

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0002.6
(260 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
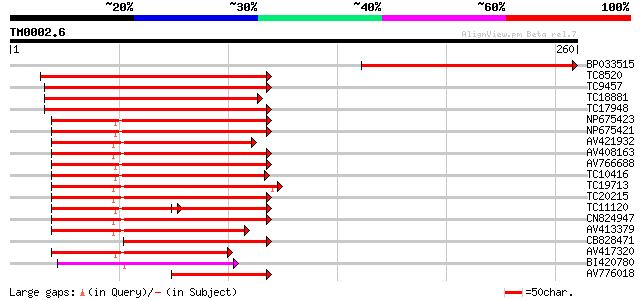
Score E
Sequences producing significant alignments: (bits) Value
BP033515 198 7e-52
TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 181 1e-46
TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 176 3e-45
TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose... 173 3e-44
TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose... 162 4e-41
NP675423 transcription factor MYB101 [Lotus japonicus] 109 5e-25
NP675421 transcription factor MYB103 [Lotus japonicus] 107 2e-24
AV421932 107 2e-24
AV408163 106 4e-24
AV766688 105 8e-24
TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial... 104 2e-23
TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%) 102 7e-23
TC20215 similar to AAS10104 (AAS10104) MYB transcription factor,... 102 7e-23
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 62 4e-21
CN824947 95 1e-20
AV413379 93 5e-20
CB828471 80 5e-16
AV417320 79 6e-16
BI420780 68 1e-12
AV776018 55 1e-08
>BP033515
Length = 464
Score = 198 bits (504), Expect = 7e-52
Identities = 99/99 (100%), Positives = 99/99 (100%)
Frame = -2
Query: 162 LMDQGYPFKRHCVEKDDEESRDGVEEAETMTEEEVLASSVPTSLSLFPMGEKKMVAVAEK 221
LMDQGYPFKRHCVEKDDEESRDGVEEAETMTEEEVLASSVPTSLSLFPMGEKKMVAVAEK
Sbjct: 463 LMDQGYPFKRHCVEKDDEESRDGVEEAETMTEEEVLASSVPTSLSLFPMGEKKMVAVAEK 284
Query: 222 EEKKMDMQEGLMRTMMQRMIAEEVRAYFDKMRFDPRLQK 260
EEKKMDMQEGLMRTMMQRMIAEEVRAYFDKMRFDPRLQK
Sbjct: 283 EEKKMDMQEGLMRTMMQRMIAEEVRAYFDKMRFDPRLQK 167
>TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (36%)
Length = 1778
Score = 181 bits (459), Expect = 1e-46
Identities = 81/106 (76%), Positives = 91/106 (85%)
Frame = +1
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
+ DR+KG WSP+EDE L KLV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP V+HR
Sbjct: 169 EMDRIKGPWSPEEDEALQKLVERHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPQVEHRA 348
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
FTP ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 349 FTPEEDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRK 486
>TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (43%)
Length = 860
Score = 176 bits (447), Expect = 3e-45
Identities = 79/104 (75%), Positives = 89/104 (84%)
Frame = +3
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WSP+EDE L +LV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP V+HR FT
Sbjct: 69 DRIKGPWSPEEDEALQRLVEKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPQVEHRAFT 248
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 249 AEEDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRK 380
>TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose-responsive
element binding factor, partial (28%)
Length = 346
Score = 173 bits (438), Expect = 3e-44
Identities = 76/100 (76%), Positives = 87/100 (87%)
Frame = +1
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DRVKG WSP+EDE L +LV HGP+NW+ IS IPGRSGKSCRLRWCNQLSP+V+HRPFT
Sbjct: 46 DRVKGPWSPEEDEALTRLVQVHGPKNWTAISKSIPGRSGKSCRLRWCNQLSPEVEHRPFT 225
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNST 116
P ED+ II+AHA GNKWATI+R+L GRTDNA+KNHWNST
Sbjct: 226 PEEDSTIIRAHARCGNKWATIARILNGRTDNAVKNHWNST 345
>TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (50%)
Length = 487
Score = 162 bits (411), Expect = 4e-41
Identities = 73/104 (70%), Positives = 85/104 (81%)
Frame = +2
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
+R+KG WS +ED L +LV +HG RNWS+IS I GRSGKSCRLRWCNQLSP V+HRPF+
Sbjct: 161 ERIKGPWSAEEDRILTRLVERHGARNWSLISRYIKGRSGKSCRLRWCNQLSPTVEHRPFS 340
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED II AHA GN+WA I+RLLPGRTDNA+KNHWNSTL+RR
Sbjct: 341 VQEDETIIAAHARFGNRWAAIARLLPGRTDNAVKNHWNSTLKRR 472
>NP675423 transcription factor MYB101 [Lotus japonicus]
Length = 1047
Score = 109 bits (272), Expect = 5e-25
Identities = 51/103 (49%), Positives = 69/103 (66%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+P+ED L+ + +HG +W + AG+ R GKSCRLRW N L PD++ FT
Sbjct: 40 KGPWTPEEDAILVDYIQKHGHGSWRALPKLAGL-NRCGKSCRLRWTNYLRPDIKRGKFTE 216
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ +II HAV GNKW+ I+ LPGRTDN IKN WN+ L+++
Sbjct: 217 EEEQLIINLHAVLGNKWSAIAGHLPGRTDNEIKNFWNTHLKKK 345
>NP675421 transcription factor MYB103 [Lotus japonicus]
Length = 924
Score = 107 bits (267), Expect = 2e-24
Identities = 51/103 (49%), Positives = 69/103 (66%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG WS +ED+ L+ + +HG +W + AG+ R GKSCRLRW N L PD++ F+
Sbjct: 40 KGPWSQEEDKVLVDYIQKHGHGSWRALPQLAGL-NRCGKSCRLRWTNYLRPDIKRGKFSD 216
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ +II HA GNKWATI+ LPGRTDN IKN WN+ L+++
Sbjct: 217 EEEQLIINLHASLGNKWATIASHLPGRTDNEIKNLWNTHLKKK 345
>AV421932
Length = 402
Score = 107 bits (267), Expect = 2e-24
Identities = 51/96 (53%), Positives = 65/96 (67%), Gaps = 2/96 (2%)
Frame = +3
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
+G WSP+EDE L++ + HG WS + AG+ R GKSCRLRW N L PD++ FTP
Sbjct: 117 RGLWSPEEDEKLIRYITTHGYGCWSEVPEKAGLQ-RCGKSCRLRWINYLRPDIRRGRFTP 293
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHW 113
E+ +II H V GN+WA I+ LPGRTDN IKN+W
Sbjct: 294 EEEKLIISLHGVVGNRWAHIASHLPGRTDNEIKNYW 401
>AV408163
Length = 425
Score = 106 bits (265), Expect = 4e-24
Identities = 50/103 (48%), Positives = 71/103 (68%), Gaps = 2/103 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG+W+ +ED+ L+ + HG W + +AG+ R GKSCRLRW N L PD++ FT
Sbjct: 116 KGAWTKEEDDRLISYIRAHGEGCWRSLPKAAGLL-RCGKSCRLRWINYLRPDLKRGNFTE 292
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED +II+ H++ GNKW+ I+ LPGRTDN IKN+WN+ +RR+
Sbjct: 293 EEDELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRK 421
>AV766688
Length = 608
Score = 105 bits (262), Expect = 8e-24
Identities = 50/103 (48%), Positives = 70/103 (67%), Gaps = 2/103 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG+W+ +ED L+ V ++G NW + AG+ R GKSCRLRW N L PD++ FT
Sbjct: 116 KGTWTAEEDRKLIAYVTRYGCWNWRQLPKFAGL-ARCGKSCRLRWLNYLRPDIKRGHFTQ 292
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ III+ H GN+W+ I+ LPGRTDN +KNHW+++L+RR
Sbjct: 293 EEEEIIIRMHKNLGNRWSIIAAELPGRTDNEVKNHWHTSLKRR 421
Score = 34.7 bits (78), Expect = 0.017
Identities = 17/55 (30%), Positives = 31/55 (55%)
Frame = +2
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQ 71
D +G ++ +E+E ++++ G R WS+I+A +PGR+ + W L VQ
Sbjct: 266 DIKRGHFTQEEEEIIIRMHKNLGNR-WSIIAAELPGRTDNEVKNHWHTSLKRRVQ 427
>TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial (41%)
Length = 1046
Score = 104 bits (259), Expect = 2e-23
