

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
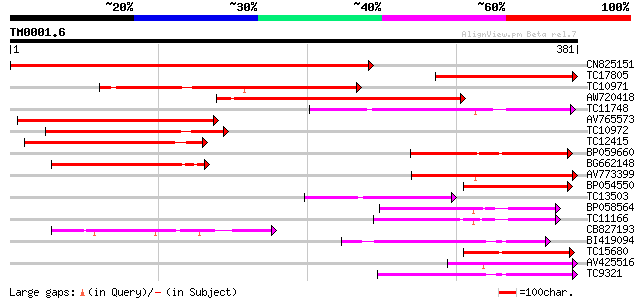
Query= TM0001.6
(381 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CN825151 503 e-143
TC17805 weakly similar to UP|O82681 (O82681) Lanatoside 15'-O-ac... 197 3e-51
TC10971 similar to UP|Q40318 (Q40318) Coil protein, partial (42%) 167 3e-42
AW720418 164 2e-41
TC11748 weakly similar to UP|O82076 (O82076) Lipase homolog (Fra... 137 4e-33
AV765573 134 3e-32
TC10972 similar to UP|Q40318 (Q40318) Coil protein, partial (32%) 133 4e-32
TC12415 similar to UP|Q9LII9 (Q9LII9) Genomic DNA, chromosome 3,... 126 5e-30
BP059660 125 1e-29
BG662148 117 4e-27
AV773399 116 5e-27
BP054550 92 1e-19
TC13503 similar to UP|Q8W0Y7 (Q8W0Y7) Enod8.3 (Fragment), partia... 82 1e-16
BP058564 79 2e-15
TC11166 weakly similar to UP|Q9FXJ3 (Q9FXJ3) F1K23.17, partial (... 77 6e-15
CB827193 76 8e-15
BI419094 75 1e-14
TC15680 weakly similar to UP|Q7Y1X1 (Q7Y1X1) ENSP-like protein, ... 75 1e-14
AV425516 71 3e-13
TC9321 similar to UP|O04320 (O04320) Proline-rich protein APG is... 68 2e-12
>CN825151
Length = 740
Score = 503 bits (1294), Expect = e-143
Identities = 244/244 (100%), Positives = 244/244 (100%)
Frame = +1
Query: 1 MGLRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGE 60
MGLRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGE
Sbjct: 7 MGLRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGE 186
Query: 61 SFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETI 120
SFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETI
Sbjct: 187 SFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETI 366
Query: 121 FQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDL 180
FQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDL
Sbjct: 367 FQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDL 546
Query: 181 SVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLP 240
SVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLP
Sbjct: 547 SVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLP 726
Query: 241 AGYL 244
AGYL
Sbjct: 727 AGYL 738
>TC17805 weakly similar to UP|O82681 (O82681) Lanatoside
15'-O-acetylesterase precursor, partial (12%)
Length = 484
Score = 197 bits (500), Expect = 3e-51
Identities = 89/95 (93%), Positives = 92/95 (96%)
Frame = +3
Query: 287 AAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWD 346
AAKYGLIS+T++EGFVDPL IC GYHVNDTHIW GTIGTANGKDVFGNACEKPSMYVSWD
Sbjct: 3 AAKYGLISHTQHEGFVDPLPICFGYHVNDTHIWFGTIGTANGKDVFGNACEKPSMYVSWD 182
Query: 347 GVHYAEAANHWVANRILNGSFTDPPTLITQACYRH 381
GVHYAEAANHWVANRILNGSFTDPPTLITQACYRH
Sbjct: 183 GVHYAEAANHWVANRILNGSFTDPPTLITQACYRH 287
>TC10971 similar to UP|Q40318 (Q40318) Coil protein, partial (42%)
Length = 615
Score = 167 bits (422), Expect = 3e-42
Identities = 88/181 (48%), Positives = 119/181 (65%), Gaps = 5/181 (2%)
Frame = +2
