

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0593a.3
(562 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
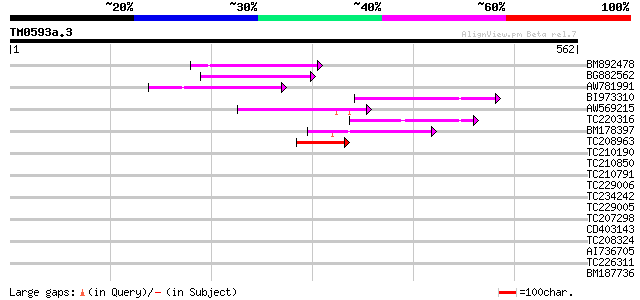
Score E
Sequences producing significant alignments: (bits) Value
BM892478 99 7e-21
BG882562 97 1e-20
AW781991 97 2e-20
BI973310 76 3e-14
AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis... 75 8e-14
TC220316 75 1e-13
BM178397 similar to GP|4914326|gb| F14N23.12 {Arabidopsis thalia... 61 1e-09
TC208963 42 7e-04
TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 35 0.12
TC210850 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 34 0.15
TC210791 34 0.20
TC229006 33 0.44
TC234242 32 0.98
TC229005 30 2.2
TC207298 30 3.7
CD403143 30 3.7
TC208324 similar to UP|WRK6_ARATH (Q9C519) WRKY transcription fa... 29 4.9
AI736705 similar to GP|11994736|db far-red impaired response pro... 29 6.4
TC226311 similar to UP|Q84TR6 (Q84TR6) Subtilase, partial (18%) 29 6.4
BM187736 similar to GP|13506731|gb| WRKY DNA-binding protein 18 ... 28 8.3
>BM892478
Length = 429
Score = 98.6 bits (244), Expect = 7e-21
Identities = 48/131 (36%), Positives = 70/131 (52%)
Frame = +3
Query: 180 LSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNK 239
L Y Q + +D FF+ N N+FW DG SR FGDV+V ++Y+K Y
Sbjct: 33 LEYFQSRQAEDTGFFYAM-EVDNGNCMNIFWADGRSRYSCSHFGDVLVLDTSYRKTVYLV 209
Query: 240 PVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAA 299
P F G NHH + + CAL+ DE+ E++ W+ + AM G+ P V+ D D +++ A
Sbjct: 210 PFATFVGVNHHKQPVLLGCALIADESEESFTWLFQTWLRAMSGRLPLTVIADQDIAIQRA 389
Query: 300 VKVVFPNATHR 310
+ VFP HR
Sbjct: 390 IAKVFPVTHHR 422
>BG882562
Length = 416
Score = 97.4 bits (241), Expect = 1e-20
Identities = 44/114 (38%), Positives = 66/114 (57%)
Frame = +3
Query: 190 DPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNH 249
+P FF+ + + L N+FW D SR Y FGDV+ F ST N Y P+V F G NH
Sbjct: 75 NPNFFYVMDLNDDGQLRNVFWIDSRSRAAYSYFGDVVAFDSTCLSNNYEIPLVAFVGVNH 254
Query: 250 HTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVV 303
H ++ + C L+ DET ETY W+ +A M G+ P+ ++ + K+M++A+ V
Sbjct: 255 HGKSVLLGCGLLADETFETYIWLFRAWLTCMTGRPPQTIITNXCKAMQSAIAEV 416
>AW781991
Length = 419
Score = 96.7 bits (239), Expect = 2e-20
Identities = 45/137 (32%), Positives = 76/137 (54%)
Frame = +3
Query: 138 ITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGEKDPMFFFKF 197
I L+ + GG VGF + D N++ + SL GD L YL+ ++P FF+
Sbjct: 6 IMSALIKEYGGISKVGFTEVDCRNYMRNNRLRSL-EGDIQLVLDYLRQMHAENPNFFYAV 182
Query: 198 TRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFA 257
++++ N+FW D +R++Y FGD + F +TY+ N+Y P F+G NHH + +F
Sbjct: 183 QGDEDQSITNVFWADPKARMNYTFFGDTVTFDTTYRSNRYRLPFAFFTGVNHHGQPVLFG 362
Query: 258 CALVCDETIETYKWVLK 274
CA + +E+ ++ W+ K
Sbjct: 363 CAFLINESEASFVWLFK 413
>BI973310
Length = 457
Score = 76.3 bits (186), Expect = 3e-14
Identities = 39/145 (26%), Positives = 74/145 (50%)
Frame = +3
Query: 342 TPERFETKWLQVIEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFI 401
T + FE+ W ++E++ + + +W++ +Y +RQ W ++RE FF I E + SF
Sbjct: 3 TIDEFESYWHSLLERFYVMDNEWLQSIYNSRQHWVPVYLRETFFGEISLNEGNEYLISFF 182
Query: 402 KSYVLCKNSILDFIYNFERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLN 461
YV ++ + +E+AV + EL +D+ + PVL T S +E+ A+ L T
Sbjct: 183 DGYVFSSTTLQVLVRQYEKAVSSWHEKELKADYDTTNSSPVLKTP-SPMEKQAASLYTRK 359
Query: 462 MFREMRHMIQRALNLNLVERSEIGT 486
+F + + + L ++ + GT
Sbjct: 360 IFMKFQEELVETLANPAIKIDDSGT 434
>AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis thaliana},
partial (19%)
Length = 425
Score = 75.1 bits (183), Expect = 8e-14
Identities = 41/138 (29%), Positives = 72/138 (51%), Gaps = 5/138 (3%)
Frame = +1
Query: 226 IVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQP 285
+VF +TY+ Y+ + + G +++ T F+CAL+ DE I+++ W LKA M GK P
Sbjct: 10 VVFDTTYRVEAYDMLLGIWLGVDNNGMTCFFSCALLRDENIQSFSWALKAFLGFMKGKAP 189
Query: 286 KAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCLE--KIKIPDFLNEFKT---LIYGN 340
+ ++ D + ++ A+ V P H C WHI + + + +E+K +Y
Sbjct: 190 QTILTDHNMWLKEAIAVELPQTKHAFCIWHILSKFSDWFSLLLGSQYDEWKAEFHRLYNL 369
Query: 341 FTPERFETKWLQVIEKYG 358
E FE W Q++++YG
Sbjct: 370 EQVEDFEEGWRQMVDQYG 423
>TC220316
Length = 988
Score = 74.7 bits (182), Expect = 1e-13
Identities = 37/127 (29%), Positives = 67/127 (52%)
Frame = +1
Query: 338 YGNFTPERFETKWLQVIEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGI 397
Y + T + W KYG+ ++ W+K+MY+ R W +++ FFAGI + E +
Sbjct: 10 YQSQTVDEXXATWNXXXNKYGLKDDAWLKEMYQKRASWVPLYLKGTFFAGI---PMNESL 180
Query: 398 NSFIKSYVLCKNSILDFIYNFERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKL 457
+SF + + + +++FI +ER +E R E DF + +P+L T +E+ +L
Sbjct: 181 DSFFGALLNAQTPLMEFIPRYERGLERRREEERKEDFNTSNFQPILQTK-EPVEEQCRRL 357
Query: 458 LTLNMFR 464
TL +F+
Sbjct: 358 YTLTVFK 378
>BM178397 similar to GP|4914326|gb| F14N23.12 {Arabidopsis thaliana}, partial
(20%)
Length = 422
Score = 61.2 bits (147), Expect = 1e-09
Identities = 35/134 (26%), Positives = 63/134 (46%), Gaps = 6/134 (4%)
Frame = +1
Query: 296 MRAAVKVVFPNATHRLCGWHIQQ------NCLEKIKIPDFLNEFKTLIYGNFTPERFETK 349
++ A+ P H C W I N + + D+ EF L Y + E FE
Sbjct: 13 LKEALSTEMPTTKHAFCIWMIVAKFPSWFNAVLGERYNDWKAEFYRL-YNLESVEDFELG 189
Query: 350 WLQVIEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKN 409
W ++ +G+ + + +Y +R +WA F+R +F AG+ TT + IN+FI+ ++ +
Sbjct: 190 WREMACSFGLHSNRHMVNLYSSRSLWALPFLRSHFLAGMTTTGQSKSINAFIQRFLSAQT 369
Query: 410 SILDFIYNFERAVE 423
+ F+ AV+
Sbjct: 370 RLAHFVEQVAVAVD 411
>TC208963
Length = 1163
Score = 42.0 bits (97), Expect = 7e-04
Identities = 21/53 (39%), Positives = 34/53 (63%)
Frame = +1
Query: 285 PKAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCLEKIKIPDFLNEFKTLI 337
P+ +V GD ++ +AV+VVFP++++ LC +HI QN K K L E + L+
Sbjct: 22 PQVIVTVGDIALMSAVQVVFPSSSNLLCRFHINQNVKAKCKSIVHLKEKQELM 180
>TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (65%)
Length = 866
Score = 34.7 bits (78), Expect = 0.12
Identities = 19/43 (44%), Positives = 21/43 (48%)
Frame = +1
Query: 62 RIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTP 104
+I + TRV C A VR S KW V F EH H LTP
Sbjct: 172 KIVRQRAETRVGCRAMIMVR-KLSSGKWVVAKFVKEHTHPLTP 297
>TC210850 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (63%)
Length = 1326
Score = 34.3 bits (77), Expect = 0.15
Identities = 18/43 (41%), Positives = 21/43 (47%)
Frame = +1
Query: 62 RIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTP 104
+I + TRV C A VR S KW + F EH H LTP
Sbjct: 154 KIVRQRAETRVGCRAMIMVR-KLSSGKWVIAKFVKEHTHPLTP 279
>TC210791
Length = 456
Score = 33.9 bits (76), Expect = 0.20
Identities = 14/44 (31%), Positives = 28/44 (62%)
Frame = +1
Query: 283 KQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCLEKIKI 326
+ P ++ D ++ AV+VVFP++++ LC +HI +N K ++
Sbjct: 25 EMPPVIITVRDIALMDAVQVVFPSSSNLLCRFHISKNVKAKSQV 156
>TC229006
Length = 991
Score = 32.7 bits (73), Expect = 0.44
