

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.16
(405 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
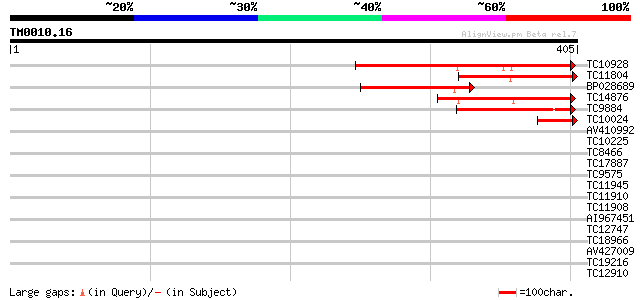
Score E
Sequences producing significant alignments: (bits) Value
TC10928 weakly similar to PIR|G96668|G96668 protein F1N19.7 [imp... 172 1e-43
TC11804 similar to GB|AAO63413.1|28950979|BT005349 At5g01420 {Ar... 102 9e-23
BP028689 100 7e-22
TC14876 weakly similar to UP|Q9ZVB8 (Q9ZVB8) At2g41330 protein (... 96 1e-20
TC9884 weakly similar to GB|AAO63337.1|28950827|BT005273 At5g585... 74 3e-14
TC10024 weakly similar to UP|Q8H5B5 (Q8H5B5) OJ1477_F01.28 prote... 41 4e-04
AV410992 35 0.017
TC10225 similar to UP|Q8LFQ6 (Q8LFQ6) Glutaredoxin, partial (69%) 34 0.038
TC8466 29 1.2
TC17887 weakly similar to UP|AAQ91191 (AAQ91191) 1-O-sinapoylglu... 29 1.6
TC9575 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase,... 28 2.1
TC11945 28 2.1
TC11910 similar to GB|CAA54498.1|602891|SCIILDNA YBL0422 {Saccha... 28 3.6
TC11908 similar to PIR|T46895|T46895 acyl-CoA dehydrogenase [va... 27 4.7
AI967451 27 6.1
TC12747 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-l... 27 6.1
TC18966 similar to UP|Q93W26 (Q93W26) AT5g53860/K19P17_2, partia... 27 8.0
AV427009 27 8.0
TC19216 similar to UP|MCPI_SOLTU (P01075) Metallocarboxypeptidas... 27 8.0
TC12910 similar to UP|Q9XEL2 (Q9XEL2) Monodehydroascorbate reduc... 27 8.0
>TC10928 weakly similar to PIR|G96668|G96668 protein F1N19.7 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 590
Score = 172 bits (435), Expect = 1e-43
Identities = 80/165 (48%), Positives = 118/165 (71%), Gaps = 8/165 (4%)
Frame = +3
Query: 248 IQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKE 307
+ F +K PPGG ++VVIYTT+LRG+RKTFEDCN+ R ++E + V + ERDV++ SGF +
Sbjct: 51 LSQFPEKSPPGGADAVVIYTTSLRGVRKTFEDCNRAREVLEGHRVVVDERDVALHSGFLK 230
Query: 308 ELRKLMGAEQV----KVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEG--IPKAVG--GC 359
E+++L+ E V +P VFVKGR +GG++E+V+L + +LG +L + + VG GC
Sbjct: 231 EVKELLAEEVVVAVAPLPRVFVKGRYLGGLEELVELNESGRLGRILNATRVERGVGRYGC 410
Query: 360 EGCGGVRFVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
+GCGG RFV C++C GSCK++ D + +C KCNENG++ CP C
Sbjct: 411 DGCGGARFVPCLDCGGSCKLVLSDVNQIERCPKCNENGLVHCPAC 545
>TC11804 similar to GB|AAO63413.1|28950979|BT005349 At5g01420 {Arabidopsis
thaliana;}, partial (12%)
Length = 548
Score = 102 bits (255), Expect = 9e-23
Identities = 46/88 (52%), Positives = 64/88 (72%), Gaps = 3/88 (3%)
Frame = +3
Query: 321 PVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKAV--GGCEGCGGVRFVMCVECNGSCK 378
P +FVKGR +GG +EVV L ++ KL L EG+P G C+ CGGVRFV+C +CNGS K
Sbjct: 42 PRLFVKGRYIGGAEEVVTLHEQGKLKKLFEGVPMDYFDGACDACGGVRFVLCFKCNGSHK 221