Identities = 49/102 (48%), Positives = 71/102 (69%), Gaps = 2/102 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG+W+ +ED+ L+ V ++G NW + AG+ R GKSCRLRW N L P+++ FT
Sbjct: 131 KGTWTAEEDKKLIAYVTRYGCWNWRQLPKFAGL-ARCGKSCRLRWLNYLRPNIKRGNFTQ 307
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRR 119
E+ III+ H GN+W+TI+ LPGRTD+ +KNHW++TL+R
Sbjct: 308 EEEEIIIRMHKKLGNRWSTIAAELPGRTDSEVKNHWHTTLKR 433
Score = 35.4 bits (80), Expect = 0.010
Identities = 17/62 (27%), Positives = 34/62 (54%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
+G+++ +E+E ++++ + G R WS I+A +PGR+ + W L V + E
Sbjct: 290 RGNFTQEEEEIIIRMHKKLGNR-WSTIAAELPGRTDSEVKNHWHTTLKRTVHEQNTVTNE 466
Query: 80 DT 81
+T
Sbjct: 467 ET 472
>TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%)
Length = 532
Score = 102 bits (254), Expect = 7e-23
Identities = 50/110 (45%), Positives = 72/110 (65%), Gaps = 4/110 (3%)
Frame = +3
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+ +ED L+ + HG W+ + SAG+ R+GKSCRLRW N L PDV+ TP
Sbjct: 111 KGPWTIEEDLILINYIANHGEGVWNTLAKSAGLK-RTGKSCRLRWLNYLRPDVRRGNITP 287
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRR--RRGESL 125
E +I++ HA GN+W+ I++ LPGRTDN IKN W + +++ ++ ESL
Sbjct: 288 EEQLLIMELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQKHIKQAESL 437
>TC20215 similar to AAS10104 (AAS10104) MYB transcription factor, partial
(35%)
Length = 464
Score = 102 bits (254), Expect = 7e-23
Identities = 50/103 (48%), Positives = 68/103 (65%), Gaps = 2/103 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
+G WS +ED+ L + Q+G +W + +AG+ R GKSCRLRW N L DV+ T
Sbjct: 68 RGRWSEEEDKILTDFIKQNGEGSWKSLPKNAGLL-RCGKSCRLRWINYLREDVKRGNITS 244
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ II++ HA GN+W+ I+ LPGRTDN IKN+WNS LRR+
Sbjct: 245 EEEEIIVKLHAALGNRWSVIAGHLPGRTDNEIKNYWNSHLRRK 373
Score = 32.7 bits (73), Expect = 0.064
Identities = 17/50 (34%), Positives = 29/50 (58%)
Frame = +2
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQL 66
D +G+ + +E+E ++KL G R WSVI+ +PGR+ + W + L
Sbjct: 218 DVKRGNITSEEEEIIVKLHAALGNR-WSVIAGHLPGRTDNEIKNYWNSHL 364
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 62.0 bits (149), Expect(2) = 4e-21
Identities = 28/62 (45%), Positives = 41/62 (65%), Gaps = 2/62 (3%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+P+ED+ L++ + +HG +W + AG+ R GKSCRLRW N L PD++ F+P
Sbjct: 106 KGPWTPEEDQKLVEHIQKHGHGSWRALPKLAGL-NRCGKSCRLRWTNYLRPDIKRGKFSP 282
Query: 78 TE 79
E
Sbjct: 283 EE 288
Score = 55.1 bits (131), Expect(2) = 4e-21
Identities = 22/46 (47%), Positives = 33/46 (70%)
Frame = +2
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
F ++ I+ H++ GNKW+TI+ LPGRTDN IKN WN+ L+++
Sbjct: 275 FLQKKEQTILHLHSILGNKWSTIATHLPGRTDNEIKNFWNTHLKKK 412
Score = 26.9 bits (58), Expect = 3.5
Identities = 14/57 (24%), Positives = 28/57 (48%)
Frame = +2
Query: 21 GSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
G++ ++++T++ L G + WS I+ +PGR+ + W L + F P
Sbjct: 269 GNFLQKKEQTILHLHSILGNK-WSTIATHLPGRTDNEIKNFWNTHLKKKLIQMGFDP 436
>CN824947
Length = 738
Score = 95.1 bits (235), Expect = 1e-20
Identities = 45/103 (43%), Positives = 67/103 (64%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
+G W+ +EDE L K + G +W + +AG+ R GKSCRLRW N L D++ +
Sbjct: 190 RGRWTAEEDELLTKYIQASGEGSWRSLPKNAGLL-RCGKSCRLRWINYLRGDLKRGNISD 366
