPTILL+MQ LY+ GGL+AEGIFRINA+NSQEE V
Sbjct: 165 SASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFV 344
Query: 194 REQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPP 253
R QLNRG+VP GV+VHCL+GLIKAWFRELPTG+LD L+PE+VM SEE+C LV+LLP
Sbjct: 345 RCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPS 524
Query: 254 TEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKT 313
TEAALLDWAINLMADV + E FNKMNARNIAMVFAPNMT M DPLTAL++AVQVM+FLKT
Sbjct: 525 TEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKT 704
Query: 314 LVVKTLREREESIVKSNPVPNL----------NSFDDDGHQSDSQ------VLPKDGSEN 357
L++KTLRER+ES+ K+ + +L + F D+ +S +Q +P D SE
Sbjct: 705 LILKTLRERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQPVDTCATMPPDKSEF 884
Query: 358 GND--CSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSP 396
C DE V+ S E + G + S E+ P
Sbjct: 885 SRMEWCVDE-KVWSSEEKGTGGGALESVSGGSSPSRYESGP 1004
>NP459444 rac GTPase activating protein 3 [Lotus japonicus]
Length = 1300
Score = 356 bits (913), Expect = 5e-99
Identities = 186/313 (59%), Positives = 233/313 (74%), Gaps = 4/313 (1%)
Frame = +2
Query: 55 KDREGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRF 114
++ E +Q S L+A +KS++ +CS D + MEIGWP+NV+HV HVTFDRF
Sbjct: 2 EEAEQNQGSPAAFLLAALKKSMV-ACSVDSPDDVISAVHPMEIGWPTNVKHVTHVTFDRF 178
Query: 115 HGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGG 174
+GFLGLP+E E VP PSAS SVFGVS ESM S+D++GNSVPTILLLMQ LY++GG
Sbjct: 179 NGFLGLPLELEVHVPAPVPSASVSVFGVSAESMHCSYDSKGNSVPTILLLMQERLYSQGG 358
Query: 175 LQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEE 234
L AEGIFRIN EN QEE +R+QLNRGVVP+ +DVHCLAGLIKAWFRELP+G+LD LSPE+
Sbjct: 359 LMAEGIFRINPENGQEEHLRDQLNRGVVPDNIDVHCLAGLIKAWFRELPSGVLDGLSPEQ 538
Query: 235 VMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHM 294
V++ +EEE QLV+ L PTE ALL+WA++LMADV + E NKM+ARNIAMVFAPNMT M
Sbjct: 539 VLECNTEEEFVQLVKQLKPTELALLNWALDLMADVVEEEEHNKMDARNIAMVFAPNMTQM 718
Query: 295 ADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGH-QSDSQVLPKD 353
+DPLTALM+AVQVM+ LKTL++KTL EREE+ + +S D + DSQ+
Sbjct: 719 SDPLTALMHAVQVMNLLKTLILKTLSEREEATTAGYSSMSSHSSDRQSEDEYDSQLEMYT 898
Query: 354 GSE---NGNDCSD 363
+E + +DC D
Sbjct: 899 SAELRGSQSDCDD 937
>AV425945
Length = 415
Score = 137 bits (344), Expect = 5e-33
Identities = 68/111 (61%), Positives = 87/111 (78%)
Frame = +1
Query: 52 EKEKDREGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTF 111
E+E+DR+ +QLSL+ LL+A RKS++ +C D + MEIGWP++V+H+ HVTF
Sbjct: 88 EEEQDRQ-NQLSLVALLLAALRKSMV-ACRVDRPDEAISTVHQMEIGWPTDVQHITHVTF 261
Query: 112 DRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTIL 162
DRF+GFLGLPVEFE E+P R PSAS SVFGVS ESMQ S+D++GNSVPTIL
Sbjct: 262 DRFNGFLGLPVEFEVEIPGRVPSASVSVFGVSAESMQCSYDSKGNSVPTIL 414
>TC16577 similar to UP|O82458 (O82458) Rac GTPase activating protein 1,
partial (16%)
Length = 726
Score = 90.5 bits (223), Expect = 6e-19
Identities = 43/54 (79%), Positives = 49/54 (90%)
Frame = +1
Query: 138 SVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEE 191
++FGVSTESMQLS+D RGNSVPTILL MQRHLYA+GGLQAEGIFRIN +N+ E
Sbjct: 169 TMFGVSTESMQLSYDTRGNSVPTILLQMQRHLYAQGGLQAEGIFRINTDNTHTE 330