Query: 61 SFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETI 120
+F Q PS +GRLI+DFI E+L +PYLSAYLNS+G+NYRHGANFA GG++IR
Sbjct: 98 TFIQIPSG---NGRLIIDFITEELEIPYLSAYLNSIGSNYRHGANFAAGGASIRPV---- 256
Query: 121 FQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALE----RSKLPVPEEFSKALYTFDIG 176
YG SPF L MQ QF Q ++ + L + + +S LP PE+FSKALYT DIG
Sbjct: 257 --YGFSPFYLGMQVAQFIQLQSHIENLLNQFSSNRTEPPFKSYLPRPEDFSKALYTIDIG 430
Query: 177 QNDLSVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPV-NLFY 235
QNDL G + +++ S+P+++ V+ +Y++G R FWIHNT PIGCLP ++FY
Sbjct: 431 QNDLGFGLMHTSEEEVLRSIPEMMRNFTYDVQVLYDVGARVFWIHNTCPIGCLPTSSIFY 610
Query: 236 K 236
+
Sbjct: 611 E 613
>AW720418
Length = 500
Score = 164 bits (415), Expect = 2e-41
Identities = 83/167 (49%), Positives = 114/167 (67%)
Frame = +3
Query: 140 FKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDI 199
FK RT+ L+ + S +P E+FS+ALY FDIGQNDLS GF+ + +Q+R S+P+I
Sbjct: 3 FKLRTQILFN-GTV*IFXSSIPRXEDFSRALYMFDIGQNDLSYGFQYSSEEQLRASLPNI 179
Query: 200 VNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVE 259
++Q + AV+ +Y G R FWIHNT PIGCLP+N G +D GCV QN +A E
Sbjct: 180 LSQFSQAVEQLYXEGARVFWIHNTGPIGCLPLNFLAYGKSKKGNVDANGCVISQNGIAHE 359
Query: 260 FNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLK 306
FN QLK++V++LR +LP A + YVD+Y AKY L+SN + GFV+PL+
Sbjct: 360 FNLQLKNQVLQLRKKLPLAKLIYVDVYKAKYELVSNARKLGFVNPLE 500
>TC11748 weakly similar to UP|O82076 (O82076) Lipase homolog (Fragment),
partial (48%)
Length = 679
Score = 137 bits (344), Expect = 4e-33
Identities = 71/183 (38%), Positives = 103/183 (55%), Gaps = 4/183 (2%)
Frame = +3
Query: 202 QLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFN 261
++ +AVK++Y G R FW+HNT P+GCLP L LD +GC+ N A FN
Sbjct: 6 EIENAVKSLYNEGARKFWVHNTGPLGCLPKILALAQKKD---LDLFGCLSSYNSAARLFN 176
Query: 262 KQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY----HVNDTH 317
+ L KLRT L +A + YVD+YA K+ LI+N GF +PL +CCG+ + D
Sbjct: 177 EALYHSSQKLRTRLKDATLVYVDIYAIKHDLIANATKYGFSNPLTVCCGFGGPPYNFDLR 356
Query: 318 IWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQA 377
+ CG G C++ S YV+WDG H+ EAAN ++A++IL+ ++ P T
Sbjct: 357 VTCGQPGY--------QVCDEGSRYVNWDGTHHTEAANTFIASKILSTDYSTPRTPFEFF 512
Query: 378 CYR 380
C+R
Sbjct: 513 CHR 521
>AV765573
Length = 440
Score = 134 bits (336), Expect = 3e-32
Identities = 69/135 (51%), Positives = 88/135 (65%)
Frame = +2
Query: 6 LFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQK 65
LF F + L + V + C FPAI+NFG SNSDTGG +AAF PP G+++ +
Sbjct: 32 LFALLFTTVLLNPDVVFATTTKTCDFPAIFNFGASNSDTGGFAAAFLQPGPPQGDTYFGR 211
Query: 66 PSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGI 125
P+ R DGR+I+DFIA+ LPYLSAYLNSLG NY HGA+FAT STIR +
Sbjct: 212 PAGRFSDGRIILDFIAQNFELPYLSAYLNSLGANYSHGASFATLSSTIRLPESVVPNGRS 391
Query: 126 SPFSLDMQFVQFKQF 140
SPF L +Q++QF+QF
Sbjct: 392 SPFFLGLQYLQFQQF 436
>TC10972 similar to UP|Q40318 (Q40318) Coil protein, partial (32%)
