

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0627.15
(557 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
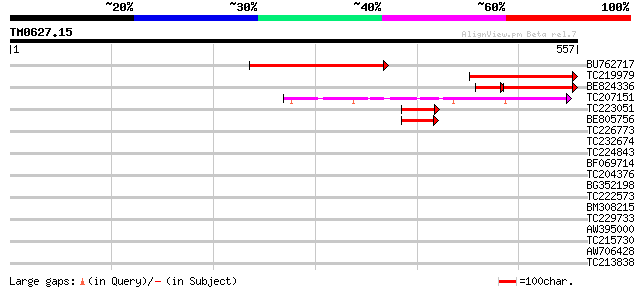
Score E
Sequences producing significant alignments: (bits) Value
BU762717 249 2e-66
TC219979 similar to UP|O23240 (O23240) Actin interacting protein... 180 1e-45
BE824336 similar to GP|16604326|gb| At4g36400/C7A10_960 {Arabido... 124 1e-36
TC207151 similar to UP|Q94AX4 (Q94AX4) AT5g06580/F15M7_11, parti... 85 7e-17
TC223051 similar to UP|O23240 (O23240) Actin interacting protein... 65 1e-10
BE805756 similar to GP|16604326|gb| At4g36400/C7A10_960 {Arabido... 63 4e-10
TC226773 homologue to UP|Q9ATR0 (Q9ATR0) Brassinosteroid biosynt... 33 0.26
TC232674 similar to UP|Q9FUJ1 (Q9FUJ1) Cytokinin oxidase (FAD-li... 33 0.33
TC224843 homologue to UP|Q8RUR2 (Q8RUR2) ADP-ribosylation factor... 33 0.44
BF069714 similar to GP|5059126|gb|A cytochrome P450 monooxygenas... 32 0.74
TC204376 similar to UP|Q9SVG3 (Q9SVG3) Reticuline oxidase-like p... 32 0.74
BG352198 GP|20161472|db ADP-ribosylation factor {Oryza sativa (j... 30 2.8
TC222573 weakly similar to UP|Q7QHR9 (Q7QHR9) EbiP8208 (Fragment... 30 2.8
BM308215 29 4.8
TC229733 similar to UP|Q84YI1 (Q84YI1) Polyphenol oxidase , pa... 29 4.8
AW395000 29 6.3
TC215730 similar to UP|Q9FN50 (Q9FN50) Emb|CAB62636.1 (AT5g23040... 28 8.2
AW706428 weakly similar to GP|11994643|dbj UTP-glucose glucosylt... 28 8.2
TC213838 similar to PIR|E86432|E86432 T5I8.15 protein - Arabidop... 28 8.2
>BU762717
Length = 415
Score = 249 bits (636), Expect = 2e-66
Identities = 125/137 (91%), Positives = 132/137 (96%)
Frame = +3
Query: 236 NVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSL 295
NVSTNAGGLRLVRYGSLHGSVLGVEAVLA+GTVLDMLKTLRKDNTGYDLKHLFIGSEGSL
Sbjct: 3 NVSTNAGGLRLVRYGSLHGSVLGVEAVLANGTVLDMLKTLRKDNTGYDLKHLFIGSEGSL 182
Query: 296 GIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVI 355
GIVT V+ILTPPKLSSVNVA LACKDY+ CQKLLQEAK KLGEILSAFEFLD QSMNLV+
Sbjct: 183 GIVTIVSILTPPKLSSVNVAFLACKDYSSCQKLLQEAKGKLGEILSAFEFLDVQSMNLVL 362
Query: 356 NHLEGARNPLPTLHDFY 372
NH+EGARNPLP+LH+FY
Sbjct: 363 NHMEGARNPLPSLHNFY 413
>TC219979 similar to UP|O23240 (O23240) Actin interacting protein, partial
(19%)
Length = 535
Score = 180 bits (457), Expect = 1e-45
Identities = 87/106 (82%), Positives = 93/106 (87%)
Frame = +2
Query: 452 EMRSRLGNAANVVGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHG 511
E R+ + ANV+GYGHLGDGNLHLNIS YDDKILS IEP+VYEWTSKH GSISAEHG