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+++I++ HA GN+W+ I+ +PGRTDN IKN+WNS L R+
Sbjct: 367 EEESLIVKLHASFGNRWSLIASHMPGRTDNEIKNYWNSHLSRK 495
>AV413379
Length = 416
Score = 92.8 bits (229), Expect = 5e-20
Identities = 46/93 (49%), Positives = 62/93 (66%), Gaps = 2/93 (2%)
Frame = +3
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG+W+ +ED+ L+ + HG W + SAG+ R GKSCRLRW N L PD++ FTP
Sbjct: 141 KGAWTKEEDDRLIAYIRAHGEGCWRSLPKSAGLL-RCGKSCRLRWINYLRPDLKRGNFTP 317
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIK 110
ED +II+ H++ GNKW+ I+ L GRTDN IK
Sbjct: 318 QEDELIIKLHSLLGNKWSLIAGRLAGRTDNEIK 416
Score = 31.6 bits (70), Expect = 0.14
Identities = 15/38 (39%), Positives = 25/38 (65%)
Frame = +3
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRS 54
D +G+++PQEDE ++KL G WS+I+ + GR+
Sbjct: 291 DLKRGNFTPQEDELIIKLHSLLG-NKWSLIAGRLAGRT 401
>CB828471
Length = 525
Score = 79.7 bits (195), Expect = 5e-16
Identities = 35/68 (51%), Positives = 47/68 (68%)
Frame = +1
Query: 53 RSGKSCRLRWCNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNH 112
R GKSCRLRW N L PD++ F+ E+ II H + GN+W+ I+ LPGRTDN IKN
Sbjct: 22 RCGKSCRLRWANYLRPDIKRGNFSREEEDEIINLHELLGNRWSAIAARLPGRTDNEIKNV 201
Query: 113 WNSTLRRR 120
W++ L++R
Sbjct: 202 WHTHLKKR 225
Score = 31.2 bits (69), Expect = 0.19
Identities = 17/55 (30%), Positives = 30/55 (53%)
Frame = +1
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQ 71
D +G++S +E++ ++ L G R WS I+A +PGR+ + W L +Q
Sbjct: 70 DIKRGNFSREEEDEIINLHELLGNR-WSAIAARLPGRTDNEIKNVWHTHLKKRLQ 231
>AV417320
Length = 369
Score = 79.3 bits (194), Expect = 6e-16
Identities = 37/85 (43%), Positives = 54/85 (63%), Gaps = 2/85 (2%)
Frame = +3
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+ +ED+ L+ + +HG NW + AG+ R GKSCRLRW N L PD++ FT
Sbjct: 117 KGPWTSEEDQILISYIQKHGHGNWRALPKHAGLL-RCGKSCRLRWINYLRPDIKRGNFTA 293
Query: 78 TEDTIIIQAHAVHGNKWATISRLLP 102
E+ +II+ H + GN+W+ I+ LP
Sbjct: 294 EEEELIIKMHELLGNRWSAIAAKLP 368
>BI420780
Length = 475
Score = 68.2 bits (165), Expect = 1e-12
Identities = 34/86 (39%), Positives = 47/86 (54%), Gaps = 3/86 (3%)
Frame = +1
Query: 23 WSPQEDETLMKLVGQHGPRNWSVISAGIP---GRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
WS +ED L V Q+GPR W+++S + R KSC RW N L P ++ T E
Sbjct: 211 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 390
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRT 105
++I A +GNKW I+ +PGRT
Sbjct: 391 QRLVILLQANYGNKWKKIAAEVPGRT 468
>AV776018
Length = 476
Score = 55.1 bits (131), Expect = 1e-08
Identities = 24/46 (52%), Positives = 32/46 (69%)
Frame = +1
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
F ED +II+ HAV GN+W+ + PGRTDN IKN WNS L+++
Sbjct: 13 FHKEEDNLIIELHAVLGNRWSQDATQWPGRTDNEIKNLWNSCLKKK 150
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,724,726
Number of Sequences: 28460
Number of extensions: 90236
Number of successful extensions: 2903
Number of sequences better than 10.0: 272
Number of HSP's better than 10.0 without gapping: 1882
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2604
length of query: 260
length of database: 4,897,600
effective HSP length: 88
effective length of query: 172
effective length of database: 2,393,120
effective search space: 411616640
effective search space used: 411616640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0002.6


