

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0009.6
(309 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
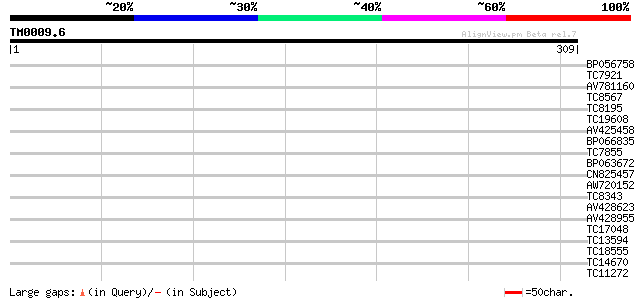
Score E
Sequences producing significant alignments: (bits) Value
BP056758 36 0.009
TC7921 homologue to UP|Q8LHP9 (Q8LHP9) OJ1008_F01.11 protein, pa... 35 0.012
AV781160 35 0.012
TC8567 weakly similar to UP|Q98Y38 (Q98Y38) ORF1, partial (4%) 35 0.016
TC8195 similar to UP|Q7SEW0 (Q7SEW0) Predicted protein, partial ... 34 0.027
TC19608 weakly similar to UP|Q7T2U0 (Q7T2U0) Nicotinic acetylcho... 34 0.036
AV425458 33 0.047
BP066835 27 0.072
TC7855 weakly similar to UP|Q8S9B4 (Q8S9B4) Matrix metalloprotei... 33 0.080
BP063672 29 0.88
CN825457 28 1.5
AW720152 28 1.5
TC8343 weakly similar to GB|BAB02401.1|9294391|AB023038 cytochro... 28 2.0
AV428623 28 2.6
AV428955 28 2.6
TC17048 homologue to UP|Q8W199 (Q8W199) Serine acetyltransferase... 28 2.6
TC13594 homologue to GB|AAP47511.1|32331967|AY164937 RTNLB18w {M... 27 4.4
TC18555 similar to GB|AAP68307.1|31711902|BT008868 At2g33570 {Ar... 27 4.4
TC14670 27 4.4
TC11272 similar to UP|Q8LIQ8 (Q8LIQ8) OJ1370_E02.2 protein (OJ13... 27 4.4
>BP056758
Length = 417
Score = 35.8 bits (81), Expect = 0.009
Identities = 20/29 (68%), Positives = 22/29 (74%)
Frame = -2
Query: 152 TINHALITRSVFINQTLVPVSLELKDTVV 180
TI+HALIT+S T V VSLELKDTVV
Sbjct: 251 TIDHALITQSCSDTMT*VLVSLELKDTVV 165
>TC7921 homologue to UP|Q8LHP9 (Q8LHP9) OJ1008_F01.11 protein, partial (7%)
Length = 1202
Score = 35.4 bits (80), Expect = 0.012
Identities = 16/30 (53%), Positives = 20/30 (66%)
Frame = +1
Query: 33 TSGGFSSTLGGSFTQISGFVLVFLLGHHHL 62
T GGFS+T+GG TQISG+ H+HL
Sbjct: 211 TVGGFSTTIGGFITQISGYFAFSSSCHYHL 300
>AV781160
Length = 352
Score = 35.4 bits (80), Expect = 0.012
Identities = 22/42 (52%), Positives = 26/42 (61%), Gaps = 7/42 (16%)
Frame = +3
Query: 38 SSTLGGSFTQISGFVLVF----LLGHH---HLPLTAATLSST 72
SST+ G TQISGFVL F L+GHH H AA ++ST
Sbjct: 225 SSTVDGFSTQISGFVLTF*LSCLVGHHSSVHASSMAAAIAST 350
>TC8567 weakly similar to UP|Q98Y38 (Q98Y38) ORF1, partial (4%)
Length = 489
Score = 35.0 bits (79), Expect = 0.016
Identities = 24/61 (39%), Positives = 31/61 (50%)
Frame = +1
Query: 30 FLPTSGGFSSTLGGSFTQISGFVLVFLLGHHHLPLTAATLSSTAIPVAAAATLASSSLFR 89
F PT GGF ST+GG TQ G L+ L+ LP + T + P +A L L+R
Sbjct: 151 FSPTIGGFPSTVGGFATQ*VGCFLLLLIFPFMLPSSPPTSTHRPPPSHLSAALC---LYR 321
Query: 90 R 90
R
Sbjct: 322 R 324
>TC8195 similar to UP|Q7SEW0 (Q7SEW0) Predicted protein, partial (3%)
Length = 933
Score = 34.3 bits (77), Expect = 0.027
Identities = 18/30 (60%), Positives = 21/30 (70%)
Frame = +2
Query: 26 ILGRFLPTSGGFSSTLGGSFTQISGFVLVF 55
+LG FL + SSTLGGS TQISGF V+
Sbjct: 92 MLGVFLSHNRWVSSTLGGSSTQISGFPFVY 181
>TC19608 weakly similar to UP|Q7T2U0 (Q7T2U0) Nicotinic acetylcholine
receptor alpha 7b subunit (Fragment), partial (3%)
Length = 530
Score = 33.9 bits (76), Expect = 0.036
Identities = 21/46 (45%), Positives = 28/46 (60%), Gaps = 3/46 (6%)