Sbjct: 5 EASCRIRHEANVIGYGHLGDGNLHLNISTSHYDDKILSHIEPYVYEWTSKHRGSISAEHG 184
Query: 512 LGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLPHSLIA 557
LGLMKANEI YSKS ETVQ+MASIKNLLDPNHILNPYKVLP SLI+
Sbjct: 185 LGLMKANEIFYSKSHETVQVMASIKNLLDPNHILNPYKVLPQSLIS 322
>BE824336 similar to GP|16604326|gb| At4g36400/C7A10_960 {Arabidopsis
thaliana}, partial (25%)
Length = 497
Score = 124 bits (311), Expect(2) = 1e-36
Identities = 61/72 (84%), Positives = 65/72 (89%)
Frame = -2
Query: 486 KILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHIL 545
+ILS I +VYEWTSKH GSISAEHGLGLMKANEI YSKS ETVQ+MASIKNLLDPNHIL
Sbjct: 316 QILSHIXXYVYEWTSKHRGSISAEHGLGLMKANEIFYSKSHETVQVMASIKNLLDPNHIL 137
Query: 546 NPYKVLPHSLIA 557
NPYKVLP SLI+
Sbjct: 136 NPYKVLPQSLIS 101
Score = 47.8 bits (112), Expect(2) = 1e-36
Identities = 20/30 (66%), Positives = 22/30 (72%)
Frame = -3
Query: 458 GNAANVVGYGHLGDGNLHLNISVPQYDDKI 487
G ANV+GYG DGNLHLNIS YDDK+
Sbjct: 480 GXXANVIGYGXXXDGNLHLNISTSHYDDKV 391
>TC207151 similar to UP|Q94AX4 (Q94AX4) AT5g06580/F15M7_11, partial (51%)
Length = 1075
Score = 85.1 bits (209), Expect = 7e-17
Identities = 85/300 (28%), Positives = 138/300 (45%), Gaps = 17/300 (5%)
Frame = +2
Query: 270 DMLKTL---RKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSV-NVALLACKDYNCC 325
D++KT RK GYDL L IGSEG+LG++T+V + +L + +++A ++
Sbjct: 2 DIVKTASRARKSAAGYDLTRLMIGSEGTLGVITEVTL----RLQKIPQYSVVAMCNFPSV 169
Query: 326 QKLLQEAKRKL--GEILSAFEFLDSQSMNLVINHLEGARNP-LPTLHDFYVLIETTGSDE 382
+ A + G +S E LD + IN G P PTL + E G++
Sbjct: 170 KDAADVAIATMMSGIQVSRVELLDEVQVK-AINIANGKNLPECPTL-----MFEFIGTEA 331
Query: 383 SSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAV------YK 436
+ +Q L S N SD V A++ W++R+ EAL A+
Sbjct: 332 YAREQTQIVRKLVSEHNG--SDFVFAEEPEAKKELWKVRK---EALWACFAMEPNLVAMT 496
Query: 437 YDLSIPLENLYNLVEEMRSRLGNAANVVGY-GHLGDGNLHLNISV-PQYDD--KILSQIE 492
D+ +PL +L +L+ + L + V H GDGN H I P ++ + ++
Sbjct: 497 TDVCVPLSHLGDLISRSKKELDASPLVCTVIAHAGDGNFHTVILFDPNQEEQRREAERLN 676
Query: 493 PFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
F+ G+ + EHG+G K + +E ++ M IK +LDPN I+NP K++P
Sbjct: 677 QFMVHAALSLEGTCTGEHGVGTGKMKYLEEELGVEALRTMKKIKAVLDPNDIMNPGKLIP 856
>TC223051 similar to UP|O23240 (O23240) Actin interacting protein, partial
(7%)
Length = 577
Score = 64.7 bits (156), Expect = 1e-10
Identities = 32/37 (86%), Positives = 34/37 (91%)
Frame = +3
Query: 386 KQKLEAFLLHSMENELISDGVLAQDINQASSFWRLRE 422
+QK EAFLL SMENE+ISDGVLAQDINQASSFW LRE
Sbjct: 219 RQKPEAFLLGSMENEMISDGVLAQDINQASSFWLLRE 329
