

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.6
(566 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
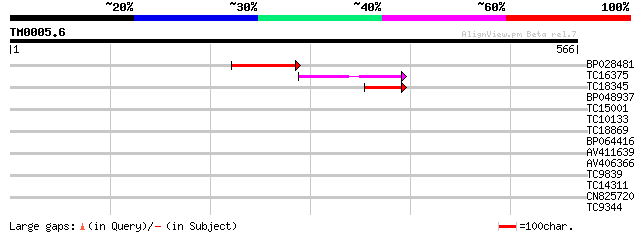
Score E
Sequences producing significant alignments: (bits) Value
BP028481 74 5e-14
TC16375 weakly similar to UP|Q9FEV9 (Q9FEV9) Microtubule-associa... 74 6e-14
TC18345 similar to UP|Q84VU1 (Q84VU1) 65 kDa microtubule associa... 41 5e-04
BP048937 37 0.009
TC15001 similar to UP|GLE1_YEAST (Q12315) RNA export factor GLE1... 32 0.21
TC10133 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, p... 30 1.4
TC18869 similar to UP|Q9SIR1 (Q9SIR1) At2g25320 protein, partial... 28 3.0
BP064416 28 3.0
AV411639 28 5.2
AV406366 27 6.8
TC9839 similar to UP|Q8S9J5 (Q8S9J5) At1g11170/T28P6_16, partial... 27 6.8
TC14311 homologue to UP|Q9FFC0 (Q9FFC0) Histone H2B like protein... 27 6.8
CN825720 27 8.9
TC9344 similar to UP|AAQ84317 (AAQ84317) Fiber NTGP1-related pro... 27 8.9
>BP028481
Length = 313
Score = 74.3 bits (181), Expect = 5e-14
Identities = 37/69 (53%), Positives = 52/69 (74%)
Frame = +1
Query: 222 DTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVD 281
DTLA L + +LK++K+QRL K+QEL L+DLW+L++TP +E+ F+HVT ISASVD
Sbjct: 1 DTLARLAKTVLTLKEDKKQRLHKLQELASQLIDLWNLMDTPPEERSLFDHVTCNISASVD 180
Query: 282 EVSTHGCLS 290
EV+ G L+
Sbjct: 181 EVTVPGALA 207
>TC16375 weakly similar to UP|Q9FEV9 (Q9FEV9) Microtubule-associated protein
MAP65-1a, partial (16%)
Length = 732
Score = 73.9 bits (180), Expect = 6e-14
Identities = 42/108 (38%), Positives = 63/108 (57%)
Frame = +2
Query: 289 LSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIE 348
L+ D+I Q E E++ L L S MKD+ K+Q +LEEI+ H+++D E A
Sbjct: 2 LAMDMI*QAEVEIESLGQLTTSMMKDISVKKQGKLEEIFSRAHVEIDPEDA--------- 154
Query: 349 SGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLD 396
+ S +L SMD Q +A+ A SR++ILD+V+K A EEE W++
Sbjct: 155 IDRMGHSSILASMDSQKAKAEA*ALSRKDILDKVDK*MLAREEESWVE 298
>TC18345 similar to UP|Q84VU1 (Q84VU1) 65 kDa microtubule associated
protein, partial (8%)
Length = 521
Score = 41.2 bits (95), Expect = 5e-04
Identities = 20/42 (47%), Positives = 29/42 (68%)
Frame = +3
Query: 355 SDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLD 396
S +L SMD Q +A+ A SR++ILD+V+K A EEE W++
Sbjct: 48 SSILASMDSQKAKAEA*ALSRKDILDKVDK*MLAREEESWVE 173
>BP048937
Length = 561
Score = 37.0 bits (84), Expect = 0.009
Identities = 17/32 (53%), Positives = 25/32 (78%)
Frame = -1
Query: 97 KQQLATIRPVLEDLRSKKDERVKEFSKIKSQI 128
++QL T PVLE L +K+ER++EFS ++SQI
Sbjct: 507 EEQLETKVPVLEQLWQQKEERIQEFSDVQSQI 412
>TC15001 similar to UP|GLE1_YEAST (Q12315) RNA export factor GLE1, partial
(4%)
Length = 988
Score = 32.3 bits (72), Expect = 0.21
Identities = 28/132 (21%), Positives = 62/132 (46%), Gaps = 7/132 (5%)
Frame = +2
Query: 298 EAEVQRLNALKASKMKDLVFKRQNEL---EEIYRGVHMDVDSEAARQILTSLIESGNIDL 354
EAE++ + A ++++ + KR E EE+ + ++ R I+ ++
Sbjct: 368 EAELKLIEEETAKRVEEAIRKRVEESLNSEEVQVEIQRRLEEGRKRLIVEVAVQLEKEKE 547