Length = 548
Score = 133 bits (335), Expect = 4e-32
Identities = 68/123 (55%), Positives = 85/123 (68%)
Frame = +1
Query: 25 NSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKL 84
+S C++PA+YNFGDSNS+TG + AAF + P G SF S R DGRLI+DFI E+L
Sbjct: 196 SSSKCSYPAVYNFGDSNSNTGVVYAAFAGLQSPGGISFFGNLSGRASDGRLIIDFITEEL 375
Query: 85 NLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKART 144
+PYLSAYLNS+G+NYRHGANFA GG++IR YG SPF L MQ QF Q ++
Sbjct: 376 EIPYLSAYLNSIGSNYRHGANFAAGGASIRP------VYGFSPFYLGMQVAQFIQLQSHI 537
Query: 145 KQL 147
+ L
Sbjct: 538 ENL 546
>TC12415 similar to UP|Q9LII9 (Q9LII9) Genomic DNA, chromosome 3, TAC
clone:K24A2, partial (22%)
Length = 366
Score = 126 bits (317), Expect = 5e-30
Identities = 67/123 (54%), Positives = 82/123 (66%)
Frame = +3
Query: 11 FLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARD 70
FL T+ + + + + +PAI+NFGDSNSDTG ISAAF + PP G++F S R
Sbjct: 18 FLPFTIVIVAAGVSFTEGSQYPAIFNFGDSNSDTGAISAAFTIVHPPNGQNFLGALSGRY 197
Query: 71 CDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSL 130
DGRLI+DFI E+L LPYL AYLNS+G NYRHGANFATGGS +I + G SPF L
Sbjct: 198 SDGRLIIDFITEELKLPYLDAYLNSVGANYRHGANFATGGS-------SILKGGYSPFHL 356
Query: 131 DMQ 133
Q
Sbjct: 357 AYQ 365
>BP059660
Length = 501
Score = 125 bits (314), Expect = 1e-29
Identities = 59/109 (54%), Positives = 74/109 (67%)
Frame = -3
Query: 270 KLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIGTANGK 329
+LR LP+A TYVD+Y AKY LISN +GFV+PL++CCG + I CG NG
Sbjct: 496 QLRRNLPKAKFTYVDVYTAKYELISNASKQGFVNPLEVCCGSYYG-YRIDCGKKAVVNGT 320
Query: 330 DVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQAC 378
V+GN C+ PS ++SWDGVHY +AAN WVA I +GS +DPP I QAC
Sbjct: 319 -VYGNPCKNPSQHISWDGVHYTQAANKWVAKHIRDGSLSDPPVPIGQAC 176
>BG662148
Length = 415
Score = 117 bits (292), Expect = 4e-27
Identities = 60/106 (56%), Positives = 78/106 (72%)
Frame = +2
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI+NFG SNSDTGG +AAFE PP+G+++ +P+ R DGR+I+DFIA+ L Y
Sbjct: 104 CDFPAIFNFGASNSDTGGYAAAFEGPKPPHGDTYFHRPAGRYSDGRIIIDFIAQSFGLTY 283
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQF 134
LSAYL+SLG+N+ HGANFAT GSTI + + G SPF L +Q+
Sbjct: 284 LSAYLDSLGSNFSHGANFATLGSTIILP-KNVKPRG-SPFFLGVQY 415
>AV773399
Length = 492
Score = 116 bits (291), Expect = 5e-27
Identities = 54/115 (46%), Positives = 77/115 (66%), Gaps = 4/115 (3%)
Frame = -1
Query: 271 LRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY---HVNDTHIWCGTIGTAN 327
LR +L A+ITYVD+Y+ KY LIS K F +PL+ CCG+ + HI CG +
Sbjct: 492 LRKKLHSASITYVDVYSVKYSLISQAKTHAFGEPLRACCGHGGKYNYYIHIGCGAKLKVH 313
Query: 328 GKDVF-GNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYRH 381
G+++ G CE PS+ V+WDGVH+ EAAN WV ++I++GSF+DPP + AC++H
Sbjct: 312 GQEILLGKPCEDPSLSVNWDGVHFTEAANKWVFDQIVDGSFSDPPIPLNMACHKH 148
>BP054550
Length = 417
Score = 92.4 bits (228), Expect = 1e-19