>BE805756 similar to GP|16604326|gb| At4g36400/C7A10_960 {Arabidopsis
thaliana}, partial (8%)
Length = 393
Score = 62.8 bits (151), Expect = 4e-10
Identities = 31/36 (86%), Positives = 33/36 (91%)
Frame = +3
Query: 386 KQKLEAFLLHSMENELISDGVLAQDINQASSFWRLR 421
+QK EAFLL SMENE+ISDGVLAQDINQASSFW LR
Sbjct: 90 RQKPEAFLLGSMENEMISDGVLAQDINQASSFWLLR 197
>TC226773 homologue to UP|Q9ATR0 (Q9ATR0) Brassinosteroid biosynthetic
protein LKB, partial (86%)
Length = 1584
Score = 33.5 bits (75), Expect = 0.26
Identities = 21/60 (35%), Positives = 30/60 (50%)
Frame = +1
Query: 248 RYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPP 307
+YG +V+ E +LADGT L KDN DL + S+G+LG++ I P
Sbjct: 652 KYGLFADTVVAYEIILADGT----LVRATKDNEYTDLYYAIPWSQGTLGLLVAAEIKLIP 819
>TC232674 similar to UP|Q9FUJ1 (Q9FUJ1) Cytokinin oxidase (FAD-linked
oxidoreductase family), partial (28%)
Length = 779
Score = 33.1 bits (74), Expect = 0.33
Identities = 32/173 (18%), Positives = 67/173 (38%), Gaps = 3/173 (1%)
Frame = +2
Query: 134 KLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNK---VI 190
+ +++P V + +K + V GN + G +++ + +M ++
Sbjct: 206 RAVIRPALAGDVERAVKEAARTTYLTVAARGNGHSINGQAMAEKGLVLDMRAMEDHFTLL 385
Query: 191 SFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYG 250
S D S + G + E+++ +E + P +GG +S + RYG
Sbjct: 386 SLDDGSLYVDVSGGALWEDVLKRCVSEFRLAPRSWTDYLGLTVGGTLSNAGVSGQAFRYG 565
Query: 251 SLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAI 303
+V +E V G L + ++ +L +G G GI+T+ +
Sbjct: 566 PQTANVTELEVVSGKGETL-----VCSESQNSELFFATLGGLGQFGIITRARV 709
>TC224843 homologue to UP|Q8RUR2 (Q8RUR2) ADP-ribosylation factor, partial
(48%)
Length = 1021
Score = 32.7 bits (73), Expect = 0.44
Identities = 44/176 (25%), Positives = 69/176 (39%), Gaps = 4/176 (2%)
Frame = -1
Query: 285 KHLFIGSEGSLGIVTKVAILTPPKL-SSVNVALLACKDYNCCQKLLQEAKRKLGEILSAF 343
K LF+G I + I P +L S +N ++ YN + L
Sbjct: 493 KILFVGKHKQYCISQFILIQHPMQLISGLNNSVPIIAVYN------------KNKTLGVL 350
Query: 344 EFLDSQSMNLVI-NHLEGARNPLPTLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELI 402
E + Q NLV+ ++ + + H IE G + +D +LE L+
Sbjct: 349 EVVPPQGTNLVLATNIPNSEANVLVFHSLN--IEPDGRNCGNDLSELE----------LV 206
Query: 403 SDGVLAQDI--NQASSFWRLREGITEALMRVGAVYKYDLSIPLENLYNLVEEMRSR 456
D L I N +S + LRE TE L + +DL IPLE ++ + + R
Sbjct: 205 QDSGLTSSIKTNHQNSHFLLREKPTEKLRECQPHFSFDLLIPLEMKCSVCRKRKER 38
>BF069714 similar to GP|5059126|gb|A cytochrome P450 monooxygenaseCYP93D1
{Glycine max}, partial (9%)
Length = 157