Query: 355 SDLLQSMDDQIRQAKEQAQSRREILDRV----EKWRFAAEEEKWLDEYERDENRYSAVRG 410
+ L+++ R+ +EQA+ +E LDR+ K A+ + L++ R+E RY +
Sbjct: 548 AALIEA-----REKEEQARKEKEDLDRMLEENRKKIEEAQRREALEQQRREEERYRELEE 712
Query: 411 AHKNLKRAEKAR 422
+ + A + +
Sbjct: 713 IQRQKEEAMRRK 748
>TC10133 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, partial
(34%)
Length = 1827
Score = 29.6 bits (65), Expect = 1.4
Identities = 76/444 (17%), Positives = 181/444 (40%), Gaps = 20/444 (4%)
Frame = +1
Query: 19 LSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAEL-- 76
L + Q + E+ ++ R L ++ +EC + +++A++KH + LA A E+
Sbjct: 193 LEESQKQYLELCTTEEARITELQKITEECNRTRQSELEASQKHLSVDSAALASAANEIQL 372
Query: 77 ----INLVSSLGESSSFSRGKG------TLKQQLATIRPVLEDLRSKKDERVKEFSKIKS 126
+ LV++ ES + LKQ L+ ++E++++ +++ F + ++
Sbjct: 373 LKVQLELVANC-ESVQTHHAESADVELLNLKQNLSETLSLVENMKN----QLRNFKEFEA 537
Query: 127 QISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTI 186
Q + E + + E+ + D+ K + KS +L ++ ++T+
Sbjct: 538 QAQGLVSETLLQLETAKRTVEILRGDVA-KSVDGYKSIALELDQS--------RAKVNTL 690
Query: 187 SELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQ 246
L ++ DF ++ + GD G + SL + L+ + +K Q
Sbjct: 691 EVLVEKLNPDF---INNNCNNSGDLVDGHRSDHVEAEICSLKSDVGRLRSAVETAEIKYQ 861
Query: 247 ELTKFLVDLWDLIETPIDEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNA 306
E + I + + + A+ + ++ S E C +++ +A+++ L A
Sbjct: 862 E---------EQIRSTVQIRNAYELMEQIKS----ESGQKECEFEAELKRKKADIEELKA 1002
Query: 307 L---KASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQI-----LTSLIESGNIDLSDLL 358
K ++++ ++ + +N ++ D E +I + +++ +D L
Sbjct: 1003NLMDKETELQGIMEENENLNSKLQVRTSCQSDHELRNEIKRLEHCVAALKADMMDKETTL 1182
Query: 359 QSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRA 418
QS+ ++ K + EI + AAE E + + S+ R A + ++
Sbjct: 1183QSISEENEMLKLERNKSEEI-----EAAKAAEHEALMKLGIAMQEVESSSRKAARVTEQL 1347
Query: 419 EKARIVVSKIPSIVENLTTKVKAW 442
E A+ S++ + + + + W
Sbjct: 1348EAAQASNSEMEAELRRMKVQSDQW 1419
>TC18869 similar to UP|Q9SIR1 (Q9SIR1) At2g25320 protein, partial (8%)
Length = 381
Score = 28.5 bits (62), Expect = 3.0
Identities = 31/133 (23%), Positives = 54/133 (40%), Gaps = 2/133 (1%)
Frame = +1
Query: 89 FSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEV 148
FSR K L +Q+ I LE LRS++D+ + S K + Q + +
Sbjct: 1 FSREKKELTEQVQEIESQLEWLRSERDDEKVKLSAEKKVL-----------QDRLHDADT 147
Query: 149 DQSDLTFKKLGELKSHLQDLQ--NEKILRQQKVKSHISTISELSAVMSIDFWKTLSEIHP 206
S L +K ELK +++ E++ + + + A ++ T EI
Sbjct: 148 QLSQLKSRKRDELKKVVKEKNALAERLKNAEAARKRFDEELKRFATENV----TREEIRQ 315
Query: 207 SLGDSSKGTPQSI 219
SL D + Q++
Sbjct: 316 SLEDEVRRLTQTV 354
>BP064416
Length = 528
Score = 28.5 bits (62), Expect = 3.0
Identities = 21/98 (21%), Positives = 43/98 (43%), Gaps = 15/98 (15%)
Frame = +1
Query: 153 LTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMS---------------IDF 197
+ F + L LQ Q++ +L+ Q++ + S ++ ++ + +
Sbjct: 178 ILFLSIIHLSEQLQYSQSQTLLKLQQLLGYPSALNTFTSTTPNTDYCNIEPTPYLTLVCY 357