Score = 56.6 bits (135), Expect = 9e-09
Identities = 29/71 (40%), Positives = 43/71 (59%), Gaps = 8/71 (11%)
Frame = +1
Query: 254 TEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMT--------HMADPLTALMYAV 305
TEA+LLDWAINLM DV Q EH ++MNA NIAMV +++ H L ++ +
Sbjct: 325 TEASLLDWAINLMVDVVQEEHLSRMNAHNIAMVLVNSVSPGIVGRKMHTYTVLVLFIFVI 504
Query: 306 QVMSFLKTLVV 316
++ +K L++
Sbjct: 505 RLQIIVKILLM 537
>TC16562 similar to GB|AAM78073.1|21928089|AY125563 AT4g24660/F22K18_140
{Arabidopsis thaliana;}, partial (29%)
Length = 621
Score = 31.6 bits (70), Expect = 0.31
Identities = 12/25 (48%), Positives = 19/25 (76%)
Frame = +3
Query: 36 LQLQTHLDLVEEDEEEEKEKDREGD 60
++ H DL EE+EEEE+E++ EG+
Sbjct: 123 MEFDDHEDLEEEEEEEEEEEEEEGE 197
>TC7948
Length = 1230
Score = 30.8 bits (68), Expect = 0.53
Identities = 11/38 (28%), Positives = 20/38 (51%)
Frame = +1
Query: 369 VSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSG 406
++++P P+P H ++G ++ GSE P L G
Sbjct: 307 ITSQPPPPTPPQHVDEGVQSSQGSEEEGLPPRILLHRG 420
>AV414308
Length = 389
Score = 29.3 bits (64), Expect = 1.5
Identities = 24/91 (26%), Positives = 33/91 (35%)
Frame = +3
Query: 318 TLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPS 377
TL R+ S SNP P + Q S K+ + S P P
Sbjct: 57 TLPWRKNSKSSSNPTPTTGATP----QKKSTTSNKESLLQNPPSPPPEASPSSPGPGSPP 224
Query: 378 PTHHTEDGCETESGSETSPTPAENFLSSGSR 408
P HH + + + TSP P++ SS R
Sbjct: 225 PPHHAPSSSWSTATATTSPGPSKPPQSSSPR 317
>TC10633
Length = 593
Score = 29.3 bits (64), Expect = 1.5
Identities = 19/57 (33%), Positives = 27/57 (47%), Gaps = 1/57 (1%)
Frame = +1
Query: 323 EESIVKSNPVPNLNSFDDDG-HQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSP 378
E S + P++ F D + S S P+ ENG++ +D VFVS P P P
Sbjct: 148 EHSTTAAPESPDIFGFSDPNPNYSQSPFEPEHVVENGDENGYDDGVFVSDGPVLPPP 318
>TC12274 similar to PIR|G88642|G88642 protein C54E4.4 [imported] -
Caenorhabditis elegans {Caenorhabditis elegans;} ,
partial (11%)
Length = 589
Score = 28.9 bits (63), Expect = 2.0
Identities = 24/116 (20%), Positives = 49/116 (41%), Gaps = 6/116 (5%)
Frame = +3
Query: 333 PNLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAE------PSQPSPTHHTEDGC 386
P + + DDD H+SD +V+ K+ +D D ++S E PS + TH
Sbjct: 213 PEVLNEDDDNHESD-EVIAKNDKYGKSDVLSSDVNYISNELLDNQTPSGLNITHEGSAAW 389
Query: 387 ETESGSETSPTPAENFLSSGSRLLIDSCPCNVVSQLCSFAIGLQDSSIATGQAKIS 442
+ + + P + F ++ + ++ S + F I L D + ++ ++
Sbjct: 390 TELTSASATEVPEKTF---HENIIENVLEFSLASNVVDFEIPLLDVKFISQRSPVT 548
>AV410260
Length = 373
Score = 28.5 bits (62), Expect = 2.6
Identities = 18/59 (30%), Positives = 25/59 (41%)
Frame = +2
Query: 347 SQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSS 405
S PK S + T S+ PS PSPT + + + S +P P + LSS
Sbjct: 185 SNASPKTNSSTSSKTPSSATP-TSSSPSAPSPTPTSRSASFSSAASAGTPPPTASALSS 358
>TC18419
Length = 409
Score = 28.5 bits (62), Expect = 2.6
Identities = 17/45 (37%), Positives = 20/45 (43%)
Frame = +1
Query: 363 DEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGS 407
D+D V P PS +DG E + PTPA NF S S
Sbjct: 253 DDDGPLVGPPPPPPSEQQPDDDG---EMFGPSPPTPASNFNDSDS 378
>AV416247
Length = 417