Query: 379 VLDE-DKKKTVKCGKCNENGIMQCPICC 405
V+ E ++K++ +C +CNENG++ CP CC
Sbjct: 222 VIAENEEKESTQCPQCNENGLIVCPYCC 305
>BP028689
Length = 563
Score = 99.8 bits (247), Expect = 7e-22
Identities = 52/89 (58%), Positives = 61/89 (68%), Gaps = 7/89 (7%)
Frame = +2
Query: 251 FEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELR 310
F++ CPP GEN VVIYTTTLRG+R+TFE CN VRA +++ V ERDVSMDSGFKEELR
Sbjct: 293 FQRICPPNGENRVVIYTTTLRGVRRTFEACNAVRAAFQAFGVLFCERDVSMDSGFKEELR 472
Query: 311 KLMGAE-------QVKVPVVFVKGRLVGG 332
L + V P VFVKG +GG
Sbjct: 473 MLFKGKGKDASMTMVVPPKVFVKGFYIGG 559
Score = 36.6 bits (83), Expect(2) = 0.001
Identities = 23/49 (46%), Positives = 26/49 (52%), Gaps = 3/49 (6%)
Frame = +2
Query: 34 VSLTSSTYGALKLDKDNDQPVIAAKPREEPEAIT---TINAWELMEGLE 79
VSLTS+TYG L LD P A P T IN+WELM GL+
Sbjct: 80 VSLTSTTYGLLTLDP--QPPHTTAPSTPTPSRFTLPEVINSWELMAGLD 220
Score = 21.9 bits (45), Expect(2) = 0.001
Identities = 9/18 (50%), Positives = 11/18 (61%)
Frame = +1
Query: 1 MGCVSSKLIRKDIKQEHV 18
MGCVSS L+ D H+
Sbjct: 1 MGCVSSNLLNHDDDFSHL 54
>TC14876 weakly similar to UP|Q9ZVB8 (Q9ZVB8) At2g41330 protein
(At2g41330/F13H10.12), partial (18%)
Length = 822
Score = 95.9 bits (237), Expect = 1e-20
Identities = 46/104 (44%), Positives = 69/104 (66%), Gaps = 5/104 (4%)
Frame = +1
Query: 306 KEELRKLMGAEQVK---VPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKAVGG--CE 360
K+EL L G + ++ +P VF++GR VGG D + +L + +LG +LEG+P+ G CE
Sbjct: 1 KKELMGLFGEKNMRNVVLPQVFIRGRCVGGADVIKQLCEVGELGKILEGLPRTKPGFVCE 180
Query: 361 GCGGVRFVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
CG +RFV C C+GS KV D+D+ +C +CNENG+++CP C
Sbjct: 181 SCGDMRFVPCGNCSGSRKVFDDDEGLLKRCLECNENGLLRCPNC 312
>TC9884 weakly similar to GB|AAO63337.1|28950827|BT005273 At5g58530
{Arabidopsis thaliana;}, partial (31%)
Length = 557
Score = 74.3 bits (181), Expect = 3e-14
Identities = 38/86 (44%), Positives = 54/86 (62%), Gaps = 1/86 (1%)
Frame = +2
Query: 320 VPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKA-VGGCEGCGGVRFVMCVECNGSCK 378
+P VF+ GR VGG +EV ++ + +L +L+ +P+ C+ CGG RFV+C EC GS K
Sbjct: 5 LPRVFIGGRYVGGAEEVRQMNEVGELKKILKALPEVDPAECDVCGGHRFVLCDECYGSRK 184
Query: 379 VLDEDKKKTVKCGKCNENGIMQCPIC 404
V E V C CNENG+++CP C
Sbjct: 185 VFTEKAGFRV-CIACNENGLVRCPSC 259
>TC10024 weakly similar to UP|Q8H5B5 (Q8H5B5) OJ1477_F01.28 protein, partial
(14%)
Length = 482
Score = 40.8 bits (94), Expect = 4e-04
Identities = 15/28 (53%), Positives = 21/28 (74%)
Frame = +2
Query: 378 KVLDEDKKKTVKCGKCNENGIMQCPICC 405
KV DED+ +C +CNENG+++CP CC
Sbjct: 2 KVFDEDEGLLKRCLECNENGLVRCPGCC 85
>AV410992
Length = 418
Score = 35.4 bits (80), Expect = 0.017
Identities = 15/24 (62%), Positives = 18/24 (74%)
