

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.3.5
(80 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
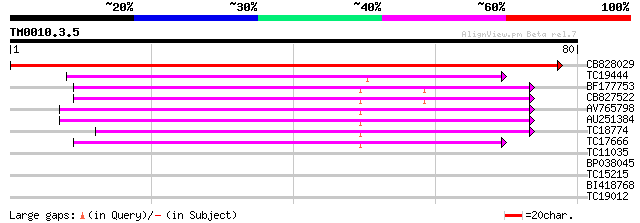
CB828029 152 1e-38
TC19444 weakly similar to UP|LSM7_HUMAN (Q9UK45) U6 snRNA-associ... 58 4e-10
BF177753 48 3e-07
CB827522 46 1e-06
AV765798 45 2e-06
AU251384 44 5e-06
TC18774 similar to GB|AAM16215.1|20334918|AY094059 At1g65700/F1E... 40 6e-05
TC17666 similar to GB|AAP37664.1|30725284|BT008305 At2g43810 {Ar... 39 2e-04
TC11035 homologue to UP|Q9C6K5 (Q9C6K5) Sm-like protein, partial... 36 0.001
BP038045 24 5.8
TC15215 similar to UP|CYP4_ARATH (P34791) Peptidyl-prolyl cis-tr... 23 7.5
BI418768 23 9.8
TC19012 23 9.8
>CB828029
Length = 502
Score = 152 bits (384), Expect = 1e-38
Identities = 73/78 (93%), Positives = 78/78 (99%)
Frame = +2
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDKNLQIKLNANRMI+GTLRG+DQFMN+V+DNTVEVNGNEKNDIGMV
Sbjct: 56 MSRSGQPPDLKKYMDKNLQIKLNANRMIIGTLRGFDQFMNLVIDNTVEVNGNEKNDIGMV 235
Query: 61 VIRGNSVVTVEALEPVTR 78
VIRGNSVVTVEALEPVT+
Sbjct: 236 VIRGNSVVTVEALEPVTK 289
>TC19444 weakly similar to UP|LSM7_HUMAN (Q9UK45) U6 snRNA-associated
Sm-like protein LSm7, partial (88%)
Length = 565
Score = 57.8 bits (138), Expect = 4e-10
Identities = 27/71 (38%), Positives = 42/71 (59%), Gaps = 9/71 (12%)
Frame = +3
Query: 9 DLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV---------NGNEKNDIGM 59
DL K++DK +Q+KL R + GTL+GYDQ +N+V+D VE + +G+
Sbjct: 138 DLAKFVDKGVQVKLTGGRQVTGTLKGYDQLLNLVLDEAVEFLRDPDDPLKTTAQTRSLGL 317
Query: 60 VVIRGNSVVTV 70
+V RG +V+ V
Sbjct: 318 IVCRGTAVMLV 350
>BF177753
Length = 491
Score = 48.1 bits (113), Expect = 3e-07
Identities = 28/69 (40%), Positives = 41/69 (58%), Gaps = 4/69 (5%)
Frame = +3
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE--VNGNEKNDI--GMVVIRGN 65
L Y+DK L + L R ++GTLR +DQF N V++ E + G+ DI G+ VIRG
Sbjct: 102 LASYLDKKLLVLLRDGRKLMGTLRSFDQFANAVLEGACERVIVGDLYCDIPLGLYVIRGE 281
Query: 66 SVVTVEALE 74
+VV + L+
Sbjct: 282 NVVLIGELD 308
>CB827522
Length = 519
Score = 45.8 bits (107), Expect = 1e-06
Identities = 26/69 (37%), Positives = 41/69 (58%), Gaps = 4/69 (5%)
Frame = +3
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE--VNGNEKNDI--GMVVIRGN 65
L Y+DK L + L R ++G LR +DQF N+V++ E + G+ D+ G+ VIRG
Sbjct: 177 LATYLDKKLLVLLRDGRKLLGLLRSFDQFANVVLEGACERVIVGDLYCDVPLGLYVIRGE 356
Query: 66 SVVTVEALE 74
+VV + L+
Sbjct: 357 NVVLIGELD 383
>AV765798
Length = 566
Score = 45.1 bits (105), Expect = 2e-06
Identities = 23/72 (31%), Positives = 42/72 (57%), Gaps = 5/72 (6%)
Frame = +1
Query: 8 PDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-----VNGNEKNDIGMVVI 62
P L+ +D+ + + N R IVGTL+G+DQ N+++D + E G + +G+ +I
Sbjct: 205 PGLESLVDQQISVITNDGRNIVGTLKGFDQATNIILDESHERVFSTKEGVQLIALGLYII 384
Query: 63 RGNSVVTVEALE 74
RG+++ V L+
Sbjct: 385 RGDNISVVGELD 420
>AU251384
Length = 364
Score = 43.9 bits (102), Expect = 5e-06
Identities = 22/72 (30%), Positives = 42/72 (57%), Gaps = 5/72 (6%)
Frame = +2
Query: 8 PDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-----VNGNEKNDIGMVVI 62
P L+ +D+ + + N R IVG L+G+DQ N+++D + E G ++ +G+ +I
Sbjct: 149 PGLESLVDQQISVITNDGRNIVGVLKGFDQATNIILDESHERVFSTKEGVQQIVLGLYII 328
Query: 63 RGNSVVTVEALE 74
RG+++ V L+
Sbjct: 329 RGDNISVVGELD 364
