

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.4
(283 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
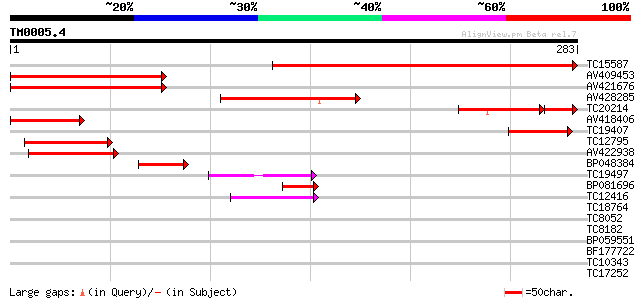
Score E
Sequences producing significant alignments: (bits) Value
TC15587 similar to UP|Q9LHG0 (Q9LHG0) Genomic DNA, chromosome 3,... 311 1e-85
AV409453 156 3e-39
AV421676 154 2e-38
AV428285 100 2e-22
TC20214 similar to UP|Q9LHG0 (Q9LHG0) Genomic DNA, chromosome 3,... 58 4e-13
AV418406 69 9e-13
TC19407 55 2e-08
TC12795 UP|Q9FMV5 (Q9FMV5) Transcription factor Hap5a-like prote... 48 2e-06
AV422938 45 1e-05
BP048384 43 5e-05
TC19497 weakly similar to UP|Q8LA02 (Q8LA02) Contains similarity... 40 4e-04
BP081696 40 6e-04
TC12416 similar to GB|AAH56066.1|33417132|BC056066 sop-prov prot... 39 8e-04
TC18764 similar to UP|Q8LBJ5 (Q8LBJ5) Nucleic acid binding prote... 35 0.014
TC8052 similar to GB|AAP13431.1|30023796|BT006323 At5g02530 {Ara... 34 0.032
TC8182 weakly similar to UP|Q93VA8 (Q93VA8) At1g76010/T4O12_22, ... 33 0.042
BP059551 33 0.042
BF177722 33 0.055
TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (... 32 0.12
TC17252 similar to UP|O04237 (O04237) Transcription factor, part... 32 0.12
>TC15587 similar to UP|Q9LHG0 (Q9LHG0) Genomic DNA, chromosome 3, P1 clone:
MQC3 (At3g12480/MQC3.32), partial (7%)
Length = 768
Score = 311 bits (796), Expect = 1e-85
Identities = 151/152 (99%), Positives = 152/152 (99%)
Frame = +2
Query: 132 LSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDTNTDMTIHDSS 191
+SHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDTNTDMTIHDSS
Sbjct: 8 VSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDTNTDMTIHDSS 187
Query: 192 ETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPPTESQHEEIPGWS 251
ETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPPTESQHEEIPGWS
Sbjct: 188 ETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPPTESQHEEIPGWS 367
Query: 252 LSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG 283
LSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG
Sbjct: 368 LSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG 463
>AV409453
Length = 402
Score = 156 bits (395), Expect = 3e-39
Identities = 78/78 (100%), Positives = 78/78 (100%)
Frame = +3
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK
Sbjct: 168 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 347
Query: 61 TMNSLHLKHCVQSYNVFD 78
TMNSLHLKHCVQSYNVFD
Sbjct: 348 TMNSLHLKHCVQSYNVFD 401
>AV421676
Length = 424
Score = 154 bits (389), Expect = 2e-38
Identities = 76/78 (97%), Positives = 78/78 (99%)
Frame = +2
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK
Sbjct: 191 MRKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 370
Query: 61 TMNSLHLKHCVQSYNVFD 78
TMN+LHLKHCVQSYNVFD
Sbjct: 371 TMNALHLKHCVQSYNVFD 424
>AV428285
Length = 239
