

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0004a.3
(148 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
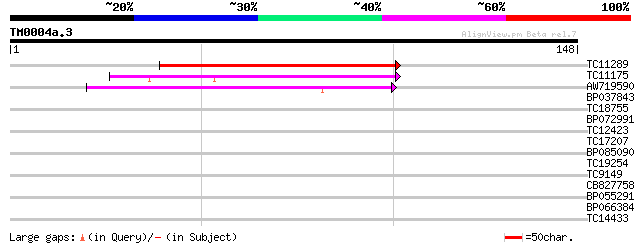
Score E
Sequences producing significant alignments: (bits) Value
TC11289 weakly similar to UP|Q9LUE5 (Q9LUE5) Emb|CAB86483.1, par... 62 5e-11
TC11175 weakly similar to GB|AAL06796.1|15809736|AY054135 AT3g60... 60 2e-10
AW719590 55 4e-09
BP037843 32 0.059
TC18755 similar to UP|AAN85378 (AAN85378) Resistance protein (Fr... 29 0.29
BP072991 29 0.38
TC12423 similar to PIR|T01558|T01558 auxin-induced protein homol... 28 0.50
TC17207 27 1.5
BP085090 27 1.5
TC19254 weakly similar to UP|Q94C33 (Q94C33) AT3g51180/F24M12_22... 27 1.9
TC9149 similar to UP|Q8LSK7 (Q8LSK7) Auxin-regulated protein, pa... 25 4.2
CB827758 25 5.5
BP055291 25 7.2
BP066384 24 9.4
TC14433 similar to UP|Q41057 (Q41057) Cysteine protease , complete 24 9.4
>TC11289 weakly similar to UP|Q9LUE5 (Q9LUE5) Emb|CAB86483.1, partial (32%)
Length = 765
Score = 61.6 bits (148), Expect = 5e-11
Identities = 26/63 (41%), Positives = 38/63 (60%)
Frame = +2
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
P GC VYVG + QRF I ++N P+F LL+E E E+G G ++LPC+V F ++
Sbjct: 305 PEGCFSVYVGQQMQRFVIKTEYVNHPLFKMLLEEAESEYGYSSQGPIVLPCNVDVFYKVL 484
Query: 100 KRL 102
+
Sbjct: 485 MEM 493
>TC11175 weakly similar to GB|AAL06796.1|15809736|AY054135
AT3g60690/T4C21_100 {Arabidopsis thaliana;}, partial
(42%)
Length = 692
Score = 59.7 bits (143), Expect = 2e-10
Identities = 33/87 (37%), Positives = 45/87 (50%), Gaps = 11/87 (12%)
Frame = +1
Query: 27 ISLRRHDSS--------DPRPPSGCLYVYVGHER---QRFAIPARFLNLPVFAGLLDETE 75
+++R H S DP+ P G L VYVG E R +P + N P+F LL ETE
Sbjct: 79 LTVRHHGSGYAQLGSNLDPKVPKGHLPVYVGEEDGELHRLLVPVIYFNHPLFGELLKETE 258
Query: 76 EEFGLRGSGGLILPCDVCFFTHIVKRL 102
+EFG GG+ +PC V F + R+
Sbjct: 259 DEFGFHHPGGITIPCRVTEFERVKTRI 339
>AW719590
Length = 496
Score = 55.5 bits (132), Expect = 4e-09
Identities = 32/82 (39%), Positives = 44/82 (53%), Gaps = 1/82 (1%)
Frame = +2
Query: 21 MMRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL 80
++R S + S P G L VYVG E +RF IP +LN P F LL + EEEFG
Sbjct: 86 LIRRASFSTTQASSKGFEVPKGHLAVYVGDEMRRFVIPVSYLNQPSFQELLYQAEEEFGY 265
Query: 81 -RGSGGLILPCDVCFFTHIVKR 101
+GGL +PC F +++ +
Sbjct: 266 DHPTGGLKIPCREDDFLNLISQ 331
>BP037843
Length = 443
Score = 31.6 bits (70), Expect = 0.059
Identities = 12/40 (30%), Positives = 23/40 (57%)
Frame = -2
Query: 70 LLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNENKY 109
LL++ EEFG SG L +PC++ F +++ + ++
Sbjct: 436 LLEKAAEEFGFDHSGALTIPCEIETFKYLLNCMENTHQQH 317
>TC18755 similar to UP|AAN85378 (AAN85378) Resistance protein (Fragment),
complete
Length = 880
Score = 29.3 bits (64), Expect = 0.29
Identities = 18/72 (25%), Positives = 32/72 (44%)
Frame = -3
Query: 19 QVMMRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEF 78
+V ++ I H SS P P Y+ + + + P + L PV+A + ++
Sbjct: 350 EVFYHYRIIIYLLHQSSFPHPSHPSNRNYLAYRTRFYQTPHQLLFTPVYAH*ISVVQQ-- 177
Query: 79 GLRGSGGLILPC 90
+G LI+PC
Sbjct: 176 --KGISSLIIPC 147
>BP072991
Length = 411
Score = 28.9 bits (63), Expect = 0.38
Identities = 16/63 (25%), Positives = 30/63 (47%), Gaps = 3/63 (4%)
Frame = +3
Query: 3 GAMKKVDK---IRQIVRLKQVMMRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPA 59