Score = 28.1 bits (61), Expect = 3.4
Identities = 27/96 (28%), Positives = 43/96 (44%), Gaps = 6/96 (6%)
Frame = +1
Query: 17 DGSHQTALINNSSSVEEGVLQLQTH-LDLVEEDEEEEKEKDREGDQLSLLTLLIATFRKS 75
D + A+ N S E+ + + LDL +++EE EKE +E Q + K
Sbjct: 142 DPIDEVAIQNLKSYKEKNFVDISKEDLDLGDKNEEREKEVKQEFAQ-------TCDWIKK 300
Query: 76 LIGSCSTSPRDSGALSSS-----SMEIGWPSNVRHV 106
+G S + S LSSS S + GW +N+ +
Sbjct: 301 RLGDKVASVQISNRLSSSPCVLVSGKFGWSANMERL 408
>TC20242 similar to UP|Q9LXK4 (Q9LXK4) Guanine nucleotide-exchange-like
protein, partial (5%)
Length = 741
Score = 28.1 bits (61), Expect = 3.4
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +1
Query: 356 ENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSP 396
+N D + E + +EP+Q +P+ + ETE G+ T P
Sbjct: 400 QNFTDITKEASQRKQSEPNQAAPSPESASANETEDGAVTKP 522
>BP058319
Length = 518
Score = 27.7 bits (60), Expect = 4.5
Identities = 10/18 (55%), Positives = 16/18 (88%)
Frame = -2
Query: 95 MEIGWPSNVRHVAHVTFD 112
+EIG P++V+HVAH+ +D
Sbjct: 415 IEIGCPTDVQHVAHIGWD 362
>AV779571
Length = 538
Score = 27.7 bits (60), Expect = 4.5
Identities = 26/93 (27%), Positives = 38/93 (39%)
Frame = +1
Query: 334 NLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSE 393
N N Q +Q D + GN S FVSA Q SPT + +S S+
Sbjct: 256 NHNQQQQQQQQQQTQQQVVDNQQIGNAGSTMSQ-FVSAPQPQSSPTQAQSSLSQQQSFSD 432
Query: 394 TSPTPAENFLSSGSRLLIDSCPCNVVSQLCSFA 426
++ P +S L+ S P + SQL + +
Sbjct: 433 SNVKPVTTIVSP-LHSLMGSFPQDETSQLLNLS 528
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 26.9 bits (58), Expect = 7.6
Identities = 15/54 (27%), Positives = 22/54 (39%), Gaps = 6/54 (11%)
Frame = -2
Query: 352 KDGSENGNDCSDEDTVFVSAEPSQPSPT------HHTEDGCETESGSETSPTPA 399
K ++G CS + T + PS P + HH + CE P+PA
Sbjct: 849 KANPQHGTKCSPKSTSPPHSPPSSPPQSSPPPSPHHAKPLCEHTQRQHQQPSPA 688
>TC11888 similar to UP|O22636 (O22636) Poly(A) polymerase (Peptidyl-prolyl
cis-trans isomerase) (PPIase) (Rotamase) , partial (57%)
Length = 931
Score = 26.6 bits (57), Expect = 9.9
Identities = 14/36 (38%), Positives = 19/36 (51%), Gaps = 1/36 (2%)
Frame = -1
Query: 13 RCSNDGSHQTALINNSSSVEEGVLQL-QTHLDLVEE 47
R + GSHQ L+ + +EE + L Q HLD E
Sbjct: 733 RAPHMGSHQPCLLQHLQHIEEALFVLVQQHLDFYPE 626
>BP032533
Length = 441
Score = 26.6 bits (57), Expect = 9.9
Identities = 16/55 (29%), Positives = 25/55 (45%)
Frame = +3
Query: 353 DGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGS 407
DGS+N + V V+ ++P+P T S S +S +P + SS S
Sbjct: 66 DGSQNQK*RTQTTPVAVAVAIAEPNPLLRTSSNSSATSASHSSSSPFSSSPSSPS 230
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.130 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,989,559
Number of Sequences: 28460
Number of extensions: 109097
Number of successful extensions: 643
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 617
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 635
length of query: 493
length of database: 4,897,600
effective HSP length: 94
effective length of query: 399
effective length of database: 2,222,360
effective search space: 886721640
effective search space used: 886721640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0010.6