Frame = +1
Query: 38 SSTLGGSFTQISGFVLVFLLG---HHHLPLTAATLSSTAIPVAAAA 80
SST+G TQISGFVL F L HH + AA+ S+ A+ ++ A
Sbjct: 148 SSTVGEFSTQISGFVLAFWLSCFVGHHPSVHAASASTLAVFTSSTA 285
>AV425458
Length = 424
Score = 33.5 bits (75), Expect = 0.047
Identities = 19/48 (39%), Positives = 26/48 (53%)
Frame = +3
Query: 17 NSKDKSNPTILGRFLPTSGGFSSTLGGSFTQISGFVLVFLLGHHHLPL 64
+S +++P + F PT GGFSSTLGG GF+ F LP+
Sbjct: 108 SSSMEASPALGR*FSPTIGGFSSTLGGFLPNKWGFLCYFSFPFLLLPI 251
>BP066835
Length = 452
Score = 27.3 bits (59), Expect(2) = 0.072
Identities = 11/11 (100%), Positives = 11/11 (100%)
Frame = +1
Query: 27 LGRFLPTSGGF 37
LGRFLPTSGGF
Sbjct: 382 LGRFLPTSGGF 414
Score = 24.3 bits (51), Expect(2) = 0.072
Identities = 11/13 (84%), Positives = 11/13 (84%)
Frame = +3
Query: 38 SSTLGGSFTQISG 50
SSTLGG TQISG
Sbjct: 414 SSTLGGFLTQISG 452
>TC7855 weakly similar to UP|Q8S9B4 (Q8S9B4) Matrix metalloproteinase,
partial (3%)
Length = 1074
Score = 32.7 bits (73), Expect = 0.080
Identities = 32/107 (29%), Positives = 44/107 (40%), Gaps = 11/107 (10%)
Frame = +2
Query: 30 FLPTSGGFSSTLGGSFTQISGFV---LVFLLGHHHLPLTAA-----TLSSTAIPVAAAAT 81
F PT GGFSSTLGGS + GF + F + T S+T P A +
Sbjct: 173 FSPTIGGFSSTLGGSLPK*VGFAFSSVCFCPSSSTMSPTTTIPDHRATSATRPPAPPATS 352
Query: 82 LASSSLFRR---HLPPGCQSSRSGTVLLRSRCRVDPYLVPLKHTGFA 125
L++ F + + P C ++ S S C D + P FA
Sbjct: 353 LSTVRPFHKTCHRVFPLCLATCSAAAFF-SSCNHDRLIFPHATVSFA 490
>BP063672
Length = 528
Score = 29.3 bits (64), Expect = 0.88
Identities = 20/47 (42%), Positives = 25/47 (52%)
Frame = +3
Query: 65 TAATLSSTAIPVAAAATLASSSLFRRHLPPGCQSSRSGTVLLRSRCR 111
T + ++T P AAA AS S RR +PP RSG +RSR R
Sbjct: 258 TPSLKNTTTTPSTAAAAAASRS--RRRIPP-----RSGCFSIRSRAR 377
>CN825457
Length = 537
Score = 28.5 bits (62), Expect = 1.5
Identities = 20/53 (37%), Positives = 29/53 (53%)
Frame = +1
Query: 42 GGSFTQISGFVLVFLLGHHHLPLTAATLSSTAIPVAAAATLASSSLFRRHLPP 94
GG TQISGF FLL L+++ +++++ A TLA + HLPP
Sbjct: 7 GGFSTQISGF---FLLS*----LSSSLATTSSLHFAVTPTLAGFISYISHLPP 144
>AW720152
Length = 503
Score = 28.5 bits (62), Expect = 1.5
Identities = 21/71 (29%), Positives = 30/71 (41%), Gaps = 5/71 (7%)
Frame = -3
Query: 221 NPCLCVQCK-----SFEVIKLELLILKCYTHSLIPIQDCSIHGPPKQFPENFLLKQPIKS 275
+P L CK + ++ LL LKC T S+ P ++ P FP N K+
Sbjct: 252 SPSLYFNCKFSFPSEMKPLERTLLFLKCETPSISPFLITAVCRMPVDFPSN---SCSFKN 82
Query: 276 TESEGMFREVC 286
S M R+ C
Sbjct: 81 KSSPWMGRKCC 49
>TC8343 weakly similar to GB|BAB02401.1|9294391|AB023038 cytochrome P450
{Arabidopsis thaliana;} , partial (60%)
Length = 1784
Score = 28.1 bits (61), Expect = 2.0
Identities = 23/97 (23%), Positives = 40/97 (40%), Gaps = 3/97 (3%)
Frame = -1
Query: 64 LTAATLSSTAIPVAAAATLASSSLFRRHLPPGCQSSRSGTVLLRSRCRVDPYLVP---LK 120
+ T+ ST + + + S +R ++ G + LR CR+ ++ L
Sbjct: 1304 IDTGTIISTPAGIVRSPSFVSLRRYRANITTGGYN-------LRLSCRIIDTILS*PSLS 1146
Query: 121 HTGFAFRRRSTTSAEPFCCKEGARVKNVNRTTINHAL 157
+GF+ TS+ F C+ G + N T N AL
Sbjct: 1145 *SGFSLPNTLKTSSLAFACQSGCLLSNNKVQTSNSAL 1035