Score = 32.0 bits (71), Expect = 0.74
Identities = 17/48 (35%), Positives = 25/48 (51%)
Frame = +2
Query: 334 RKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSD 381
R++GE+L+AF D + ++ N PTLH F +IE SD
Sbjct: 14 REVGELLAAFNLWDVIEFMMPLDLQRFRTNNFPTLHKFDTMIEKVLSD 157
>TC204376 similar to UP|Q9SVG3 (Q9SVG3) Reticuline oxidase-like protein,
partial (63%)
Length = 1418
Score = 32.0 bits (71), Expect = 0.74
Identities = 32/135 (23%), Positives = 55/135 (40%), Gaps = 2/135 (1%)
Frame = +1
Query: 175 VFDEVIVSLSSMN-KVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQI 233
+ +E V L N + I+ D + + V EAG L + + + ++ G + +
Sbjct: 25 ISEEPFVILDMFNYRRITVDVKNEVAVVEAGATLGEVYYRIWEKSKVLGFPAGVCPTVGV 204
Query: 234 GGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEG 293
GG+ S G L +YG +V+ + V G +L+ + G DL G G
Sbjct: 205 GGHFSGGGYGNMLRKYGLSVDNVIDAQIVDVKGNLLN------RKTMGEDLFWAIRGGGG 366
Query: 294 -SLGIVTKVAILTPP 307
S G++ I P
Sbjct: 367 ASFGVILSFTIKLVP 411
>BG352198 GP|20161472|db ADP-ribosylation factor {Oryza sativa (japonica
cultivar-group)}, partial (19%)
Length = 471
Score = 30.0 bits (66), Expect = 2.8
Identities = 25/84 (29%), Positives = 37/84 (43%), Gaps = 2/84 (2%)
Frame = -1
Query: 375 IETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDI--NQASSFWRLREGITEALMRVG 432
IE G + S+D +LE L+ D L I N +S + LRE E L
Sbjct: 246 IEPNGRNCSNDLSELE----------LVEDSGLTSSIKTNHQNSHFLLREKPAEKLRECQ 97
Query: 433 AVYKYDLSIPLENLYNLVEEMRSR 456
+ +DL IPLE ++ + + R
Sbjct: 96 PHFSFDLLIPLEMKCSVCRKRKER 25
>TC222573 weakly similar to UP|Q7QHR9 (Q7QHR9) EbiP8208 (Fragment), partial
(17%)
Length = 432
Score = 30.0 bits (66), Expect = 2.8
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = -2
Query: 38 SQTNIAHGEHRVSSLFNPLRFSVKN 62
+QT HG+ R+SSLFNPL KN
Sbjct: 116 TQTLTNHGKVRISSLFNPLSLVQKN 42
>BM308215
Length = 415
Score = 29.3 bits (64), Expect = 4.8
Identities = 19/82 (23%), Positives = 35/82 (42%)
Frame = +1
Query: 141 NTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILV 200
N +V ++ +S V P G N +++S ++NK++ DK + +
Sbjct: 202 NKRKVKAATRFSHSIPKLVCPDGENG------------LLISTKNLNKILKIDKEARTMT 345
Query: 201 CEAGCILENIISFLDNEGFIMP 222
++G L IIS G +P
Sbjct: 346 VQSGVSLREIISKGAEAGLALP 411
>TC229733 similar to UP|Q84YI1 (Q84YI1) Polyphenol oxidase , partial (28%)
Length = 1170
Score = 29.3 bits (64), Expect = 4.8
Identities = 17/66 (25%), Positives = 29/66 (43%), Gaps = 11/66 (16%)
Frame = +1
Query: 13 TPFFNPRRNPSLLPST-----------PHFDENWWHSQTNIAHGEHRVSSLFNPLRFSVK 61