Identities = 37/73 (50%), Positives = 51/73 (69%)
Frame = -3
Query: 306 KICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNG 365
+ICCGY+ + + CG NGK+++ CE PS Y+ WDGV Y EAAN W+A+RIL+G
Sbjct: 415 EICCGYYKDGHPVNCGKKEIINGKEIYAGPCEDPSQYIRWDGVPYPEAANRWIAHRILHG 236
Query: 366 SFTDPPTLITQAC 378
SF+DPP +T +C
Sbjct: 235 SFSDPPLPLTHSC 197
>TC13503 similar to UP|Q8W0Y7 (Q8W0Y7) Enod8.3 (Fragment), partial (30%)
Length = 705
Score = 82.4 bits (202), Expect = 1e-16
Identities = 45/102 (44%), Positives = 60/102 (58%)
Frame = -2
Query: 199 IVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAV 258
++N+ +K IY+L R+F IHNT PIGCLP L N P D YGC N +A
Sbjct: 416 VLNE*NMVIKLIYDLRARSFGIHNT*PIGCLPAILT---NFPKAERDHYGCAT*YNEVAK 246
Query: 259 EFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEG 300
FN++LK+ + +LR ELP AAI YVD+Y+ K N + G
Sbjct: 245 YFNQKLKEALAQLRRELPLAAIIYVDIYSPKLDHFRNLEKYG 120
>BP058564
Length = 468
Score = 78.6 bits (192), Expect = 2e-15
Identities = 46/127 (36%), Positives = 65/127 (50%), Gaps = 5/127 (3%)
Frame = -2
Query: 249 CVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFV-DPLKI 307
C+K N+ + L+ + +LR P I Y D + A + + +N GF + LK+
Sbjct: 467 CLKWLNMFHEHHDDLLQIELNQLRVLYPLTNIIYADFFNAALQIYKSPENFGFGGNALKV 288
Query: 308 CCG----YHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRIL 363
CCG Y+ NDT + CG G AC+ PS YVSWDG H EAA W+ +L
Sbjct: 287 CCGGGGPYNYNDTAV-CGNSGVI--------ACDDPSQYVSWDGYHLTEAAYRWITKGLL 135
Query: 364 NGSFTDP 370
+GS+T P
Sbjct: 134 DGSYTTP 114
>TC11166 weakly similar to UP|Q9FXJ3 (Q9FXJ3) F1K23.17, partial (15%)
Length = 495
Score = 76.6 bits (187), Expect = 6e-15
Identities = 47/130 (36%), Positives = 69/130 (52%), Gaps = 4/130 (3%)
Frame = +2
Query: 245 DPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDP 304
D GCVK N A +N++L+ + +LR P A I Y D Y A L N + GF
Sbjct: 2 DQAGCVKWLNEFAEFYNQKLQLEIHRLRELHPHANIIYADYYNAALPLYQNPQKFGFRG- 178
Query: 305 LKICCG----YHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVAN 360
L+ICCG Y+ N + CG+ G AC+ PS Y+ WDG+H EAA ++++
Sbjct: 179 LEICCGMGGPYNYNAS-AGCGSPGVI--------ACDDPSEYIGWDGIHLTEAAYGFISD 331
Query: 361 RILNGSFTDP 370
++NG ++ P
Sbjct: 332 GLMNGPYSFP 361
>CB827193
Length = 528
Score = 76.3 bits (186), Expect = 8e-15
Identities = 61/168 (36%), Positives = 84/168 (49%), Gaps = 17/168 (10%)
Frame = +1
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIP------PPYGESFPQKPSARDCDGRLIVDFIAE 82
C+F +I++FGDS +DTG + + +P PPYG ++ +PSAR DGR+I+D+IAE
Sbjct: 67 CSFSSIFSFGDSLADTGNLYLS-SALPSHHCFFPPYGRTYFHRPSARCSDGRIIIDYIAE 243
Query: 83 KLNLPYLSAYLNSL------GTNYRHGANFATGGSTIRRQNETIFQ-YGIS---PFSLDM 132
L LP++ YL N GANFA G+T ET FQ GI +SL +
Sbjct: 244 SLGLPFVKPYLEIKKHGGLENFNVEGGANFAVIGAT--ALEETFFQDKGIQLPVNYSLPV 417
Query: 133 QFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTF-DIGQND 179
Q FK E +AL S E +L+ +IG ND
Sbjct: 418 QLNWFK-----------ELLSALCNSSTSCHEVLGNSLFIVGEIGGND 528