GA+K + I++++++ +++ Q+S + YVY GH QRF I
Sbjct: 189 GALKYTSNQIIVEPILKIRRLFIKYTQLSKTLGSKTSQSAIFTLSYVYQGHRGQRFVIIF 368
Query: 60 RFL 62
F+
Sbjct: 369 HFI 377
>TC12423 similar to PIR|T01558|T01558 auxin-induced protein homolog
A_TM018A10.6 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (43%)
Length = 317
Score = 28.5 bits (62), Expect = 0.50
Identities = 12/20 (60%), Positives = 15/20 (75%)
Frame = +1
Query: 61 FLNLPVFAGLLDETEEEFGL 80
++N P+F LL E EEEFGL
Sbjct: 205 YINHPLFMHLLKEAEEEFGL 264
>TC17207
Length = 751
Score = 26.9 bits (58), Expect = 1.5
Identities = 13/36 (36%), Positives = 18/36 (49%)
Frame = +3
Query: 88 LPCDVCFFTHIVKRLHKNENKYGKLTLEQFVITFSD 123
L C C T++ K LHKN G + ++ FSD
Sbjct: 432 LKCPSCV-TNLFKPLHKNSRHLGTFEVR*IILNFSD 536
>BP085090
Length = 278
Score = 26.9 bits (58), Expect = 1.5
Identities = 14/44 (31%), Positives = 22/44 (49%)
Frame = -2
Query: 70 LLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNENKYGKLT 113
LL++ +EFG G ++ C V F +V + N +G LT
Sbjct: 187 LLEKAYDEFGFEPRNGRVVLCSVSAFQEVVDAIECN---HGSLT 65
>TC19254 weakly similar to UP|Q94C33 (Q94C33) AT3g51180/F24M12_220, partial
(4%)
Length = 502
Score = 26.6 bits (57), Expect = 1.9
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = -3
Query: 34 SSDPRPPSGCLYVYVGHERQRFAIP 58
+SDPRPP+ C++V E + F IP
Sbjct: 278 TSDPRPPAKCIFV---AELETFNIP 213
>TC9149 similar to UP|Q8LSK7 (Q8LSK7) Auxin-regulated protein, partial
(35%)
Length = 602
Score = 25.4 bits (54), Expect = 4.2
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = -1
Query: 57 IPARFLNLPVFAGLLDETEEE 77
+ A +LP+F G ++ETEEE
Sbjct: 374 VVAAVFSLPLFGGRVEETEEE 312
>CB827758
Length = 485
Score = 25.0 bits (53), Expect = 5.5
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = -1
Query: 83 SGGLILPCDVCFF 95
+G L+L CDVCFF
Sbjct: 101 TGMLLLSCDVCFF 63
>BP055291
Length = 496
Score = 24.6 bits (52), Expect = 7.2
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +3
Query: 83 SGGLILPCDVCFFTHIVKRLHKNENKYGKL 112
SGG + +CF I+ + KN+ KY K+
Sbjct: 114 SGGPKVLLSLCFM*PILNTMLKNDKKYSKI 203
>BP066384
Length = 430
Score = 24.3 bits (51), Expect = 9.4
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = -2
Query: 34 SSDPRPPSGCLYVYVGHERQR 54
SS P PP C+Y H+RQR
Sbjct: 87 SSPPPPPPRCIYT---HKRQR 34
>TC14433 similar to UP|Q41057 (Q41057) Cysteine protease , complete
Length = 1392
Score = 24.3 bits (51), Expect = 9.4
Identities = 21/68 (30%), Positives = 30/68 (43%), Gaps = 5/68 (7%)
Frame = +2
Query: 22 MRWKQISLRRHDSSDPRPPSGCLYVYVGHER-QRFAIPARFLNLPVFAGLLDETEEEFGL 80
+RW +SLRR D R P ++ + R R ++ L+ P A L D + L
Sbjct: 101 LRWCSVSLRRQDRPASRTPIQFVWSQIWRSRFFR*SVKLAMLS-PSHASLPDTASDTIPL 277
Query: 81 R----GSG 84
R GSG
Sbjct: 278 RRFSIGSG 301
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.141 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,622,282
Number of Sequences: 28460
Number of extensions: 33959
Number of successful extensions: 233
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 232
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 232
length of query: 148
length of database: 4,897,600
effective HSP length: 82
effective length of query: 66
effective length of database: 2,563,880
effective search space: 169216080
effective search space used: 169216080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0004a.3