Frame = +2
Query: 257 PGGENSVVIYTTTLRGIRKTFEDC 280
PG EN VV+Y T+LR +R TFE C
Sbjct: 347 PGTENRVVVYFTSLRVVRPTFEGC 418
>TC10225 similar to UP|Q8LFQ6 (Q8LFQ6) Glutaredoxin, partial (69%)
Length = 725
Score = 34.3 bits (77), Expect = 0.038
Identities = 23/75 (30%), Positives = 39/75 (51%), Gaps = 5/75 (6%)
Frame = +3
Query: 280 CNKVRAIIES-----YCVQMRERDVSMDSGFKEELRKLMGAEQVKVPVVFVKGRLVGGVD 334
CN+ +A+ + Y V++ ERD S ++ L ++G V P VF+ G+ +GG D
Sbjct: 210 CNRAKAVFKELNQVPYVVELDERDDG--SKIQDYLINIVGKRTV--PQVFINGKHLGGSD 377
Query: 335 EVVKLEDEEKLGVLL 349
+ V+ + L LL
Sbjct: 378 DTVEAYESGLLAKLL 422
>TC8466
Length = 1265
Score = 29.3 bits (64), Expect = 1.2
Identities = 17/39 (43%), Positives = 23/39 (58%)
Frame = -1
Query: 74 LMEGLEDGVPISNQPKKSPKSSSPFLRGFMNSDTRSPLK 112
L+ GLE GV Q +K+ S PFLRGF + ++ LK
Sbjct: 617 LINGLEQGV----QQRKTLNSFIPFLRGFKHVGVKNLLK 513
>TC17887 weakly similar to UP|AAQ91191 (AAQ91191)
1-O-sinapoylglucose:choline sinapoyltransferase, partial
(11%)
Length = 595
Score = 28.9 bits (63), Expect = 1.6
Identities = 22/83 (26%), Positives = 41/83 (48%), Gaps = 1/83 (1%)
Frame = +1
Query: 261 NSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDS-GFKEELRKLMGAEQVK 319
N + YTTTL + +++ + K YC + D+S+ G + ++ L + K
Sbjct: 103 NKTLAYTTTLSNVVESYRNLTKANLQALVYCSDL---DMSVPQLGTQHWIKSLNMSISDK 273
Query: 320 VPVVFVKGRLVGGVDEVVKLEDE 342
FV+G+ V G EV K++++
Sbjct: 274 WRAWFVEGQ-VAGFTEVYKMKED 339
>TC9575 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(10%)
Length = 764
Score = 28.5 bits (62), Expect = 2.1
Identities = 9/23 (39%), Positives = 13/23 (56%)
Frame = +1
Query: 356 VGGCEGCGGVRFVMCVECNGSCK 378
V C C ++ + CV C GSC+
Sbjct: 145 VSACASCPSIQAIFCVACCGSCE 213
>TC11945
Length = 542
Score = 28.5 bits (62), Expect = 2.1
Identities = 20/75 (26%), Positives = 31/75 (40%), Gaps = 10/75 (13%)
Frame = +2
Query: 340 EDEEKLGVLLEGIPKAVGG-----CEGCGGVRFVMCVECNGSCKVLDEDK-----KKTVK 389
ED KLG +G+ ++ C C G ++C EC+G + E + + K
Sbjct: 101 EDSSKLGAS-DGVDRSKNKDGTTKCVQCRGEGRLLCTECDGGGEPNIEPQFMELVEDGTK 277
Query: 390 CGKCNENGIMQCPIC 404
C C G C +C
Sbjct: 278 CPYCEGLGYTPCDLC 322
>TC11910 similar to GB|CAA54498.1|602891|SCIILDNA YBL0422 {Saccharomyces
cerevisiae;} , partial (6%)
Length = 638
Score = 27.7 bits (60), Expect = 3.6
Identities = 18/57 (31%), Positives = 26/57 (45%), Gaps = 1/57 (1%)
Frame = +1
Query: 349 LEGIPKAVGGCEGCGGVRFVMCVECN-GSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
L+G P C G GG++ CV CN G K + + C C +G++ C C
Sbjct: 337 LKGDPVPCERCAGNGGMK---CVFCNDGKMK----QEMGLIDCKVCKGSGLILCKKC 486
>TC11908 similar to PIR|T46895|T46895 acyl-CoA dehydrogenase [validated] -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (36%)
Length = 512