Query: 198 WKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLK 235
L+++H + PQS S+D+L S G + SLK
Sbjct: 358 EDNLTQLHVVGNNEFTPLPQSFSSDSLFSTXGTLSSLK 471
>AV411639
Length = 349
Score = 27.7 bits (60), Expect = 5.2
Identities = 10/24 (41%), Positives = 14/24 (57%)
Frame = -1
Query: 3 ATPPSFSPSRTTCASLLSQLQMIW 26
++PPSF+P R C + Q IW
Sbjct: 346 SSPPSFTPLRPACPTRFLQANQIW 275
>AV406366
Length = 412
Score = 27.3 bits (59), Expect = 6.8
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = -3
Query: 388 AAEEEKWLDEYERDENRYSA 407
AAE+E W+ + RDENR A
Sbjct: 107 AAEDESWMARHVRDENRSRA 48
>TC9839 similar to UP|Q8S9J5 (Q8S9J5) At1g11170/T28P6_16, partial (17%)
Length = 672
Score = 27.3 bits (59), Expect = 6.8
Identities = 13/33 (39%), Positives = 20/33 (60%), Gaps = 1/33 (3%)
Frame = +1
Query: 373 QSRREILDRVEKWRFA-AEEEKWLDEYERDENR 404
QS E+ E+W A AE+ W+D ++RD+ R
Sbjct: 184 QSSWELQIFKERWNKAVAEDRNWVDPFKRDQRR 282
>TC14311 homologue to UP|Q9FFC0 (Q9FFC0) Histone H2B like protein, partial
(97%)
Length = 626
Score = 27.3 bits (59), Expect = 6.8
Identities = 17/56 (30%), Positives = 26/56 (46%), Gaps = 2/56 (3%)
Frame = +1
Query: 195 IDFWKTLSEIHPSLGDSSK--GTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQEL 248
I +K L ++HP +G SSK G S ND L L + Q+ + +E+
Sbjct: 211 IYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNQKPTITSREI 378
>CN825720
Length = 699
Score = 26.9 bits (58), Expect = 8.9
Identities = 28/129 (21%), Positives = 55/129 (41%)
Frame = +1
Query: 42 QLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFSRGKGTLKQQLA 101
QL + L+I +++ A A L L EA S ++R L++
Sbjct: 163 QLSPDLLNIMEQRLSAIEHRSACLSNLLNQPEAT----------PSEYARANKELRKLSG 312
Query: 102 TIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSSEEVDQSDLTFKKLGEL 161
++ ++ +LR+K+ KE +K+ +++ SSE+ D + ++ G+
Sbjct: 313 SVE-LINELRAKQ----KEIDGLKTLMAE-------------SSEDKDMLSMATEEFGQA 438
Query: 162 KSHLQDLQN 170
K + LQN
Sbjct: 439 KEEERRLQN 465
>TC9344 similar to UP|AAQ84317 (AAQ84317) Fiber NTGP1-related protein,
complete
Length = 995
Score = 26.9 bits (58), Expect = 8.9
Identities = 22/61 (36%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Frame = +2
Query: 314 DLVFKRQNELEEIYRGVHMDVDSEAAR-QILTSLIESGNIDLSDLLQSMDDQIRQAKEQA 372
D + K Q EL+E +H +DS AR + L SL+E SDL + +QAK+
Sbjct: 452 DKLMKIQKELDETKIILHKTIDSVLARGEKLDSLVEKS----SDLSAASQMFYKQAKKTN 619
Query: 373 Q 373
Q
Sbjct: 620 Q 622
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.128 0.347
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,864,261
Number of Sequences: 28460
Number of extensions: 87055
Number of successful extensions: 444
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 440
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 443
length of query: 566
length of database: 4,897,600
effective HSP length: 95
effective length of query: 471
effective length of database: 2,193,900
effective search space: 1033326900
effective search space used: 1033326900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0005.6