>AV428623
Length = 291
Score = 27.7 bits (60), Expect = 2.6
Identities = 14/33 (42%), Positives = 21/33 (63%)
Frame = +1
Query: 67 ATLSSTAIPVAAAATLASSSLFRRHLPPGCQSS 99
+T SS+A+P+AA+A ASS+ H P +S
Sbjct: 127 STASSSAVPIAASAPPASSAFTLPHSAPTALNS 225
>AV428955
Length = 363
Score = 27.7 bits (60), Expect = 2.6
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = +3
Query: 33 TSGGFSSTLGGSFTQISGFVLVFLLGHHHL 62
T GGFS+T+ G T ISG + H+HL
Sbjct: 117 TVGGFSTTIRGFITHISGNLGFSSSCHYHL 206
>TC17048 homologue to UP|Q8W199 (Q8W199) Serine acetyltransferase, partial
(47%)
Length = 676
Score = 27.7 bits (60), Expect = 2.6
Identities = 30/105 (28%), Positives = 40/105 (37%), Gaps = 5/105 (4%)
Frame = +1
Query: 51 FVLVFLLGHH---HLPLTAATLSSTAIPVAAAATLASSSLFRRHLPPG--CQSSRSGTVL 105
F L + LG+ + P T + T P + TL+SS PP CQ + S T
Sbjct: 73 FSLCWFLGNSTPPYQPRHRVTAAVTVSPPSL*TTLSSS-------PPID*CQPANSATHR 231
Query: 106 LRSRCRVDPYLVPLKHTGFAFRRRSTTSAEPFCCKEGARVKNVNR 150
R RC P K G RR A + +RV + R
Sbjct: 232 RRRRCLTPPPAAGSKKRGGCGRRSRRRHAVTLSLNQRSRVTSTPR 366
>TC13594 homologue to GB|AAP47511.1|32331967|AY164937 RTNLB18w {Medicago
truncatula;} , partial (41%)
Length = 556
Score = 26.9 bits (58), Expect = 4.4
Identities = 14/43 (32%), Positives = 21/43 (48%)
Frame = -3
Query: 79 AATLASSSLFRRHLPPGCQSSRSGTVLLRSRCRVDPYLVPLKH 121
AA ++SS++ PP C R+ +L S R Y +KH
Sbjct: 500 AAAVSSSNI*TTKWPPKCTRQRASKMLNSSTIRRQIYFTHMKH 372
>TC18555 similar to GB|AAP68307.1|31711902|BT008868 At2g33570 {Arabidopsis
thaliana;}, partial (19%)
Length = 759
Score = 26.9 bits (58), Expect = 4.4
Identities = 12/26 (46%), Positives = 15/26 (57%)
Frame = +3
Query: 59 HHHLPLTAATLSSTAIPVAAAATLAS 84
HHH LTA++ SSTA+ AS
Sbjct: 276 HHHRNLTASSCSSTAVTTTTGRARAS 353
>TC14670
Length = 435
Score = 26.9 bits (58), Expect = 4.4
Identities = 17/51 (33%), Positives = 27/51 (52%), Gaps = 4/51 (7%)
Frame = +3
Query: 59 HHHLPLTA----ATLSSTAIPVAAAATLASSSLFRRHLPPGCQSSRSGTVL 105
HHH+P ++ +TL+S I + A T + + RR L C +S S + L
Sbjct: 108 HHHVPTSSLTGTSTLASLKISSSPATTTSMRTATRRRL--SCSASASPSTL 254
>TC11272 similar to UP|Q8LIQ8 (Q8LIQ8) OJ1370_E02.2 protein (OJ1354_H07.12
protein), partial (59%)
Length = 1070
Score = 26.9 bits (58), Expect = 4.4
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +2
Query: 64 LTAATLSSTAIPVAAAATLASSSLFRRHLPPGCQSSRSGTV 104
+ +A +S +P A ++ S +FR L + SRSGT+
Sbjct: 308 VASADAASRPVPAHKAVLVSRSPVFRAMLENDMEESRSGTI 430
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,318,092
Number of Sequences: 28460
Number of extensions: 101574
Number of successful extensions: 630
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 623
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 628
length of query: 309
length of database: 4,897,600
effective HSP length: 90
effective length of query: 219
effective length of database: 2,336,200
effective search space: 511627800
effective search space used: 511627800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0009.6