Score = 27.3 bits (59), Expect = 4.7
Identities = 9/19 (47%), Positives = 12/19 (62%)
Frame = -3
Query: 13 IKQEHVIIDNCGGGGRYLS 31
+K +H I+ NC GRY S
Sbjct: 114 VKAQHAIVSNCASEGRYFS 58
>AI967451
Length = 425
Score = 26.9 bits (58), Expect = 6.1
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = +3
Query: 317 QVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLL 349
Q VP VF+ G +GG D L ++ KL LL
Sbjct: 258 QRSVPNVFIGGNHIGGCDSTKALHNQGKLVPLL 356
>TC12747 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-like
protein, partial (43%)
Length = 624
Score = 26.9 bits (58), Expect = 6.1
Identities = 11/28 (39%), Positives = 15/28 (53%)
Frame = -2
Query: 359 CEGCGGVRFVMCVECNGSCKVLDEDKKK 386
CEG V ++C E +C +LD KK
Sbjct: 329 CEGIQAVHALLCYEFERTCNILDGFPKK 246
>TC18966 similar to UP|Q93W26 (Q93W26) AT5g53860/K19P17_2, partial (35%)
Length = 554
Score = 26.6 bits (57), Expect = 8.0
Identities = 12/34 (35%), Positives = 18/34 (52%)
Frame = +1
Query: 359 CEGCGGVRFVMCVECNGSCKVLDEDKKKTVKCGK 392
C C G + + C C GS +V + K T+K G+
Sbjct: 340 CRTCNGWQALRCTMCRGSGRVHYQVKSCTLKRGE 441
>AV427009
Length = 404
Score = 26.6 bits (57), Expect = 8.0
Identities = 13/37 (35%), Positives = 16/37 (43%)
Frame = +1
Query: 201 SQSPLFDPELVASYEKELSEEEEQVKRMVWATPKTRR 237
S P FDP L + E R +ATP+ RR
Sbjct: 139 SSLPTFDPTLTPIIDPTYFNESSGTHRQSYATPRQRR 249
>TC19216 similar to UP|MCPI_SOLTU (P01075) Metallocarboxypeptidase inhibitor
(MCPI) (Carboxypeptidase inhibitor), partial (33%)
Length = 406
Score = 26.6 bits (57), Expect = 8.0
Identities = 12/26 (46%), Positives = 17/26 (65%), Gaps = 2/26 (7%)
Frame = +2
Query: 373 CNGSCKVLDEDKKKTVKC--GKCNEN 396
CNG CK LD+ + + C GKCN++
Sbjct: 128 CNGPCKTLDDCSGQLI-CINGKCNDD 202
>TC12910 similar to UP|Q9XEL2 (Q9XEL2) Monodehydroascorbate reductase,
partial (24%)
Length = 482
Score = 26.6 bits (57), Expect = 8.0
Identities = 16/47 (34%), Positives = 24/47 (51%), Gaps = 1/47 (2%)
Frame = +2
Query: 138 VRTGGVRRLDYSNSPKGILKPSNLSPNASKN-SGIPAKGSPICARRK 183
V T V +L S+S ++ S SP + + +P PIC+RRK
Sbjct: 176 VETPPVMQLALSSSTAWLMAASVSSPKRHMHLTSVPLSQKPICSRRK 316
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.134 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,587,941
Number of Sequences: 28460
Number of extensions: 88994
Number of successful extensions: 424
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 411
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 415
length of query: 405
length of database: 4,897,600
effective HSP length: 92
effective length of query: 313
effective length of database: 2,279,280
effective search space: 713414640
effective search space used: 713414640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0010.16