>TC18774 similar to GB|AAM16215.1|20334918|AY094059 At1g65700/F1E22_3
{Arabidopsis thaliana;}, partial (86%)
Length = 572
Score = 40.4 bits (93), Expect = 6e-05
Identities = 21/67 (31%), Positives = 38/67 (56%), Gaps = 5/67 (7%)
Frame = -1
Query: 13 YMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-----VNGNEKNDIGMVVIRGNSV 67
Y + + + N R IVGTL+G+DQ N+++D + E G + +G+ +IRG+++
Sbjct: 470 YFAEQISVITNDGRNIVGTLKGFDQATNIILDESHERVFSTKEGVQLIALGLYIIRGDNI 291
Query: 68 VTVEALE 74
V L+
Sbjct: 290 SVVGELD 270
>TC17666 similar to GB|AAP37664.1|30725284|BT008305 At2g43810 {Arabidopsis
thaliana;}, partial (95%)
Length = 770
Score = 38.5 bits (88), Expect = 2e-04
Identities = 23/62 (37%), Positives = 34/62 (54%), Gaps = 1/62 (1%)
Frame = +3
Query: 10 LKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVE-VNGNEKNDIGMVVIRGNSVV 68
LK + + +KLN+ G L D +MN+ ++ T E VNG KN G IRGN+V+
Sbjct: 186 LKSIRGRPVVVKLNSGVDYRGILACLDGYMNIAMEQTEEYVNGQLKNKYGDAFIRGNNVL 365
Query: 69 TV 70
+
Sbjct: 366 YI 371
>TC11035 homologue to UP|Q9C6K5 (Q9C6K5) Sm-like protein, partial (96%)
Length = 576
Score = 36.2 bits (82), Expect = 0.001
Identities = 25/90 (27%), Positives = 46/90 (50%), Gaps = 15/90 (16%)
Frame = +2
Query: 6 QPPDLKKY-MDKNLQIKLNANRMIVGTLRGYDQFMNMVVDN------TVEVNG------- 51
+P DL + +D+ + +KL ++R + G L YDQ +NM++ + TVE++
Sbjct: 44 EPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEEIV 223
Query: 52 -NEKNDIGMVVIRGNSVVTVEALEPVTRTS 80
K + + +RG+ V+ V P RT+
Sbjct: 224 RTTKRTVPFLFVRGDGVILV---SPPLRTA 304
>BP038045
Length = 556
Score = 23.9 bits (50), Expect = 5.8
Identities = 9/19 (47%), Positives = 14/19 (73%)
Frame = +1
Query: 28 IVGTLRGYDQFMNMVVDNT 46
I GTL DQ++N+ ++NT
Sbjct: 103 IRGTLHSVDQYLNIKLENT 159
>TC15215 similar to UP|CYP4_ARATH (P34791) Peptidyl-prolyl cis-trans
isomerase, chloroplast precursor (PPIase) (Rotamase)
(Cyclophilin) (Cyclosporin A-binding protein) , partial
(45%)
Length = 654
Score = 23.5 bits (49), Expect = 7.5
Identities = 20/71 (28%), Positives = 31/71 (43%), Gaps = 5/71 (7%)
Frame = -3
Query: 10 LKKYMDKNL-----QIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMVVIRG 64
L+ Y K L K+N+ R+ TLRG +Q + +N V + G +++R
Sbjct: 520 LRSYSHKTLLKTGFDFKINSKRISEHTLRGDNQNIGNCTENQKNVIM*CTSA*GSIILRQ 341
Query: 65 NSVVTVEALEP 75
S V A P
Sbjct: 340 FSTVNNSARLP 308
>BI418768
Length = 546
Score = 23.1 bits (48), Expect = 9.8
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = +1
Query: 14 MDKNLQIKLNANRMIVGTLRGYDQFMNMVV 43
+D++ K N +IVG LR FM MVV
Sbjct: 280 LDQHSI*KGQCNALIVGRLRRVSGFMPMVV 369
>TC19012
Length = 274
Score = 23.1 bits (48), Expect = 9.8
Identities = 10/18 (55%), Positives = 12/18 (66%)
Frame = -1
Query: 46 TVEVNGNEKNDIGMVVIR 63
TVE N EK D+G+ V R
Sbjct: 217 TVESNRGEKRDLGVSVFR 164
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.133 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 829,035
Number of Sequences: 28460
Number of extensions: 5343
Number of successful extensions: 28
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 27
length of query: 80
length of database: 4,897,600
effective HSP length: 56
effective length of query: 24
effective length of database: 3,303,840
effective search space: 79292160
effective search space used: 79292160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0010.3.5