>BI419094
Length = 585
Score = 75.5 bits (184), Expect = 1e-14
Identities = 46/141 (32%), Positives = 66/141 (46%), Gaps = 1/141 (0%)
Frame = +3
Query: 224 APIGCLPVNL-FYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITY 282
AP+GCLP + + H D CV N AV FN++L L+ LP +
Sbjct: 12 APVGCLPAAITLFGH-------DSNQCVARLNNDAVNFNRKLNTTSQSLQKSLPGLKLVL 170
Query: 283 VDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMY 342
+D+Y Y L++ GF + + CCG + +T I C N K + C S Y
Sbjct: 171 LDIYQPLYDLVTKPSENGFAEARRACCGTGLLETSILC------NQKSI--GTCANASEY 326
Query: 343 VSWDGVHYAEAANHWVANRIL 363
V WDG H +EAAN +A ++
Sbjct: 327 VFWDGFHPSEAANQVLAGDLI 389
>TC15680 weakly similar to UP|Q7Y1X1 (Q7Y1X1) ENSP-like protein, partial
(7%)
Length = 641
Score = 75.5 bits (184), Expect = 1e-14
Identities = 34/74 (45%), Positives = 48/74 (63%)
Frame = +1
Query: 306 KICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNG 365
+ CCG ++ I CG T +G V C+ PS ++SWDGVHY+E AN +A +IL+G
Sbjct: 1 EFCCGNYLGGYKISCGQNTTESGT-VSDKTCKNPSEFLSWDGVHYSEEANLLIAKQILSG 177
Query: 366 SFTDPPTLITQACY 379
+F+DPP I QAC+
Sbjct: 178 AFSDPPVTIRQACF 219
>AV425516
Length = 415
Score = 70.9 bits (172), Expect = 3e-13
Identities = 34/90 (37%), Positives = 52/90 (57%), Gaps = 3/90 (3%)
Frame = -3
Query: 295 NTKNEGFVDPLKICCGYHVNDTH---IWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYA 351
+++ +GF PL CCGY + + CG +G+++F +CE PS V WDG HY
Sbjct: 404 HSEYQGFEIPLINCCGYGGKYNYTDGVACGQNIKVDGEEIFVGSCESPSTKVIWDGTHYT 225
Query: 352 EAANHWVANRILNGSFTDPPTLITQACYRH 381
EAAN V + I G+F+DPP + +C ++
Sbjct: 224 EAANKIVFDLISTGAFSDPPIPLNMSCSKN 135
>TC9321 similar to UP|O04320 (O04320) Proline-rich protein APG isolog,
partial (37%)
Length = 541
Score = 68.2 bits (165), Expect = 2e-12
Identities = 45/137 (32%), Positives = 62/137 (44%), Gaps = 3/137 (2%)
Frame = +2
Query: 248 GCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKI 307
GCV N A FNK++ KL+ + P I D+Y Y L+ N GF + +
Sbjct: 8 GCVSRFNTDAQAFNKKINSAAAKLQKQHPGLKIVIFDIYKPLYDLVQTPSNFGFAEARRG 187
Query: 308 CCGY-HVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANR-ILNG 365
CCG V T + C N K + C + YV WD VH +EAAN +A+ IL G
Sbjct: 188 CCGTGTVETTSLLC------NPKSI--GTCSNATQYVFWDSVHPSEAANQVLADALILQG 343
Query: 366 -SFTDPPTLITQACYRH 381
+ ++ CY H
Sbjct: 344 IALIG*GLIVIYLCYNH 394
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,912,444
Number of Sequences: 28460
Number of extensions: 98899
Number of successful extensions: 576
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 541
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 544
length of query: 381
length of database: 4,897,600
effective HSP length: 92
effective length of query: 289
effective length of database: 2,279,280
effective search space: 658711920
effective search space used: 658711920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0001.6