Score = 100 bits (250), Expect = 2e-22
Identities = 50/73 (68%), Positives = 55/73 (74%), Gaps = 3/73 (4%)
Frame = +3
Query: 106 IPKRRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRG---RTAVRETPH 162
+PKRRKAAGDD NDSDEE KR KM EL HT STG+GRGRGRGRGRGRG RT +ET
Sbjct: 3 LPKRRKAAGDDGNDSDEEAKRSKMLELGHTSSTGKGRGRGRGRGRGRGRGARTVEKETSV 182
Query: 163 QAVEPEPCASVQQ 175
VE EPC S+ +
Sbjct: 183 HQVESEPCTSIPE 221
>TC20214 similar to UP|Q9LHG0 (Q9LHG0) Genomic DNA, chromosome 3, P1 clone:
MQC3 (At3g12480/MQC3.32), partial (5%)
Length = 576
Score = 57.8 bits (138), Expect(2) = 4e-13
Identities = 30/44 (68%), Positives = 34/44 (77%), Gaps = 1/44 (2%)
Frame = +1
Query: 225 KKASTTAKQEISE-PPTESQHEEIPGWSLSDVDKMAIDSVQLAN 267
K AST A+ + E TE +HEEIPGWSLSDVDKMAID +QLAN
Sbjct: 1 KGASTAAQASLPELAETEIKHEEIPGWSLSDVDKMAIDHMQLAN 132
Score = 32.3 bits (72), Expect(2) = 4e-13
Identities = 12/16 (75%), Positives = 15/16 (93%)
Frame = +2
Query: 268 LGTHIEEDEEDYDEEG 283
LG+ +E+DEEDYDEEG
Sbjct: 134 LGSRVEDDEEDYDEEG 181
>AV418406
Length = 188
Score = 68.9 bits (167), Expect = 9e-13
Identities = 33/37 (89%), Positives = 36/37 (97%)
Frame = +1
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSK 37
M+KKLDTRFPAARIKKIMQ DED+GKIALAVP+LVSK
Sbjct: 76 MRKKLDTRFPAARIKKIMQTDEDIGKIALAVPLLVSK 186
>TC19407
Length = 418
Score = 54.7 bits (130), Expect = 2e-08
Identities = 24/32 (75%), Positives = 28/32 (87%)
Frame = +3
Query: 250 WSLSDVDKMAIDSVQLANLGTHIEEDEEDYDE 281
WSLSDV+KMAID +QLANL T I++D EDYDE
Sbjct: 3 WSLSDVEKMAIDPIQLANLNTRIDDDVEDYDE 98
>TC12795 UP|Q9FMV5 (Q9FMV5) Transcription factor Hap5a-like protein, partial
(40%)
Length = 405
Score = 47.8 bits (112), Expect = 2e-06
Identities = 23/44 (52%), Positives = 32/44 (72%)
Frame = +2
Query: 8 RFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTY 51
+ P ARIKKIM+ADEDV I+ P+L +KA ELF+ +L R++
Sbjct: 197 QLPLARIKKIMKADEDVRMISAEAPILFAKACELFILELTIRSW 328
>AV422938
Length = 466
Score = 45.1 bits (105), Expect = 1e-05
Identities = 22/45 (48%), Positives = 32/45 (70%)
Frame = +3
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEIT 54
P ARIKKIM+ADEDV I+ PV+ ++A E+F+ +L R++ T
Sbjct: 330 PLARIKKIMKADEDVRMISAEAPVVFARACEMFILELTLRSWNHT 464
>BP048384
Length = 465
Score = 43.1 bits (100), Expect = 5e-05
Identities = 19/25 (76%), Positives = 22/25 (88%)
Frame = -3
Query: 65 LHLKHCVQSYNVFDFLRDVVSKVPD 89
L L HCV+S NVFDFL+D+VSKVPD
Sbjct: 445 LILFHCVESSNVFDFLKDIVSKVPD 371
>TC19497 weakly similar to UP|Q8LA02 (Q8LA02) Contains similarity to
RNA-binding protein from Arabidopsis thaliana gi|2129727
and contains RNA recognition PF|00076 domain, partial
(11%)
Length = 436
Score = 40.0 bits (92), Expect = 4e-04
Identities = 24/54 (44%), Positives = 28/54 (51%)
Frame = +2
Query: 100 SADDRTIPKRRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRG 153