+P ++PRRNP + P+T P ++N T++ G R S +VK
Sbjct: 829 SPLYDPRRNPDITPTTLVDLNYGRGKEPSVEQNLGVMYTSVDSGAKRASLFHGKPFLAVK 1008
Query: 62 NQQMGV 67
++ V
Sbjct: 1009QPELRV 1026
>AW395000
Length = 315
Score = 28.9 bits (63), Expect = 6.3
Identities = 19/80 (23%), Positives = 35/80 (43%)
Frame = +1
Query: 110 NVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLV 169
+V ++ L N+D+M Y G +K+ + P N + + + V PQ NT V
Sbjct: 4 DVYPRQQGLLGYNSDYMVWYRGKTKMFVDP-NNANTTALGDVVETL*YMVSPQWRNTWTV 180
Query: 170 GGSVPVFDEVIVSLSSMNKV 189
VP D++ + ++
Sbjct: 181 DDLVPYVDKLAIISEEQERI 240
>TC215730 similar to UP|Q9FN50 (Q9FN50) Emb|CAB62636.1 (AT5g23040/MYJ24_3),
partial (82%)
Length = 1255
Score = 28.5 bits (62), Expect = 8.2
Identities = 13/33 (39%), Positives = 22/33 (66%)
Frame = -3
Query: 297 IVTKVAILTPPKLSSVNVALLACKDYNCCQKLL 329
+ TK+AI K +S++ +LL+C +Y C +K L
Sbjct: 581 LCTKLAINIFSKAASIDSSLLSCPEYCCNRKFL 483
>AW706428 weakly similar to GP|11994643|dbj UTP-glucose glucosyltransferase
{Arabidopsis thaliana}, partial (7%)
Length = 331
Score = 28.5 bits (62), Expect = 8.2
Identities = 17/51 (33%), Positives = 24/51 (46%), Gaps = 1/51 (1%)
Frame = -1
Query: 14 PFFNPRRNPSLLPSTPH-FDENWWHSQTNIAHGEHRVSSLFNPLRFSVKNQ 63
PFF P R+P+LL P E Q + H + RV + + +V NQ
Sbjct: 172 PFFRPHRHPNLLHIHPQLLVEKMLRKQRSCCHRQPRVHAFHLRVPTTVSNQ 20
>TC213838 similar to PIR|E86432|E86432 T5I8.15 protein - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (33%)
Length = 543
Score = 28.5 bits (62), Expect = 8.2
Identities = 26/114 (22%), Positives = 49/114 (42%), Gaps = 3/114 (2%)
Frame = +3
Query: 202 EAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEA 261
+AG L + + + G + +GG++S G + +YG +V+ +
Sbjct: 147 QAGATLGEVYYRIAEKSKTHAFPAGVCHTVGVGGHISGGGYGNMMRKYGLSVDNVIDAQM 326
Query: 262 VLADGTVLDMLKTLRKDNTGYDLKHLFIGSEG-SLGIVT--KVAILTPPKLSSV 312
V G +LD + + G DL G G S G+V K+ ++ P++ +V
Sbjct: 327 VDVQGRLLD------RKSMGEDLFWAITGGGGASFGVVLAYKIKLVRVPEIVTV 470
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,997,286
Number of Sequences: 63676
Number of extensions: 322991
Number of successful extensions: 1629
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1627
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1628
length of query: 557
length of database: 12,639,632
effective HSP length: 102
effective length of query: 455
effective length of database: 6,144,680
effective search space: 2795829400
effective search space used: 2795829400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0627.15


