

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.1
(346 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
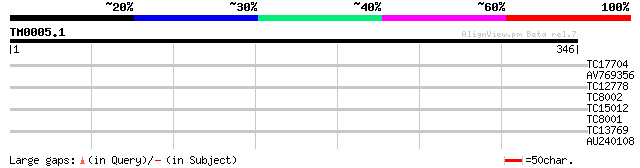
TC17704 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (7%) 31 0.35
AV769356 28 2.3
TC12778 27 3.9
TC8002 similar to GB|AAA28385.1|156990|DROBCD segmentation prote... 27 3.9
TC15012 27 5.0
TC8001 similar to UP|Q8LC02 (Q8LC02) Lil3 protein, partial (56%) 27 6.6
TC13769 weakly similar to GB|AAM19768.1|20453045|AY094388 AT3g56... 27 6.6
AU240108 26 8.6
>TC17704 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (7%)
Length = 283
Score = 30.8 bits (68), Expect = 0.35
Identities = 24/92 (26%), Positives = 42/92 (45%), Gaps = 7/92 (7%)
Frame = +2
Query: 153 VQRQTLCAQATEASGVSTNDMGFSSSALIKG-------SGNLNEVSHKPALISEKPSAAM 205
+ Q + A A ++ G S+S L+ GN + + +SE+P +
Sbjct: 5 IGHQQTGSAAAAAQSLAIGTPGISASPLLAEFSGPDGVHGNTFPPTSGKSTVSEQP---L 175
Query: 206 QRLIGVFTSMSSEALGASIGKIREVVHLNDAI 237
+RLI S++ AL A++ + VV +ND I
Sbjct: 176 ERLIKAAKSLTPNALSAAVSDMGSVVSMNDRI 271
>AV769356
Length = 574
Score = 28.1 bits (61), Expect = 2.3
Identities = 10/30 (33%), Positives = 15/30 (49%)
Frame = +1
Query: 13 YQQICSKLRQHESSPHEPKTFNVEKFRGHK 42
+ +CS HE H PK N+ + GH+
Sbjct: 469 FSHLCSSFPCHEFKKHSPKAINIPFWCGHQ 558
>TC12778
Length = 462
Score = 27.3 bits (59), Expect = 3.9
Identities = 13/45 (28%), Positives = 24/45 (52%)
Frame = +1
Query: 80 HQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCL 124
+ L+ +H+ +Q VLP +LG Q+ A+QT++ + L
Sbjct: 319 NSGLTYDASQHTNPLQGSFDVLPSILGHQDFSSAQQTAENRDTTL 453
>TC8002 similar to GB|AAA28385.1|156990|DROBCD segmentation protein PRD 4
{Drosophila melanogaster;} , partial (50%)
Length = 640
Score = 27.3 bits (59), Expect = 3.9
Identities = 22/71 (30%), Positives = 33/71 (45%)
Frame = -1
Query: 75 NSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYG 134
+S+ QH+ ++ H + S L +LG QN H T Q LQ+QQ KQ
Sbjct: 547 SSLPQHRYQHPNRCCHFSGTASSPMCLQPILGPQN-HRESSTPQ-----LQQQQQMKQTH 386
Query: 135 NSEQGVNQLSI 145
S+Q + +I
Sbjct: 385 LSQQRFSPAAI 353
>TC15012
Length = 1089
Score = 26.9 bits (58), Expect = 5.0
Identities = 20/62 (32%), Positives = 28/62 (44%), Gaps = 10/62 (16%)
Frame = -1
Query: 195 ALISEKPSAAMQRLIGVFTSMSSEALG----------ASIGKIREVVHLNDAIPAAKFID 244
ALI+E PSA M+ L+ S+ ++L A GK V L DA F+
Sbjct: 282 ALINEAPSAVMRSLLRATPSLDGKSLKTKYATSNLPVALFGKWSREVSLFDATSCKAFLV 103
Query: 245 EP 246
+P
Sbjct: 102 DP 97
>TC8001 similar to UP|Q8LC02 (Q8LC02) Lil3 protein, partial (56%)
Length = 1039
Score = 26.6 bits (57), Expect = 6.6
Identities = 22/71 (30%), Positives = 33/71 (45%)
Frame = -3
Query: 75 NSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYG 134
+S+ QH+ ++ H + S L +LG QN H T Q LQ+QQ KQ
Sbjct: 347 SSLPQHR*QHPNRCCHFSGTASSPMCLQPILGPQN-HRESSTPQ-----LQQQQQMKQTH 186
Query: 135 NSEQGVNQLSI 145
S+Q + +I
Sbjct: 185 LSQQRFSPAAI 153
>TC13769 weakly similar to GB|AAM19768.1|20453045|AY094388
AT3g56200/F18O21_160 {Arabidopsis thaliana;}, partial
(20%)
Length = 438
Score = 26.6 bits (57), Expect = 6.6
Identities = 15/52 (28%), Positives = 25/52 (47%)
Frame = +2
Query: 192 HKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAAKFI 243
H P L KP+A + G ++++ +GA I I ++ + IPA I
Sbjct: 62 HVPLLPGSKPAAPPASVSGAVFNVANSIVGAGIMSIPALLKVLGVIPAFAMI 217
>AU240108
Length = 300
Score = 26.2 bits (56), Expect = 8.6
Identities = 13/40 (32%), Positives = 17/40 (42%)
Frame = +1
Query: 243 IDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTSRSCAR 282
ID PL +P+R+PQG R + R C R
Sbjct: 97 IDNPLPSLRPCREPARVPQGCTHRRG*GCLRLRRQRCCHR 216
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.130 0.362
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,930,837
Number of Sequences: 28460
Number of extensions: 57019
Number of successful extensions: 360
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 357
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 360
length of query: 346
length of database: 4,897,600
effective HSP length: 91
effective length of query: 255
effective length of database: 2,307,740
effective search space: 588473700
effective search space used: 588473700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0005.1