S DD R+ + D + SD G+ G GRGRGRGRGRGRGRG
Sbjct: 23 SRDDAINNARKILSKGDIDGSDT----GRERTPREGGGLGRGRGRGRGRGRGRG 172
Score = 35.8 bits (81), Expect = 0.008
Identities = 15/16 (93%), Positives = 15/16 (93%)
Frame = +2
Query: 139 GRGRGRGRGRGRGRGR 154
G GRGRGRGRGRGRGR
Sbjct: 122 GLGRGRGRGRGRGRGR 169
>BP081696
Length = 383
Score = 39.7 bits (91), Expect = 6e-04
Identities = 17/18 (94%), Positives = 17/18 (94%)
Frame = -3
Query: 137 STGRGRGRGRGRGRGRGR 154
S GRGRGRGRGRGRGRGR
Sbjct: 186 SRGRGRGRGRGRGRGRGR 133
Score = 38.1 bits (87), Expect = 0.002
Identities = 16/19 (84%), Positives = 16/19 (84%)
Frame = -3
Query: 136 GSTGRGRGRGRGRGRGRGR 154
G RGRGRGRGRGRGRGR
Sbjct: 195 GGRSRGRGRGRGRGRGRGR 139
>TC12416 similar to GB|AAH56066.1|33417132|BC056066 sop-prov protein
{Xenopus laevis;} , partial (32%)
Length = 416
Score = 39.3 bits (90), Expect = 8e-04
Identities = 21/44 (47%), Positives = 22/44 (49%)
Frame = +3
Query: 111 KAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGR 154
K +GD S E G GRGRG GRGRGRGRGR
Sbjct: 75 KMSGDGRGASGEPSAEGFGDGFGRGRGEGRGRGEGRGRGRGRGR 206
Score = 37.0 bits (84), Expect = 0.004
Identities = 16/20 (80%), Positives = 16/20 (80%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGRTAVR 158
GRG GRGRGRGRGRGR R
Sbjct: 165 GRGEGRGRGRGRGRGRRGRR 224
Score = 27.7 bits (60), Expect = 2.3
Identities = 12/15 (80%), Positives = 12/15 (80%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRG 153
GRGRGRGRGR RG
Sbjct: 183 GRGRGRGRGRRGRRG 227
>TC18764 similar to UP|Q8LBJ5 (Q8LBJ5) Nucleic acid binding protein-like,
partial (35%)
Length = 848
Score = 35.0 bits (79), Expect = 0.014
Identities = 15/16 (93%), Positives = 15/16 (93%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGR 154
GRGRGRGRGRGRGR R
Sbjct: 174 GRGRGRGRGRGRGRFR 221
Score = 30.4 bits (67), Expect = 0.36
Identities = 13/16 (81%), Positives = 13/16 (81%)
Frame = +3
Query: 143 GRGRGRGRGRGRTAVR 158
GRGRGRGRGRGR R
Sbjct: 174 GRGRGRGRGRGRGRFR 221
>TC8052 similar to GB|AAP13431.1|30023796|BT006323 At5g02530 {Arabidopsis
thaliana;}, partial (55%)
Length = 1389
Score = 33.9 bits (76), Expect = 0.032
Identities = 18/30 (60%), Positives = 18/30 (60%), Gaps = 2/30 (6%)
Frame = +2
Query: 127 GKMPELSHTGSTGRGRG--RGRGRGRGRGR 154
G L G GRGRG RGRGRGRG GR
Sbjct: 722 GAFGRLRGGGDGGRGRGHRRGRGRGRGHGR 811
Score = 33.5 bits (75), Expect = 0.042
Identities = 16/21 (76%), Positives = 16/21 (76%), Gaps = 2/21 (9%)
Frame = +2
Query: 136 GSTGRG--RGRGRGRGRGRGR 154
G GRG RGRGRGRG GRGR
Sbjct: 755 GGRGRGHRRGRGRGRGHGRGR 817
Score = 33.5 bits (75), Expect = 0.042
Identities = 17/35 (48%), Positives = 19/35 (53%)
Frame = +2
Query: 140 RGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQ 174
RGRGRGRG GRGRG E +E S+Q
Sbjct: 779 RGRGRGRGHGRGRGEKVSAEDLDADLEKYHAESMQ 883