Identities = 13/40 (32%), Positives = 23/40 (57%)
Frame = +2
Query: 63 IKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGL 102
+++ + TR C A ++++ KS KW + F +HNH L
Sbjct: 293 VRKPRASTREGCKAMIHIKYN-KSGKWVITKFVKDHNHPL 409
>TC234242
Length = 478
Score = 31.6 bits (70), Expect = 0.98
Identities = 14/43 (32%), Positives = 24/43 (55%)
Frame = +3
Query: 202 EENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNF 244
E+ + ++FWC S A V + STY+ N+Y P+++F
Sbjct: 81 EDVVRDIFWCHPDSVKLVNACNLVFLIDSTYRTNRYRLPLLDF 209
>TC229005
Length = 679
Score = 30.4 bits (67), Expect = 2.2
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +1
Query: 70 TRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGL 102
TR C A +++ KS KW + F +HNH L
Sbjct: 7 TREGCKAMIHIKYD-KSGKWVIAKFVKDHNHPL 102
>TC207298
Length = 1011
Score = 29.6 bits (65), Expect = 3.7
Identities = 11/28 (39%), Positives = 19/28 (67%)
Frame = +3
Query: 204 NLENLFWCDGTSRLDYQAFGDVIVFYST 231
++++ FW SR+D+ FGD++VF T
Sbjct: 342 HIQHTFWLICLSRVDWWQFGDLVVFQGT 425
>CD403143
Length = 341
Score = 29.6 bits (65), Expect = 3.7
Identities = 17/43 (39%), Positives = 27/43 (62%), Gaps = 1/43 (2%)
Frame = -2
Query: 500 TQLSSWSFMTRIRTCLIVIVGFLNMLAY-LVLTLYVL*ELRT* 541
T LS W +T + CL+ + G L +++Y L+L+L L +L T*
Sbjct: 181 TVLSLWRIITPLFYCLLTVTGLLCLISY*LILSLINLPQL*T* 53
>TC208324 similar to UP|WRK6_ARATH (Q9C519) WRKY transcription factor 6 (WRKY
DNA-binding protein 6) (AtWRKY6), partial (19%)
Length = 1638
Score = 29.3 bits (64), Expect = 4.9
Identities = 12/34 (35%), Positives = 19/34 (55%)
Frame = +2
Query: 73 SCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAA 106
+CP + +V+ A+ + +E HNH L PAA
Sbjct: 770 ACPVRKQVQRCAEDESVVITTYEGNHNHSLPPAA 871
>AI736705 similar to GP|11994736|db far-red impaired response protein;
Mutator-like transposase-like protein; phytochrome A
signaling, partial (14%)
Length = 383
Score = 28.9 bits (63), Expect = 6.4
Identities = 14/39 (35%), Positives = 18/39 (45%)
Frame = +3
Query: 70 TRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHV 108
++ C A V+ KW + F EHNH L PA V
Sbjct: 75 SKTDCKASMHVK-RRSDGKWVIHSFVKEHNHELLPAQAV 188
>TC226311 similar to UP|Q84TR6 (Q84TR6) Subtilase, partial (18%)
Length = 943
Score = 28.9 bits (63), Expect = 6.4
Identities = 20/41 (48%), Positives = 27/41 (65%)
Frame = -2
Query: 509 TRIRTCLIVIVGFLNMLAYLVLTLYVL*ELRT*MNSLLILF 549
T +R L+V+VGFL + AYL+ T *EL T* N+ L +F
Sbjct: 225 TNVR*NLMVVVGFLWLFAYLI-TDDAS*ELIT*ENNNLSIF 106
>BM187736 similar to GP|13506731|gb| WRKY DNA-binding protein 18 {Arabidopsis
thaliana}, partial (26%)
Length = 437
Score = 28.5 bits (62), Expect = 8.3
Identities = 12/29 (41%), Positives = 16/29 (54%)
Frame = +3
Query: 73 SCPAKFRVRFHAKSSKWKVVLFEPEHNHG 101
SCP K +V+ + V +E EHNHG
Sbjct: 216 SCPVKKKVQRSVEDPSVLVTTYEGEHNHG 302
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.331 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,750,275
Number of Sequences: 63676
Number of extensions: 361458
Number of successful extensions: 2281
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 2269
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2278
length of query: 562
length of database: 12,639,632
effective HSP length: 102
effective length of query: 460
effective length of database: 6,144,680
effective search space: 2826552800
effective search space used: 2826552800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0593a.3