Score = 30.4 bits (67), Expect = 0.36
Identities = 12/13 (92%), Positives = 12/13 (92%)
Frame = +2
Query: 139 GRGRGRGRGRGRG 151
GRGRGRG GRGRG
Sbjct: 782 GRGRGRGHGRGRG 820
Score = 27.7 bits (60), Expect = 2.3
Identities = 11/16 (68%), Positives = 11/16 (68%)
Frame = +2
Query: 134 HTGSTGRGRGRGRGRG 149
H GRGRG GRGRG
Sbjct: 773 HRRGRGRGRGHGRGRG 820
>TC8182 weakly similar to UP|Q93VA8 (Q93VA8) At1g76010/T4O12_22, partial
(13%)
Length = 611
Score = 33.5 bits (75), Expect = 0.042
Identities = 16/18 (88%), Positives = 16/18 (88%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGRTA 156
GRGRGRG GRGRGRGR A
Sbjct: 207 GRGRGRG-GRGRGRGRNA 257
Score = 31.6 bits (70), Expect = 0.16
Identities = 16/20 (80%), Positives = 16/20 (80%), Gaps = 1/20 (5%)
Frame = +3
Query: 136 GSTGRGRGRGRGRGRG-RGR 154
G GRGRG GRGRGRG RGR
Sbjct: 72 GYGGRGRGFGRGRGRGFRGR 131
>BP059551
Length = 454
Score = 33.5 bits (75), Expect = 0.042
Identities = 17/26 (65%), Positives = 17/26 (65%)
Frame = -1
Query: 139 GRGRGRGRGRGRGRGRTAVRETPHQA 164
GRGRGRGRG RGRGR P QA
Sbjct: 352 GRGRGRGRGGYRGRGRGFRSNGPIQA 275
>BF177722
Length = 399
Score = 33.1 bits (74), Expect = 0.055
Identities = 20/60 (33%), Positives = 27/60 (44%)
Frame = -3
Query: 126 RGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDTNTDM 185
R + P G T RGR RG R R +V + + E PCA V GI + + D+
Sbjct: 196 RHERPGRRAHGRTVTAAARGRRRGGSRRRESVSDEGIELAEERPCAGVIAGIGNPSLLDL 17
>TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (Fibrillarin
2) (AtFib2), partial (85%)
Length = 1116
Score = 32.0 bits (71), Expect = 0.12
Identities = 18/30 (60%), Positives = 19/30 (63%), Gaps = 1/30 (3%)
Frame = +1
Query: 136 GSTGRGRG-RGRGRGRGRGRTAVRETPHQA 164
G RGRG RGRGRG G GR R TP +A
Sbjct: 61 GGGFRGRGDRGRGRGDGGGRGGGRGTPFKA 150
>TC17252 similar to UP|O04237 (O04237) Transcription factor, partial (21%)
Length = 492
Score = 32.0 bits (71), Expect = 0.12
Identities = 22/69 (31%), Positives = 27/69 (38%), Gaps = 3/69 (4%)
Frame = +1
Query: 88 PDYSHGHGHSEPSADDRT---IPKRRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGR 144
P HG + P+A +P + +S E RG GRG GR
Sbjct: 286 PKKGQEHGKAGPAAQQNKPAQLPSKPLPPTQAVRESKNEASRG-----------GRGGGR 432
Query: 145 GRGRGRGRG 153
G GRG GRG
Sbjct: 433 GGGRGFGRG 459
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,201,986
Number of Sequences: 28460
Number of extensions: 54126
Number of successful extensions: 893
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 472
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 679
length of query: 283
length of database: 4,897,600
effective HSP length: 89
effective length of query: 194
effective length of database: 2,364,660
effective search space: 458744040
effective search space used: 458744040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0005.4


